Professional Documents
Culture Documents
S3T SBoytcheva EPaskaleva IvNikolova GAngelova-Extracting Patient-Related Description From Medical Records in Bulgarian
Uploaded by
Mulya Nurmansyah ArdisasmitaCopyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
S3T SBoytcheva EPaskaleva IvNikolova GAngelova-Extracting Patient-Related Description From Medical Records in Bulgarian
Uploaded by
Mulya Nurmansyah ArdisasmitaCopyright:
Available Formats
Extracting Patient-Related Description from
Medical Records in Bulgarian
Svetla Boytcheva1, Elena Paskaleva2, Ivelina Nikolova2, and Galia Angelova2
1
State University of Library Studies and Information Technologies
119 Tzarigradsko Shose Blvd., 1784 Sofia, Bulgaria,
svetla.boytcheva@gmail.com
2
Institute for Parallel Processing, Bulgarian Academy of Sciences
25A Acad. G. Bonchev Str., 1113 Sofia, Bulgaria,
{hellen,iva,galia}@lml.bas.bg
Abstract. This paper deals with the extraction of medical information
from hospital patient records. It proposes a cascade approach for the
extraction of multi-layer knowledge statements because the subject is
too complex. We sketch the Information Extraction view to text analysis, where patient-related facts are recognised using predefined regular
expressions and templates. A laboratory prototype for patient status extraction is presented together with the first evaluation results.
1 Introduction
Patient-related medical records, which are supported by personal GPs, hospitals, medical examining commissions etc., remain the most important source of
information about diseases, treatments, case histories, and medication effects.
Unfortunately these records contain unstructured data presented in a variety of
formats: free texts in natural language, numeric values of clinical data, images,
graphics and tables. In addition, the documents are kept as disconnected sets of
files and databases, including items in large paper archives. All these complications make the automated analysis of medical records a challenging task.
Recent advances of language technologies provided a new perspective to
medical text processing. Great efforts have been made to translate the information, which is locked-up into personal medical documents, to certain semistructured representations; one of the primary research task is to develop software tools for semi-automatic information extraction from free texts. It turns
out, however, that the medical language is rather complex since there are no
conventions and standardisation of the wording and expressions in the textual
patient descriptions. In addition the automatic text processing requires substantial background knowledge about the sophisticated medical domain but the acquisition of the necessary conceptual resources is difficult and very expensive.
Another obstacle is the variety of practices for including text descriptions in
the patient records, which are too specific for different countries and different
languages. Despite all these complications there are numerous projects in the
challenging domain of medical text processing.
Here we present the initial results of a project which deals with automatic
text processing of patient records in Bulgarian language. This article is
structured as follows. Section 2 overviews some related work, including the
basic Natural Language Processing (NLP) techniques which provide medical
22
S. Boytcheva, E. Paskaleva, Iv. Nikolova, and G. Angelova
Information Extraction (IE). Section 3 sketches the general project framework
of patient-related knowledge acquisition and discusses the raw data features.
Section 4 presents the patient status data, some techniques for their extraction
and evaluation of the current experiments. Section 5 contains the conclusion.
2 Related work
In general the approaches for automatic processing of patient-related texts can
be grouped in two categories:
(i) Tools for automatic extraction of diagnosis, treatment, medications, manipulations etc. in order to encode the information with respect to some establish1ed classification schemes, which are provided by financing or statistical
institutions. These approaches apply large terminology-based nomenclatures
such as SNOMED (the Systematized Nomenclature of Medicine) and ICD (the
International Classifcation of Diseases), which are uniffied multilingual classification systems, and are developed to support the health management and
health statistics. Usually the extraction is focused on the recognition of medical terms in the free text of patient records. The industrial systems for automatic analysis of medical documents belong primarily to this category of tools.
However, the software is far from being perfect; successful recognition of the
complex medical terminology is possible for some 70% of the terms, which are
increased up to 90% after human intervention and editing [1];
(ii) Research prototypes for studying the application of language technologies in the medical domain or prototypes, oriented to medical research and
knowledge discovery in medicine. These approaches reflect the AI view to text
processing and aim to translate the text to internal structured representations,
to make inferences, to discover interconnections between facts and concepts
which could remain unnoticed otherwise, and to spot previously unknown
regularities. This research is oriented mostly to medical texts in English and
integrates some public linguistic resources and language technologies as well
as large public archives of medical abstracts. Certain investigations are oriented
to deeper study of isolated constructions, e.g. extraction of causal-effect relations in the medical domain [2]. This kind of research prototypes often remain
as isolated laboratory systems which are not integrated in holistic software solutions.
We consider in more depth the Information Extraction approach, which is
relevant for our research work. IE relies on partial text analysis in order to
extract only relevant facts from the document, ignoring the remaining parts
which are considered irrelevant [3]. Usually the IE systems are set to recognise
specific facts only, by knowing in advance the words that may signal these facts
in the text. Thus, for instance, an IE module can analyse patient records in order
to discover the status of patient skin. Main IE tasks are Named Entity recognition, Coreference resolution, Template construction and Template filling, the
latter being the process of mapping the text elements to template fields, which is
performed in general with accuracy of less than 70% [4]. Various systems employ the IE approach to medical texts in different analysis stages, for instance
CLEF (Clinical E-Science Framework) for extracting data from clinical records
of cancer patients [5], AMBIT (Acquiring Medical and Biomedical Information
from Text) [6] and MiTAP (MITRE Text and Audio Processing) for monitoring
Extracting Patient-Related Description from Medical Records in Bulgarian
23
infectious disease outbreaks and other global events [7]. The IE partial analysis
is often based on pattern matching involving cascade regular expressions which
are defined in terms of lexical categories and/or particular important words.
The manually-produced patterns provide better matches but their adaptation to
new domains is difficult. Some patterns can be extracted semi-automatically
by general meta-rules but they are not too precise. Medical IE is a hot research
area, to be explored independently for every natural language.
We note that partial text analysis is typical for many research prototypes beyond the IE context, e.g. one could extract and analyse only data about the patient smoking status using machine learning techniques [8]. Other studies concern complex linguistic constructions like the negation in medical texts [9].
3 Project settings - from data to knowledge
The project investigates information extraction from hospital records of patients, who are diagnosed with diabetes and are treated in the Specialised University Hospital for Active Treatment in Endocrinology, in the Clinical Centre
of Endocrinology and Gerontology - Sofia. The hospital information system
integrates most clinical data but is in use only recently, so family histories are
not supported. Perhaps the most significant task is to analyse the hospitalisation
effects: how the treatment affects a patient who enters the hospital with status
A and leaves it with status B. In this way the automatic extraction of information concerning the patient status is a very important task. Our research corpus
consists of pseudonymised patient records, which can be linked to records of
previous patient visits to the same hospital.
One of the first essential findings is that the analysis of text words and sentences is insufficient since the temporal and causal relations form the core of
the treatment descriptions. Therefore we have designed a 4-layered view to
the patient record content: (i) at the lowest level, text words/phrases/sentences
are analysed and specific templates are filled in by information about different
states and events; (ii) temporal interdependencies between the states and the
events are to be recognised at layer 2; (iii) causal-effect relations are recognised
at layer 3 and (iv) more complex implicit facts are inferred at layer 4. This ambitious scenario exploits the internal structure of the patient record documents,
which are divided into sections. Certainly, we do not aim at a full discovery
of all patient-related information; the project will focus on certain important
aspects only. More details about the multi-layered conceptual representation
are given in [10].
Processing patient records in Bulgarian is far from trivial because of the
variety of terminological expressions. About 1% of the medical terminology
is given in Latin, sometimes with different transcriptions in Cyrillic. Several
wordforms per term are used for most of the medical terminology in Bulgarian
(66% of the terms) as well as term abbreviations both in Bulgarian and Latin
(for about 3% of the term tokens). Some 16% of the tokens are numerical values of clinical test data. The major part of the text consists of sentence phrases,
especially in some sections, without agreement between the sentence parts.
Especially for Bulgarian, there is a lack of well-developed, stable language
technologies with satisfactory precision for partial syntax analysis. About 2/3
of the whole text describe connected states and events, which have happened
24
S. Boytcheva, E. Paskaleva, Iv. Nikolova, and G. Angelova
to the patient in different periods of time. We have already said that the rich
temporal structure of the patient records requires extracting the information
in multi-layered internal representations. Due to all reasons listed above, it is
necessary to elaborate NLP techniques for conceptual structures extraction using partial analysis of the patient record texts. We consider normalised texts,
with sucient punctuation marks and without spelling errors, because we aim
at research studies. Our present experimental corpus is formed by some 6400
words, with some 2000 of them being medical terms. Further discussion of the
text features is given in [11].
4 Patient Status Data Extraction
The Patient Records (PRs) in Bulgarian hospitals usually contain 2-3 pages. The
text is organised in the following sections: General information about the patient, Diagnosis, Anamnesis, Accompanying/Past diseases; Family anamnesis;
Risk factors, Status, Examinations and Clinical Data; Consultations; Debate.
Here we report results about extracting patient status data. The relevant PR
section contains mostly short declarative sentences in present tense which describe the status of different anatomic organs. Several organs are referred to in
the PRs of patients with diabetes. The Status section consists of descriptions of
the typical characteristics of 20-30 different anatomic organs and status conditions. The full description contains more than 45 different status observations.
The explanation detailness depends on the status of the corresponding anatomic
organs: for some organs only general characteristics are presented, while full
description is given for other organs which are important for the particular disease. For the missing organ description we assume that there is no deviation
from their normal status, therefore the system automatically includes certain
default values. When an organ description is located in the text, its phrases and
sentences are analysed by a cascade of regular expressions.
The analysis is based on a terminological bank of medical terms, derived
from the International Classification of Diseases (ICD-10) in Bulgarian language. It contains 10970 terms, a partial taxonomy of body parts and a medical goods list. A lexicon of 30000 Bulgarian lexemes, which is part of a large
general-purpose lexical database with 70000 lexemes, provides the necessary
linguistic resource for morphological analysis.
There are four manners to present the patient organs and their features in
our corpus:
-- General - by giving some default value, e.g. (with unchanged characteristics typical for the
age), (with preserved characteristics),
(with normal characteristics). For filling
in the obligatory IE template fields in these cases, we need predefined
default values for the respective organs status. The medical experts in
the project have developed a scale of normal, bed, worse and worst conditions (Fig. 1). The scale for the different organs can vary depending on
the possible values. Some words from the PRs are chosen as representative for the corresponding status scale interpretation and the other text
expressions are automatically classified into these typical status grades
by using especially designed regular expressions for shallow parsing.
Extracting Patient-Related Description from Medical Records in Bulgarian
25
General statements happen relatively often, for instance the skin status
for 26% of the PRs in our corpus is represented by general explanations.
Fig. 1 illustrates the scales for skin and gives examples for words signaling the respective status.
Scale
0
-1
-2
-3
Hydration
normal
mild dehydration
moderate dehydration
Turgor
good / preserved
doughy
decreased
Elasticity
good / preserved
doughy
reduced
severe dehydration
poor
poor
Fig. 1. Status types for skin characteristics
---
By diagnosis - sometimes a diagnosis is given instead of organ description, e.g. (onychomicosis) or
(obesity first degree).
Explicit - in this case the PR text contains particular specific values.
The characteristic name might be missing since the attribute is sufficient: e.g. pale skin instead of skin with pale colour. The attributes
are described by a variety of expressions, e.g. for the volume of the
thyroid gland the value (normal) can be represented as
; ; (not enlarged; not palpate enlarged; not palpate). There are several characteristics (about 15%) that have numerical values like
150/110 mmHg, 110/90 mmHg.
(BP in lying position 150/110 mmHg, in standing position 110/90
mmHg). Some typical organ descriptions in the PR texts are given below as regular expressions. Let us denote the Anatomic Organs by AO,
their characteristics by Ch and their attribute features by F. Then the
explicit status-related expressions can be grouped into the following
categories:
AO [-] [with of F] Ch1,[with of F] Ch2, ...
AO [-] [with of F] Ch1. [with of F] Ch2. ...
AO1 and AO2 [-] Ch1, Ch2, Ch3, ...
AO1 Ch11, Ch12, ..., AO2 Ch12, Ch22,..., AO3 Ch13 ...
About 73% of all PRs in our training corpus contain organ descriptions
in this format, which excludes the application of deeper syntactic analysis at least to the text paragraphs concerning organ descriptions. The
more complicated phrases and sentences are analysed by rules for recognising the characteristics and their values scope. Some values are not
described for all patients. Fig. 2 presents figures about the percentage
of occurances of major skin characteristics in the PRs corpus. To tackle
the problem, at present we are exploring the issue of correlating values,
where missing information for some attributes can be filled in using
status data interdependencies. For example for skin, we notice that the
turgor and elasticity are usually discussed together and their values depend of skin hydration chacteristics. Collecting statistical observations
about attribute interdependencies would be helpful for the IE process
as a whole.
26
S. Boytcheva, E. Paskaleva, Iv. Nikolova, and G. Angelova
Hypodermic Fat Tissue
63.24%
41.18%
Elasticity
42.65%
Turgor
Hydration
13.24%
66.18%
Colour
0%
20%
40%
80%
60%
Fig. 2. Percentage of PRs with explicit statements about skin status in the training corpus
--
Partial - the PR text contains descriptions about organ parts, not for the
whole anatomic organ. For instance, the skin status can be expressed by
phrases like (diffuse redness of the
face). In this case we need to generate dynamically an IE template with
some additional fields (Fig. 3).
Skin
Colour:
light rose
Turgor:
preserved
Elasticity:
preserved
Hydration:
normal
Face:
diffuse
redness
Fig. 3. Template with specific fields about face skin
The evaluation is performed with the help of medical experts who judge
the successful extraction of status attributes. We use a corpus of 197 PRs as a
training set and another 43 PRs as a test set. The assessment is done by organs
since their descriptions in the text are most often separated and can be analysed
independently. The PRs without any description of the organ status are remove
from the evaluation figures (Fig. 4).
skin
thyroid gland
limbs
Correctly recognised
characteristics
94.82%
87.35%
84.62%
Correctly processed PRs
94.03%
94.64%
76.75%
Fig. 4. Percentage of correctly extracted status attributes
Extracting Patient-Related Description from Medical Records in Bulgarian
27
The cases of incorrect extraction are due to more complex sentence structures in the PR text which need to be processed by a syntactic analyser.
5 Conclusion and Future Work
The presented approach uses techniques for filling dynamicaly generated templates depending on the available information in the PR Status section. The
reported work in progress tackles only some PR fragments. Further we aim
at temporal and cause-effect relations extraction. The project objective is to
develop algorithms for discovering more complex relations and other dependencies that are not explicitly stated in the text, which is a target for the future
project stages.
Acknowledgements. This research work is a part of the project EVTIMA (Effective search of conceptual information with applications in medical informatics) which is funded by the Bulgarian National Science Fund by grant No DO
02-292/December 2008.
References
1.
Natural Language Processing in Medical Coding. White paper of Language and
Computing (www.landcglobal.com). April 2004.
2. Christopher, S., G. Khoo, Syin Chan, Yun Niu. Extracting Causal Knowledge from
a Medical Database Using Graphical Patterns. In: Proceedings of 38th Annual
Meeting of the ACL, Hong Kong, 2000.
3. Grishman, R. Information Extraction: Techniques and Challenges. In: Pazienza,
M.-T. (Ed.), Information Extraction (Int. Summer School SCIE-97), Springer,
1997.
4. Cunnigham, H., Information extraction - an user guide. Research Memo CS-99-07,
University of Sheffield, 1999 (http://www.dcs.shef.ac.uk/hamish/IE).
5. Harkema, H., A. Setzer, R. Gaizauskas, M. Hepple, R. Power, and J. Rogers. Mining and Modelling Temporal Clinical Data. In: Proceedings of the 4th UK e-Science
All Hands Meeting, Nottingham, UK, 2005.
6. Gaizauskas, R., M. Hepple, N. Davis, Y. Guo, H. Harkema, A. Roberts, and
I.Roberts. AMBIT: Acquiring Medical and Biological Information from Text. In:
S.J. Cox (ed.) Proc. 2nd UK e-Science All Hands Meeting, Nottingham, UK, 2003.
7. Damianos, L., J. Ponte, S. Wohlever, F. Reeder, D. Day, G. Wilson, and L.
Hirschman. MiTAP for Bio-Security: A Case Study. AI Magazine 2002, 23(4), pp.
13-29.
8. Savova, G., P. Ogren, P. Duffy, J. Buntrock and C. Chute. Mayo Clinic NLP System
for Patient Smoking Status Identification. J. of the Am. Medical Informatics Association, Vol. 15 No. 1 Jan/Feb 2008, pp. 25-28.
9. Boytcheva, S., A. Strupchanska, E. Paskaleva, and D. Tcharaktchiev, Some Aspects
of Negation Processing in Electronic Health Records. In: Proc. of Int. Workshop
Language and Speech Infrastructure for Information Access in the Balkan Countries, September 2005, Borovets, Bulgaria, pp. 1-8.
10. Boytcheva, S. and G. Angelova. Towards Extraction of Conceptual Structures from
Electronic Health Records. In: Rudolph, S., Dau, F. and S. Kuznetsov (Eds.) Conceptual Structures: Leveraging Semantic Technologies, Proc. of the 17th International Conference on Conceptual Structures ICCS-2009, Moscow, Springer, LNAI
Vol. 5662, July 2009, pp. 100-113.
28
S. Boytcheva, E. Paskaleva, Iv. Nikolova, and G. Angelova
11. Boytcheva, S., I. Nikolova and E. Paskaleva. Context Related Extraction of Conceptual Information from Electronic Health Records. In: Priss, U. and G. Angelova
(Eds.) Conceptual Structures for Extracting Natural Language Semantics, Proc. of
the Int. Workshop SENSE-09, Moscow, Published on CEUR-Workshop online proceedings http://CEUR-WS.org/Vol-476/, Aachen, Vol. 476, July 2009.
You might also like
- Cek SubmitDocument40 pagesCek SubmitMulya Nurmansyah ArdisasmitaNo ratings yet
- Basic Probability Theory For Bio Medical Engineers - JohnD. EnderleDocument136 pagesBasic Probability Theory For Bio Medical Engineers - JohnD. Enderlejustwoow100% (1)
- Dosen Pembimbing Yang TersediaDocument187 pagesDosen Pembimbing Yang TersediaMulya Nurmansyah ArdisasmitaNo ratings yet
- UmlDocument120 pagesUmlRudrani SarkarNo ratings yet
- Alamat SikdaDocument1 pageAlamat SikdaMulya Nurmansyah ArdisasmitaNo ratings yet
- 27 1Document213 pages27 1Mulya Nurmansyah ArdisasmitaNo ratings yet
- Advanced Probability Theory For Bio Medical Engineers - John D. EnderleDocument108 pagesAdvanced Probability Theory For Bio Medical Engineers - John D. Enderleatul626100% (2)
- UnixDocument1 pageUnixMulya Nurmansyah ArdisasmitaNo ratings yet
- I Jcs It 2014050540Document5 pagesI Jcs It 2014050540Mulya Nurmansyah ArdisasmitaNo ratings yet
- Schema Integration Between Object-Oriented Databases: Ching-Ming ChaoDocument8 pagesSchema Integration Between Object-Oriented Databases: Ching-Ming ChaoMulya Nurmansyah ArdisasmitaNo ratings yet
- Prototyping Tools and TechniquesDocument41 pagesPrototyping Tools and Techniquesblaze3No ratings yet
- Chart Suggestions - A Thought-StarterDocument1 pageChart Suggestions - A Thought-StarterDoc AllaínNo ratings yet
- Rescuing Data From Decaying and Moribund Clinical Information SystemsDocument6 pagesRescuing Data From Decaying and Moribund Clinical Information SystemsMulya Nurmansyah ArdisasmitaNo ratings yet
- Create DFDs for Lemonade StandDocument20 pagesCreate DFDs for Lemonade Standsharoo501No ratings yet
- Ijcsi 9 5 1 329 338Document10 pagesIjcsi 9 5 1 329 338Mulya Nurmansyah ArdisasmitaNo ratings yet
- Professionalism in Interactions With Other Health ProfessionalsDocument16 pagesProfessionalism in Interactions With Other Health ProfessionalsMulya Nurmansyah ArdisasmitaNo ratings yet
- J Am Med Inform Assoc 2011 Borlawsky Amiajnl 2011 000354Document5 pagesJ Am Med Inform Assoc 2011 Borlawsky Amiajnl 2011 000354Mulya Nurmansyah ArdisasmitaNo ratings yet
- PRISM DescriptionOfToolsDocument38 pagesPRISM DescriptionOfToolsMulya Nurmansyah ArdisasmitaNo ratings yet
- Film Cari Versi RendahDocument1 pageFilm Cari Versi RendahMulya Nurmansyah ArdisasmitaNo ratings yet
- SA HB 137-2013 E Health Interoperability FrameworkDocument72 pagesSA HB 137-2013 E Health Interoperability FrameworkMulya Nurmansyah ArdisasmitaNo ratings yet
- Ehealth in IndonesiaDocument24 pagesEhealth in IndonesiaOemmaemNo ratings yet
- Denah Dasar1Document1 pageDenah Dasar1Mulya Nurmansyah ArdisasmitaNo ratings yet
- ,17 (5 29 (510 (17$/1 ( 27,$7,1 %2' $) &7&,1%,1) '2& 217+ (:+2) 5$0 (:25.&219 (17,21 1ryhpehu 2172%$&&2&21752/ 7klugvhvvlrq 3urylvlrqdodjhqgdlwhpDocument47 pages,17 (5 29 (510 (17$/1 ( 27,$7,1 %2' $) &7&,1%,1) '2& 217+ (:+2) 5$0 (:25.&219 (17,21 1ryhpehu 2172%$&&2&21752/ 7klugvhvvlrq 3urylvlrqdodjhqgdlwhpMulya Nurmansyah ArdisasmitaNo ratings yet
- Patients' Rights in Medical CareDocument17 pagesPatients' Rights in Medical CareMulya Nurmansyah ArdisasmitaNo ratings yet
- Asap CobaDocument1 pageAsap CobaMulya Nurmansyah ArdisasmitaNo ratings yet
- Modeling His 1Document8 pagesModeling His 1Mulya Nurmansyah ArdisasmitaNo ratings yet
- ,17 (5 29 (510 (17$/1 ( 27,$7,1 %2' $) &7&,1%,1) '2& 217+ (:+2) 5$0 (:25.&219 (17,21 1ryhpehu 2172%$&&2&21752/ 7klugvhvvlrq 3urylvlrqdodjhqgdlwhpDocument47 pages,17 (5 29 (510 (17$/1 ( 27,$7,1 %2' $) &7&,1%,1) '2& 217+ (:+2) 5$0 (:25.&219 (17,21 1ryhpehu 2172%$&&2&21752/ 7klugvhvvlrq 3urylvlrqdodjhqgdlwhpMulya Nurmansyah ArdisasmitaNo ratings yet
- File FTPDocument1 pageFile FTPMulya Nurmansyah ArdisasmitaNo ratings yet
- Trollandtoad Heroclix Figures and ComicsDocument8 pagesTrollandtoad Heroclix Figures and ComicsMulya Nurmansyah ArdisasmitaNo ratings yet
- Shoe Dog: A Memoir by the Creator of NikeFrom EverandShoe Dog: A Memoir by the Creator of NikeRating: 4.5 out of 5 stars4.5/5 (537)
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeFrom EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeRating: 4 out of 5 stars4/5 (5794)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceFrom EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceRating: 4 out of 5 stars4/5 (890)
- The Yellow House: A Memoir (2019 National Book Award Winner)From EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Rating: 4 out of 5 stars4/5 (98)
- The Little Book of Hygge: Danish Secrets to Happy LivingFrom EverandThe Little Book of Hygge: Danish Secrets to Happy LivingRating: 3.5 out of 5 stars3.5/5 (399)
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryFrom EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryRating: 3.5 out of 5 stars3.5/5 (231)
- Never Split the Difference: Negotiating As If Your Life Depended On ItFrom EverandNever Split the Difference: Negotiating As If Your Life Depended On ItRating: 4.5 out of 5 stars4.5/5 (838)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureFrom EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureRating: 4.5 out of 5 stars4.5/5 (474)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersFrom EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersRating: 4.5 out of 5 stars4.5/5 (344)
- Grit: The Power of Passion and PerseveranceFrom EverandGrit: The Power of Passion and PerseveranceRating: 4 out of 5 stars4/5 (587)
- On Fire: The (Burning) Case for a Green New DealFrom EverandOn Fire: The (Burning) Case for a Green New DealRating: 4 out of 5 stars4/5 (73)
- The Emperor of All Maladies: A Biography of CancerFrom EverandThe Emperor of All Maladies: A Biography of CancerRating: 4.5 out of 5 stars4.5/5 (271)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaFrom EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaRating: 4.5 out of 5 stars4.5/5 (265)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreFrom EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreRating: 4 out of 5 stars4/5 (1090)
- Team of Rivals: The Political Genius of Abraham LincolnFrom EverandTeam of Rivals: The Political Genius of Abraham LincolnRating: 4.5 out of 5 stars4.5/5 (234)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyFrom EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyRating: 3.5 out of 5 stars3.5/5 (2219)
- The Unwinding: An Inner History of the New AmericaFrom EverandThe Unwinding: An Inner History of the New AmericaRating: 4 out of 5 stars4/5 (45)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)From EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Rating: 4.5 out of 5 stars4.5/5 (119)
- Her Body and Other Parties: StoriesFrom EverandHer Body and Other Parties: StoriesRating: 4 out of 5 stars4/5 (821)
- P.3 Comprehension SchemeDocument4 pagesP.3 Comprehension Schemeice200mNo ratings yet
- Out of The Classroom and Into The City The Use ofDocument13 pagesOut of The Classroom and Into The City The Use ofLuciaNo ratings yet
- Trauma On Brain DevelopmentDocument29 pagesTrauma On Brain DevelopmentChirra Williams100% (1)
- Executive Forum: Leading in a VUCA WorldDocument6 pagesExecutive Forum: Leading in a VUCA WorldZulkarnainNo ratings yet
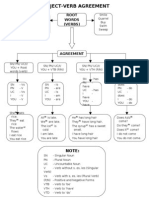
- Subject-Verb Agreement RulesDocument1 pageSubject-Verb Agreement RulesMohd AsyrafNo ratings yet
- Dement and Kleitman Short QuestionsDocument12 pagesDement and Kleitman Short QuestionsUsamaNo ratings yet
- Embodied Affectivity: On Moving and Being Moved: Thomas Fuchs and Sabine C. KochDocument12 pagesEmbodied Affectivity: On Moving and Being Moved: Thomas Fuchs and Sabine C. Kochma ceciNo ratings yet
- Interpretative Sociology and Axiological Neutrality in Max Weber and Werner SombartDocument17 pagesInterpretative Sociology and Axiological Neutrality in Max Weber and Werner Sombartionelam8679No ratings yet
- German Learning Course MaterialDocument66 pagesGerman Learning Course MaterialRobin100% (1)
- Module 25 Introduction To StylisticsDocument31 pagesModule 25 Introduction To Stylisticsshella mae pormento100% (2)
- Module 4-Section 1 GLOBAL CULTURE AND MEDIADocument7 pagesModule 4-Section 1 GLOBAL CULTURE AND MEDIAGIL BARRY ORDONEZNo ratings yet
- Multipurpose Multiplication TablesDocument54 pagesMultipurpose Multiplication Tablesnicksonwong89No ratings yet
- Silviana Rahayu Hersa - 8.3 - Remedial Pas PDFDocument14 pagesSilviana Rahayu Hersa - 8.3 - Remedial Pas PDFDesfi Nurrachma hersaNo ratings yet
- Catch Up Friday Gr10 Mar1Document3 pagesCatch Up Friday Gr10 Mar1milafer dabanNo ratings yet
- Conditionals 2nd BachilleratoDocument2 pagesConditionals 2nd BachilleratoFatima VazquezNo ratings yet
- What New Information Did I Learn From This Activity? DiscussDocument2 pagesWhat New Information Did I Learn From This Activity? DiscussJulius CabinganNo ratings yet
- A Description of Colloquial GuaranDocument125 pagesA Description of Colloquial GuaranBerat GültürkNo ratings yet
- Emotional Intelligence Participant Manual CoulibalyDocument20 pagesEmotional Intelligence Participant Manual Coulibalybrocklyn311No ratings yet
- Cause & Effect: English LessonDocument11 pagesCause & Effect: English LessonAditya DelfiantoNo ratings yet
- DLL - MTB 3 - Q2 - Catch Up ActivityDocument3 pagesDLL - MTB 3 - Q2 - Catch Up ActivityHazel Israel BacnatNo ratings yet
- IntroductionDocument8 pagesIntroductionturdygazyyeva.aigerimNo ratings yet
- Business Communication NotesDocument14 pagesBusiness Communication NotesAnimesh MayankNo ratings yet
- Pedagogical Grammar Dr. Lazgeen Mohammed Raouf Aims, Objectives and OutcomesDocument22 pagesPedagogical Grammar Dr. Lazgeen Mohammed Raouf Aims, Objectives and OutcomesMohammed DoskiNo ratings yet
- Deep Learning: Data Mining: Advanced AspectsDocument131 pagesDeep Learning: Data Mining: Advanced AspectsmakeosNo ratings yet
- Syllabus in Assessment and Evaluation in MathematicsDocument7 pagesSyllabus in Assessment and Evaluation in MathematicsJhielaMaeMacaraigNo ratings yet
- Kenneth E. Bruscia-Defining Music Therapy-Barcelona Pub (2014)Document422 pagesKenneth E. Bruscia-Defining Music Therapy-Barcelona Pub (2014)Cristina Maria Oltean100% (14)
- Ipn-Upiicsa Teacher's Training Course 1 Outline Centro de Idiomas PropedeuticsDocument2 pagesIpn-Upiicsa Teacher's Training Course 1 Outline Centro de Idiomas PropedeuticsLillian LöweNo ratings yet
- 001-Supervision Essentials For Emotion-Focused TherapyDocument181 pages001-Supervision Essentials For Emotion-Focused TherapyJenny Jin100% (1)
- TRỌNG ÂM TỪ 3 ÂM TIẾT 12Document2 pagesTRỌNG ÂM TỪ 3 ÂM TIẾT 12tuannguyen7422No ratings yet
- Fs 5 - Episode 1Document3 pagesFs 5 - Episode 1api-33461747095% (20)