Professional Documents
Culture Documents
Am002683 PDF
Uploaded by
pedro0723Original Description:
Original Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
Am002683 PDF
Uploaded by
pedro0723Copyright:
Available Formats
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, June 2001, p.
26832691
0099-2240/01/$04.000 DOI: 10.1128/AEM.67.6.26832691.2001
Copyright 2001, American Society for Microbiology. All Rights Reserved.
Vol. 67, No. 6
Isolation and Characterization of Polycyclic Aromatic
Hydrocarbon-Degrading Bacteria Associated with
the Rhizosphere of Salt Marsh Plants
L. L. DAANE,1 I. HARJONO,1 G. J. ZYLSTRA,1,2
AND
GGBLOM1,2*
M. M. HA
Biotechnology Center for Agriculture and the Environment1 and Department of Biochemistry and
Microbiology,2 Cook College, Rutgers University, New Brunswick, New Jersey 08901
Received 25 September 2000/Accepted 12 March 2001
Polycyclic aromatic hydrocarbon (PAH)-degrading bacteria were isolated from contaminated estuarine
sediment and salt marsh rhizosphere by enrichment using either naphthalene, phenanthrene, or biphenyl as
the sole source of carbon and energy. Pasteurization of samples prior to enrichment resulted in isolation of
gram-positive, spore-forming bacteria. The isolates were characterized using a variety of phenotypic, morphologic, and molecular properties. Identification of the isolates based on their fatty acid profiles and partial 16S
rRNA gene sequences assigned them to three main bacterial groups: gram-negative pseudomonads; grampositive, non-spore-forming nocardioforms; and the gram-positive, spore-forming group, Paenibacillus.
Genomic digest patterns of all isolates were used to determine unique isolates, and representatives from each
bacterial group were chosen for further investigation. Southern hybridization was performed using genes for
PAH degradation from Pseudomonas putida NCIB 9816-4, Comamonas testosteroni GZ42, Sphingomonas
yanoikuyae B1, and Mycobacterium sp. strain PY01. None of the isolates from the three groups showed homology
to the B1 genes, only two nocardioform isolates showed homology to the PY01 genes, and only members of the
pseudomonad group showed homology to the NCIB 9816-4 or GZ42 probes. The Paenibacillus isolates showed
no homology to any of the tested gene probes, indicating the possibility of novel genes for PAH degradation.
Pure culture substrate utilization experiments using several selected isolates from each of the three groups
showed that the phenanthrene-enriched isolates are able to utilize a greater number of PAHs than are the
naphthalene-enriched isolates. Inoculating two of the gram-positive isolates to a marine sediment slurry spiked
with a mixture of PAHs (naphthalene, fluorene, phenanthrene, and pyrene) and biphenyl resulted in rapid
transformation of pyrene, in addition to the two- and three-ringed PAHs and biphenyl. This study indicates
that the rhizosphere of salt marsh plants contains a diverse population of PAH-degrading bacteria, and the use
of plant-associated microorganisms has the potential for bioremediation of contaminated sediments.
significant cause of PAH removal (6). Numerous studies have
been conducted on microbial consortia and enrichment, and
several diverse genera of bacteria have been isolated (for a
review, see reference 6). Recent work has indicated that the
stimulation of microbial activity in the rhizosphere of plants
can also stimulate biodegradation of various toxic organic compounds (1, 4, 37, 39, 41, 45, 47). This general rhizosphere
effect is well known in terrestrial systems. The rhizosphere
soil has been described as the zone of soil under the direct
influence of plant roots and usually extends a few millimeters
from the root surface and is a dynamic environment for microorganisms (7). The rhizosphere microbial community is
comprised of microorganisms with different types of metabolism and adaptive responses to variation in environmental conditions. The production of mucilaginous material and the exudation of a variety of soluble organic compounds by the plant
root play an important part in root colonization and maintenance of microbial growth in the rhizosphere. Microbial activity is thus generally higher in the rhizosphere due to readily
biodegradable substrates exuded from the plant (35). Recent
studies indicate that degradation of PAHs and polychlorinated
biphenyls (PCBs) in soils can be enhanced by plants (37, 47). In
the case of PCBs, plant compounds have been shown to induce
the cometabolic degradation of PCBs (15, 16).
The activities of microbial communities in sediments and
tidal marshes are important in improving sediment quality
Contamination of estuarine sediments by highly hydrophobic, low-availability, toxic, organic contaminants has become a
significant environmental concern. For example, industrial activities in the New Jersey/New York Harbor estuary have resulted in widespread contamination, and the management of
contaminated sediments has become a significant problem.
Polycyclic aromatic hydrocarbons (PAHs) are of particular
concern because of their toxic, mutagenic, and carcinogenic
properties (32). PAHs have also been found to bioaccumulate
in aquatic organisms (29). Although PAHs are persistent,
mainly due to their high hydrophobicity, a variety of microorganisms (bacteria and fungi) can degrade certain PAHs (for
reviews, see references 6 and 44). There is thus a significant
interest in studying microorganisms present in contaminated
environments as a means for bioremediation.
The fate of PAHs and other organic contaminants in the
environment is associated with both abiotic and biotic processes, including volatilization, photooxidation, chemical oxidation, bioaccumulation, and microbial transformation. Microbial activity has been deemed the most influential and
* Corresponding author. Mailing address: Department of Biochemistry and Microbiology, Cook College, Rutgers University, 76 Lipman
Dr., New Brunswick, NJ 08901-8525. Phone: (732) 932-9763, ext. 326.
Fax: (732) 932-8965. E-mail: haggblom@aesop.rutgers.edu.
Present address: Avon Products, Inc., Suffern, NY 10901-5605.
2683
2684
DAANE ET AL.
(pollution degradation), in the biological transformations of
major nutrients influencing their availability for plant growth,
and in their role in binding sediment particles to improve
rooting medium structure. Wetlands have a higher biological
activity than many other ecosystems, and they support enhanced biotransformation of toxic chemicals (25). Experiments
with Spartina salt marsh mesocosms indicated that biodegradation of oil was influenced by flooding and fertilization conditions (28, 49).
There is extensive information on PAH degradation by both
gram-positive and gram-negative bacteria, mostly isolated
from soil environments. There is, however, little information
on the microbial communities in salt marsh ecosystems and
their role in the biodegradation of toxic organic contaminants.
In addition, little is known about the ability of these microorganisms to degrade polyaromatic compounds. In the present
study, we isolated PAH-degrading bacteria from the salt marsh
rhizosphere. We were particularly interested in isolating sporeforming bacteria for two reasons. First, the ability of a bacterium to produce spores and therefore survive adverse conditions would be an attractive feature for remediation of
contaminated sites. Secondly, only a few studies have examined
the degradation of PAHs by spore-forming bacteria, and none,
to our knowledge, involve plant-microbe interactions. Therefore, isolation of spore-forming, PAH-degrading bacteria and
subsequent genetic studies of their degradation pathways may
lead to the discovery of novel genes involved. In addition, we
wanted to evaluate the biodegradation potential of select isolates and determine if reinoculation of the rhizosphere would
enhance degradation of contaminated marine sediment.
MATERIALS AND METHODS
Plant and sediment samples. Salt marsh plant samples were obtained at two
sites. The first site, considered to be uncontaminated, was the University of
Delaware Marine Station at Lewes, Del. A total of five different plant samples
were collected and included Distichlis spicata, Juncus gerardi, Phragmites australis,
Spartina alterniflora, and Sporobolus airoides. A minimum of two plants of each
species was obtained. The second site was Piles Creek, a contaminated tributary
of Arthur Kill, in Linden, N.J. Estuarine sediment and S. alterniflora plant
samples were obtained from the intertidal area. A total of three sediment and
three plant samples were taken from separate areas within 50 feet of each other.
The plant and sediment samples were transported on ice back to the laboratory
where they were stored at 4C until analyzed.
Contaminated sediment for the microbial-sediment slurry biotransformation
experiment was obtained from Newtown Creek in the New York Harbor, Brooklyn, N.Y. Material was collected by the Army Corps of Engineers, and a chemical
analysis showed the dredged sediment to contain 2 to 7 ppm of the PAHs
naphthalene, anthracene, and phenanthrene. The sediment texture consisted of
50% sand, 41% silt, and 9% clay, as determined by the Soil Testing Laboratory
at the New Jersey Agricultural Experiment Station. The structure is considered
a silt loam, organic matter was 3.7%, and the pH was 7.90.
Bacterial strains, plasmids and media. Pseudomonas putida NCIB 9816-4 (9),
Comamonas testosteroni GZ42 (18), Sphingomonas yanoikuyae B1 (14), and Mycobacterium sp. strain PY01 (Y.-S. Oh and G. J. Zylstra, unpublished data) were
chosen as representatives of different genotypes of PAH-degrading organisms.
The genes have previously been cloned from these strains into Escherichia coli,
and the cloned fragments were used as gene probes for Southern hybridization
experiments. Plasmid pDTG112 contains the cloned nah genes from P. putida
NCIB 9816-4 (40). A 3.6-kb SalI fragment contains three of the four genes for
naphthalene dioxygenase: nahAa (reductase), nahAb (ferredoxin), and nahAc
(dioxygenase large subunit) (42). Plasmid pGJZ1822, cloned from C. testosteroni
GZ42, has a 6.7-kb SstI-XhoI fragment that contains the genes necessary for the
first few steps in PAH degradation (19). Plasmid pGJZ1512, cloned from S.
yanoikuyae B1, has a 567-bp SphI fragment that contains bphC encoding a
meta-ring cleavage enzyme involved in PAH degradation (26). A 600-bp fragment containing a gene encoding a dioxygenase large subunit involved in PAH
APPL. ENVIRON. MICROBIOL.
degradation by Mycobacterium sp. strain PY01 (J. F. Cigolini and G. J. Zylstra,
Abstr. 99th Gen. Meet. Am. Soc. Microbiol., abstr. Q-180, 1999) was PCR
amplified with the primers 5-GTTGACCCGTGACGC-3 and 5-CTCACTCA
AGGCCGG-3. The PAH-degrading strain C. testosteroni GZ39 was used as a
control in hybridization experiments (18).
Mineral salts basal (MSB) medium (43) was used for enrichment cultures and
isolation. Solid minimal medium contained 2% noble agar (Difco Laboratories,
Detroit, Mich.). Stock concentrations (100 mg ml1) of naphthalene, biphenyl,
and phenanthrene were dissolved in dimethyl formamide and were added to
liquid medium at a final concentration of 1 mg ml1. Naphthalene and biphenyl
were added in the vapor phase as crystals in the petri dish lid for solid medium.
Phenanthrene was added as a 2% noble agar overlayer onto MSB agar plates at
a final concentration of 1 mg ml1. All PAHs were obtained from Aldrich
Chemical Co. (Milwaukee, Wis.) and were a minimum of 98% pure.
Bacterial strains were grown on Trypticase soy (Beckton Dickinson & Co.,
Cockeysville, Md.) agar (TSA) plates for various phenotypic and fatty acid
methyl ester (FAME) analyses. For plasmid preparations, bacterial strains were
grown in Luria-Bertani medium (38) containing either kanamycin (50 g ml1;
Sigma Chemical Co., St. Louis, Mo.) or ampicillin (100 g ml1; Sigma Chemical
Co.) added from filter-sterilized stock solutions. All bacterial strains and isolates
were routinely grown aerobically at 30C, except for E. coli strains, which were
grown at 37C. Strains were maintained on minimal medium at 4C, and longterm storage was in 50% glycerol at 80C.
Enrichment and isolation of PAH-degrading bacteria. Bacterial enrichment
cultures were set up in cotton-plugged Erlenmeyer flasks containing 50 ml of
MSB medium using naphthalene, phenanthrene, or biphenyl as the sole source
of carbon. Plant rhizosphere and contaminated estuarine sediment were used as
sources of the inoculum for the enrichment cultures. Rhizosphere samples were
obtained by splitting open the plant root mass, so that the inner roots that had
not come in contact with sampling or storage devices could be aseptically cut and
collected. One gram of root material was gently shaken to remove loose sediment
material and was added to each enrichment culture. Sediment samples were
added directly to each enrichment culture using a 1-g subsample. A total of two
enrichment cultures for each substrate were set up with each rhizosphere and
sediment sample. One set of enrichment cultures was placed directly in a 28C
environmental chamber and shaken at 150 rpm. The second set of enrichments
was heat treated for 10 min at 80C before incubation to enhance isolation of
spore-forming bacteria.
The cultures were monitored for the presence of microorganisms by microscopy and gram staining and were subcultured (1:50) into fresh MSB medium
containing the selected PAH when growth was detected. After three to four
subcultures, the cultures were plated onto MSB agar plates containing the same
substrate as the enrichment. Piles Creek sediment and S. alterniflora rhizosphere
isolates are denoted as PS and PR, respectively. Delaware isolates obtained from
the rhizosphere of S. airoides, S. alterniflora, D. spicata, J. gerardi, or P. australis
are denoted as SA, Salt, DS, JG, or PA, respectively. In addition, strains isolated
using naphthalene are indicated by an N, while phenanthrene- and biphenylenriched isolates are denoted by P and B, respectively.
Phenotypic characterizations. Phase-contrast microscopy (1,000 magnification) was used for the morphologic examination of bacterial cells. The strains to
be tested were inoculated into 2 ml of TS broth and incubated on a rotary wheel
at 37C. The cells were examined microscopically for motility and morphology at
24, 48, and 72 h and were examined again microscopically after Gram staining.
FAME analysis. Whole-cell fatty acid analyses were performed on all of the
PAH-degrading isolates by growing the cells at 28C for 24 h on TSA plates.
Cellular fatty acids were saponified, methylated, extracted, and analyzed by gas
chromatography following the procedures given for the Sherlock Microbial Identification System (MIDI, Inc., Newark, Del.). Identification and comparison were
made to the Aerobe (TSBA version 3.9) database of the Sherlock Microbial
Identification System. The Dendrogram program of the MIDI software package
was used to produce unweighted-pair matchings based on fatty acid compositions.
DNA preparation. Bacterial strains were grown on either TSA or MSB agar
plates supplemented with naphthalene, phenanthrene, or biphenyl. Total
genomic DNA was extracted as described by Wilson (48), with the exception of
pretreatment with 1.5 mg of lysozyme for 2 h. With the exception of plasmid
pDTG112, plasmid DNA was obtained using the QIAprep Spin Miniprep Kit
(Qiagen Inc., Santa Clarita, Calif.) according to the manufacturers directions.
For plasmid pDTG112, the plasmid DNA was purified from the total DNA using
equilibrium centrifugation in a CsCl-ethidium bromide gradient as described by
Sambrook et al. (38). Restriction enzyme digests were performed on the isolated
plasmids, and the desired fragments were purified using the Qiaquick gel extraction kit (Qiagen).
VOL. 67, 2001
PAH-DEGRADING BACTERIA FROM SALT MARSH RHIZOSPHERE
Restriction fragment length polymorphism (RFLP). Total genomic restriction
enzyme digests were performed on all of the PAH-degrading isolates using either
the BamHI, EcoRI, HindIII, or NotI enzyme as recommended by the supplier
(Gibco-BRL, Madison, Wis.). Approximately 2 to 3 g of each isolate was
digested. Electrophoretic analysis of the digested DNA was done on a 0.8%
agarose gel at 25 V for 16 to 20 h. The restriction enzyme digest patterns were
visualized by UV light following ethidium bromide staining.
16S rDNA analysis. Sequence analyses of 16S ribosomal DNA (rDNA) were
performed on several of the isolates by amplifying the 16S rRNA genes by PCR
using the eubacterial primers 27f and 1522r (24). The PCR mixtures (100 l)
contained 10 mM Tris-HCl (pH 8.3), 50 mM KCl, 1.5 mM MgCl2, a 200 M
concentration of each deoxynucleoside triphosphate, 50 pmol of each primer, 2.5
U of Taq DNA polymerase (Gibco-BRL, Grand Island, N.Y.), and 10 ng of
DNA template. PCR amplification was performed using a Perkin-Elmer GeneAmp PCR System 2400 thermocycler (Applied Biosystems, Foster City, Calif.)
programmed as follows: 1 min of denaturation at 94C; followed by 25 cycles of
96C for 1 min, 55C for 1 min, and 72C for 1 min; with a final extension at 72C
for 10 min. The amplified 1.5-kb PCR products were excised from a 1% agarose
gel and purified using the QIAquick gel extraction kit (Qiagen) according to the
manufacturers instructions.
Partial DNA sequences using the primers 27f and 685r (24) were determined
directly from the purified PCR products with automated fluorescent Taq cycle
sequencing using the ABI 373A Sequencer (Applied Biosystems).
Partial 16S rRNA sequences (ranging from 301 to 1,500 bp) of each isolate
were analyzed using the Ribosomal Database Project Sequence Match and
Similarity Matrix programs (31) to obtain the most closely matched species.
Southern hybridization using PAH gene probes. Southern blots were performed on all unique isolates, as determined by RFLP, using gene probes for
four different types of PAH dioxygenases. This was accomplished by digesting 2
to 3 g of genomic DNA from each of the tested isolates with EcoRI according
to the manufacturers directions. In addition, genomic DNA obtained from P.
putida NCIB 9816-4, C. testosteroni GZ42, C. testosteroni GZ39, S. yanoikuyae B1,
and Mycobacterium sp. strain PY01 was digested and used as positive and negative controls for each hybridization. The digests were run on 0.8% agarose gels
in 40 mM Tris20 mM acetate2 mM EDTA buffer at 25 V for 16 to 20 h, stained
with ethidium bromide, and visualized under UV light prior to transferring onto
positively charged nylon membranes (Boehringer Mannheim, Indianapolis, Ind.)
using a VacuGene XL Vacuum Blotting System (Pharmacia Biotech, Piscataway,
N.J.) The transferred DNA was fixed to the nylon membranes by exposure to a
UV cross-linker (Spectronics Corp., Westbury, N.Y.) using a 2 optimal crosslink (120 mJ cm2). The Southern hybridization protocol was performed using
nonradioactive digoxigenin (Boehringer Mannheim) as recommended by the
supplier. Gene probes were prepared from the PAH dioxygenase gene fragments
by a nonradioactive random primed DNA-labeling method with alkali-labile
digoxigenin 11dUTP. Prehybridization and hybridization were performed at
37C. Following hybridization, the nylon membrane was washed under lowstringency conditions (0.1 SSC [1 SSC is 0.15 M NaCl plus 0.015 M sodium
citrate] and 0.1% sodium dodecyl sulfate) at room temperature.
Pure culture transformation of PAHs in liquid medium. Several of the isolates
were analyzed for their ability to utilize a variety of aromatic compounds, including naphthalene, phenanthrene, fluorene, pyrene, and biphenyl, individually
and as a mixture, in MSB medium. The isolates tested included the gramnegative pseudomonads PR-N7, PR-N9, PR-N10, and PS-P2; the gram-positive
non-spore formers PR-N15 and PR-P3; and the spore former PR-P1. Each
isolate was tested in duplicate for each substrate or substrates over a time course
of 11 days. Cells were grown in 100 ml of MSB containing 5 mM succinate,
pelleted by centrifugation (4,000 g, 10 min), washed with MSB, pelleted again,
and suspended in fresh MSB to give a cell density of approximately 107 cells ml1
for use in transformation experiments. Sterile 7-ml serum vials containing the
appropriate carbon source at a final concentration of 100 M were added from
hexane stock solutions, and the material was allowed to evaporate prior to the
addition of 2 ml of washed cell suspension. In addition, two controls were added
to the experimental design and included a no-cell control (MSB medium only)
and a dead-cell control (MSB 4% formaldehyde cells). The samples were
incubated at 28C on a rotary shaker at 90 rpm, and the amount of substrate
remaining was analyzed over time. At each time point two vials were taken, were
extracted with 1 ml of hexane, and were analyzed for the appropriate aromatic
substrate using a Hewlett-Packard 5890 Series II Gas Chromatograph using an
HP-5MS column and equipped with a 5971 series mass selective detector. Prior
to extraction, 20 M 2-methylnaphthalene was added from a hexane stock
solution as an internal standard. Results are reported as the percentage remaining relative to uninoculated controls after 11 days of incubation.
2685
Microbial-sediment slurry biotransformation experiment. An aerobic slurry
system was designed to determine the rate and extent of microbial transformation of contaminated sediment. The rhizosphere-associated phenanthrene-degrading strains PR-P1 and PR-P3 were grown to mid-log phase in TS broth and
were harvested by centrifugation at 4,000 g for 10 min. The cells were washed
and pelleted by centrifugation twice using 1/10 salts (8) as the diluent, diluted
into 1/10 salts, and inoculated at three different cell concentrations into a PAHspiked sediment slurry.
Approximately equal volumes (500 ml) of estuarine sediment material, obtained from Newtown Creek, and water were mixed under forced aeration for 1
week prior to bacterial inoculation of slurry microcosms. The microcosms consisted of 50-ml glass serum bottles containing 5 ml of aerated slurry spiked with
a mixture of PAHs (naphthalene, fluorene, phenanthrene, and pyrene) and
biphenyl from an acetone stock at a concentration of approximately 50 ppm of
each. Each bacterial isolate was added to the spiked slurry at 3 inoculum densities, 105, 107, or 109 cells ml1. Two controls were used consisting of an
uninoculated nonsterile slurry control and a sterile (autoclaved three times over
3 days) slurry control. All the bottles were sealed with Teflon stoppers pierced
with an 18-gauge cotton-plugged needle, and the cultures were incubated in a
30C environmental chamber at 90 rpm. The loss of PAHs and biphenyl was
analyzed over time.
Entire microcosms were sacrificed in triplicate and extracted for PAH analyses
at nine time points over a 56-day period (7-day sampling interval). Twenty parts
per million of 2-methylnaphthalene was added as an internal standard, and the
content of the entire vial was extracted overnight with 4 ml of hexane and 2 ml
of acetone by shaking on a wrist-action shaker at 200 rpm. The samples were
centrifuged for 5 min at 300 g, and the organic phase was removed. The slurry
was further extracted using 2 ml of hexane for 4 h by shaking at 200 rpm. The
organic phases were combined and analyzed for PAH and biphenyl content by
gas chromatography, as described above. Statistical analyses were performed
using the Minitab Software Package Release 11.21.
Nucleotide sequence accession numbers. The 16S rRNA sequences for the
PAH-degrading isolates have been deposited in GenBank under accession numbers AF353676 to AF353705.
RESULTS
Isolation of PAH-degrading bacterial strains. The PAHdegrading strains isolated in this study are listed in Table 1. A
total of 42 isolates were obtained from S. alterniflora plant and
sediment samples collected from contaminated Piles Creek in
New Jersey, using either naphthalene, phenanthrene, or biphenyl as the sole source of carbon. A total of nine isolates were
obtained from the four different plant rhizospheres collected
from Lewes, Del. However, no PAH-degrading isolates were
obtained from the rhizosphere of P. australis from this site. The
lower number and diversity of bacteria isolated from this site
may be expected due to its uncontaminated nature. Heat treatment prior to enrichment and isolation was included for each
of the substrates tested, which resulted in isolation of several
spore-forming bacteria. The isolates were characterized on the
basis of their morphologic and phenotypic properties and by
using a combination of molecular techniques.
FAME analysis. The tentative identification and similarity
index (SI) of all of the isolates using the aerobic (TSBA)
library version 3.9 of the Microbial Identification System is
shown in Table 1. A pairwise-comparison Euclidean-distance
dendrogram based on whole-cell fatty acid compositions of the
PAH-degrading isolates (except isolate PS-P1) was constructed, revealing that the isolates fall within three main
groups (Fig. 1). Group 1 included the endospore-forming
Paenibacillus, group 2 included the gram-positive nocardioforms, and group 3 included the gram-negative pseudomonads.
One of the isolates, PR-P12, falling outside these groups was
tentatively identified as Flavobacterium resinovorum (SI
0.79) (Table 1). Isolate PR-P3 clustered within the Paenibacil-
2686
DAANE ET AL.
APPL. ENVIRON. MICROBIOL.
TABLE 1. PAH-degrading isolates obtained from contaminated estuarine sediment and salt marsh plant rhizosphere and comparison of
bacterial identification using FAME and partial 16S rRNA gene sequences of several of the strains
Isolatea
Source
PAHb
DS-N1
DS-N2
DS-N3
JG-N1
JG-N2
JG-N3
SA-N1
Salt-N1
Salt-N2
PR-N1
PR-N2
PR-N3
PR-N4
PR-N5
PR-N6
PR-N7
PR-N8
PR-N9
PR-N10
PR-N11
PR-N12
PR-N13
PR-N14
PR-N15
PR-N16
PR-N17
PR-N18
PR-N19
PR-N20
PR-N21
PR-N22
PR-P1
PR-P2
PR-P3
PR-P6
PR-P9
PR-P10
PR-P11
PR-P12
PR-P13
PR-B1
PR-B2
PS-N1
PS-N2
PS-N3
PS-N4
PS-N5
PS-N6
PS-P1
PS-P2a
PS-P2b
D. spicata
D. spicata
D. spicata
J. gerardi
J. gerardi
J. gerardi
S. airoides
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
S. alterniflora
Sediment
Sediment
Sediment
Sediment
Sediment
Sediment
Sediment
Sediment
Sediment
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
N
P
P
P
P
P
P
P
P
P
B
B
N
N
N
N
N
N
P
P
P
FAME analysisc
Partial 16S rRNA gene sequenced
Best match
SI
Best match
% Similarity
Brevibacillus brevis
P. validuse
P. validus
P. validus
P. validus
P. validus
Paenibacillus sp.
Pseudomonas fluorescens
P. putida
Paenibacillus sp.
Paenibacillus sp.
P. validus
P. validus
Paenibacillus sp.
Pseudomonas chlororaphis
P. stutzeri
P. putida
P. putida
P. putida
Paenibacillus sp.
Paenibacillus sp.
P. validus
N. asteroides
N. brasiliensis
Paenibacillus sp.
N. asteroides
Nocardia sp.
P. validus
P. validus
P. validus
P. validus
P. validus
P. validus
A. oxydans
P. validus
P. validus
P. validus
P. validus
F. resinovorum
P. validus
P. validus
B. brevis
P. pabuli
P. fluorescens
N. asteroides
P. stutzeri
P. stutzeri
P. validus
Nocardioform
Alcaligenes xylosoxidans
A. xylosoxidans
0.45
0.92
0.81
0.80
0.81
0.87
Poor
0.59
0.80
Poor
Poor
0.87
0.86
Poor
0.85
0.88
0.87
0.87
0.82
Poor
Poor
0.81
0.77
0.21
Poor
0.57
0.14
0.93
0.84
0.75
0.81
0.85
0.90
0.46
0.88
0.91
0.91
0.70
0.79
0.91
0.87
0.34
0.26
0.47
0.71
0.89
0.86
0.33
NDg
0.65
0.56
P. validus
P. validus
P. validus
NDf
P. validus
ND
P. validus
ND
ND
P. validus
ND
ND
P. validus
P. validus
ND
P. stutzeri
ND
P. putida
P. putida
ND
ND
P. validus
Rhodococcus ruber
Tsukamurella wratislaviensis
P. validus
ND
ND
P. validus
P. validus
P. validus
P. validus
P. validus
ND
A. oxydans
ND
P. validus
P. validus
P. validus
S. subarctica
P. validus
ND
P. validus
P. validus
ND
ND
ND
ND
P. validus
ND
Alcaligenes faecalis
ND
99.6
96.7
97.9
96.0
96.0
97.3
98.0
96.9
99.0
99.0
98.0
98.2
97.1
99.8
97.3
97.3
98.0
98.9
98.5
98.5
95.4
99.5
99.8
99.5
99.3
97.9
98.4
97.1
96.4
98.4
DS, JG, SA, and salt isolates obtained from plants collected from Lewes, Del. PR and PS designates isolates obtained at Piles Creek, N.J.
PAH carbon source in enrichment culture. N, naphthalene; P, phenanthrene; and B, biphenyl.
FAME identification was performed using the aerobic (TSBA) library version 3.9 of the Microbial Identification System (MIDI, Inc.). SI values below 0.2 are listed
as poor, and the match is given only to the genus.
d
16S rRNA genes were amplified by PCR using universal primers. The PCR products were partially sequenced (500 bp, ranging from 292 to 1,488 bp) and were
compared to the Ribosomal Database Project database to obtain the most closely matched species.
e
P. validus listed as the invalid synonym P. gordonae in MIDI library.
f
ND, not determined.
g
Identification based on presence of fatty acid 10-methyl 18:0 (TSBA).
b
c
lus group but had the closest match to Arthrobacter oxydans
(SI 0.46). In addition, one of the isolates, PS-P1, was not
identified on several attempts by the MIDI Microbial Identification System but was considered a member of the nocardio-
form group based on the presence of 10-methyl 18:0 fatty acid
(tuberculostearic acid).
The dendrogram (Fig. 1) also shows subclusters within each
of these groups. In the spore-forming Paenibacillus group there
VOL. 67, 2001
PAH-DEGRADING BACTERIA FROM SALT MARSH RHIZOSPHERE
2687
FIG. 1. Euclidean-distance dendrogram based on the whole-cell fatty acid compositions of the sediment and rhizosphere PAH-degrading
isolates. Sediment and rhizosphere isolates are denoted as PS and PR, respectively. In addition, strains isolated using naphthalene are indicated
by an N, while phenanthrene- and biphenyl-enriched isolates are denoted by a P and B, respectively. , isolate PR-P3 is an Arthrobacter sp.
were two subgroups. The members of the smaller subgroup,
consisting of isolates PS-N1, PR-N16, PR-N5, PR-N11, PR-N2,
PR-N12, PS-N6, PR-N1, and SA-N1, all exhibited a mucoid
colony morphology when grown on solid media and had a low
SI to either Paenibacillus alvei, Paenibacillus pabuli, or Paenibacillus validus (Table 1). In contrast, the larger subgroup
within the Paenibacillus cluster exhibited a nonmucoid, mot-
tled colony morphology when grown on solid media, and with
the exception of PR-B2 and DS-N1, had a high SI to P. validus
(Table 1). The five isolates found within the nocardioform
group also formed two subgroups. Isolates PR-N18 and PRN15 both had a cream-colored, waxy colony morphology and
were identified, with low SIs, as Nocardia asteroides and Nocardia brasiliensis, respectively (Table 1). The remaining three
2688
DAANE ET AL.
nocardioforms had an orange, waxy colony morphology and
were identified with a high SI as N. asteroides (Table 1). In the
pseudomonad group, the three isolates PR-N7, PS-N4, and
PS-N5 were identified with a high match to Pseudomonas
stutzeri (Table 1). These three isolates cluster closely together
in relation to the remaining pseudomonad isolates. Of the P.
stutzeri strains, PS-N5 and PR-N7 exhibit a small, crinkled, tan
colony morphology, while PS-N4 exhibits a small, tan colony
morphology. The crinkled, tan morphology can give rise to
noncrinkled colonies resembling PS-N4. This dual colony morphology is commonly seen in P. stutzeri (Norberto Palleroni,
personal communication).
Genomic restriction enzyme digests. RFLP analysis of total
genomic DNA digested with BamHI, EcoRI, HindIII, and NotI
enzymes was used as a screening tool to determine unique
isolates. A large number of gels were performed encompassing
all of the cultured isolates. The gels were examined visually,
and an isolate was considered unique based on the presence of
a unique banding pattern. Isolates which shared a banding
pattern and had similar FAME profiles were grouped together,
and a single isolate from the group was chosen as a representative strain for sequencing and Southern hybridization experiments. The RFLP data are summarized below. All of the six
nocardioform isolates were found to be unique. Of the 12
original pseudomonad isolates, 10 were found to have different
restriction enzyme banding patterns. Shared RFLP patterns
were seen between the pseudomonad isolates PS-N4 and
PS-N5 and between PS-P2a and PS-P2b. The large Paenibacillus group, containing 31 isolates, was found to have 19 isolates
exhibiting unique RFLP patterns. Shared banding patterns
were seen between the mucoid colony morphology Paenibacillus isolates PR-N5, PR-N11, PR-N2, and PR-N12. In addition,
shared banding patterns were seen between the nonmucoid
colony morphology Paenibacillus isolates PR-B2 and PR-B1;
JG-N1, DS-N1, and JG-N3; PR-N19 and PR-N20; PR-N21 and
PR-N22; PR-P1, PR-P2, and PR-P6; and PR-N13, PR-N3, and
PR-N4.
16S rRNA gene analyses. Several of the unique isolates from
each of the three major groups were further identified by
partial sequencing of their 16S rRNA gene. In general, the 16S
rDNA sequence analysis confirmed the identification based on
fatty acid profiles. Within the Paenibacillus group the isolates
identified as P. validus by FAME also had a high sequence
similarity to P. validus. The smaller subgroup of isolates having
a low SI to P. alvei, P. pabuli, or P. validus by FAME had a low
similarity to P. validus based on their partial 16S rRNA gene
sequence (Table 1). Strain PR-P3, which was an outlier within
the Paenibacillus group, was identified with the closest match
by both FAME and 16S rDNA analysis to A. oxydans. The
norcardioform isolates PR-N14 and PR-N15 were identified as
belonging to the genera Rhodococcus and Tsukamurella, respectively. The identification of the Pseudomonas isolates as P.
putida and P. stutzeri by FAME was confirmed by 16S rDNA
phylogenetic analysis. The outlying isolate PR-P12 was identified with a high 16S rDNA sequence similarity to Sphingomonas subarctica.
Southern hybridization with PAH gene probes. All of the
unique isolates, as determined by RFLP, were tested for the
presence of previously described genes for PAH dioxygenases.
A large number of Southern hybridizations were performed,
APPL. ENVIRON. MICROBIOL.
FIG. 2. Southern hybridization of selected isolates with the nah
genes for naphthalene degradation from P. putida NCIB 9816-4.
EcoRI-digested genomic DNA was probed with a 3.5-kb SalI fragment
from P. putida NCIB 9816-4 containing the genes for naphthalene
dioxygenase. M1 and M2 are the digoxigenin-labeled and high-molecular-weight markers, respectively. Lanes 1 to 4 are the pseudomonad
isolates PR-N7, PR-N9, PR-N10, and PS-P2; lane 5 is the sphingomonad isolate PR-P12; lanes 6 and 7 are the nocardioform isolates
PR-N14 and PR-N15; lane 8 is Arthrobacter sp. strain PR-P3; and lanes
9 and 10 represent the mucoid and nonmucoid Paenibacillus isolates
PR-N5 and PR-P1, respectively. P is the positive control P. putida
NCIB 9816-4, and N1 and N2 are the negative controls C. testosteroni
GZ39 and S. yanoikuyae B1, respectively.
with much of the data revealing no hybridization results.
Therefore, we present the data in text form with one figure
exemplifying the nah gene hybridization results for isolates
representing the three major bacterial groups (pseudomonads,
nocardiofoms, and Paenibacillus).
All of the unique pseudomonad isolates with the exception
of PS-P2 hybridized to the classical nah genes from P. putida
NCIB 9816-4. The unique pseudomonad isolates PR-N10, PSN2, PR-N6, Salt-N1, and Salt-N2 gave the same pattern of
hybridization as the control strain P. putida NCIB 9816-4 (Fig.
2). This double-banding pattern was a result of an EcoRI
digestion of the genomic DNA. The positive bands were at 18.5
and 3.7 kb. Pseudomonad isolate PR-N7 also gave a doublebanding pattern to the nah gene probe; however, the bands
were at 9.0 and 1.9 kb. In addition, all of the pseudomonad
isolates, except for PS-P2 and PS-N5, hybridized to the nag
genes cloned from C. testosteroni GZ42. Of the pseudomonad
isolates hybridizing to the nag gene, six isolates (Salt-N1, SaltN2, PR-N6, PS-N2, PR-N10, and PR-N7) gave the same singleband pattern, while the remaining two isolates (PR-N8 and
PR-N9) gave a different single-band pattern distinguishable
from the other six isolates. However, the nocardioforms, Paenibacillus isolates, Arthrobacter isolate PR-P3, or the sphingomonad isolate PR-P12 did not hybridize to either of these
PAH dioxygenase genes (Fig. 2). None of the tested PAHdegrading isolates hybridized to the gene probe from S.
yanoikuyae B1, and only norcardioform isolates PR-N18 and
PS-P1 hybridized to the PAH oxygenase gene cloned from
Mycobacterium sp. strain PY01.
Transformation studies using pure cultures. Isolates representing each group (pseudomonad, nocardioform, and Paenibacillus) and the Arthrobacter isolate PR-P3 were tested for the
VOL. 67, 2001
PAH-DEGRADING BACTERIA FROM SALT MARSH RHIZOSPHERE
TABLE 2. Abilities of several isolates to utilize a variety of PAHs
Isolatea
Pseudomonads
PR-N7
PR-N9
PR-N10
PS-P2
Gram-positive nonspore formers
PR-N15
PR-P3
Spore former
PR-P1
% PAH remainingb
Naphthalene Biphenyl Fluorene Phenanthrene Pyrene
0
0
40
44
85
69
90
51
100
78
97
79
85
71
89
28
90
90
100
100
0
0
84
84
80
42
68
1
100
100
2689
spore-forming A. oxydans strain PR-P3 (Fig. 3). A large variation was seen between the triplicate samples analyzed for the
inoculated microcosms. A two-sample t test was performed on
the pyrene transformation data. The t test indicated that the
pyrene concentrations in the PR-P1- and PR-P3-inoculated microcosms were, by day 35, significantly different (95%
confidence, P 0.05) from the sterile and nonsterile control
microcosms. By day 50 the PR-P1 and PR-P3 high inoculum
density microcosms were found to contain mean concentrations of 3 and 11 ppm of pyrene, respectively (not significantly
different from each other).
DISCUSSION
29
100
Substrate utilization experiments were performed using dense-cell suspensions in MSB medium containing the appropriate PAH at a final concentration
of 100 M. Samples were incubated at 28C at 90 rpm, were extracted with
hexane, and were analyzed by gas chromatography.
b
Percentage remaining was calculated relative to uninoculated controls after
11 days of incubation.
ability to degrade PAHs in cell suspensions. The isolates were
tested for the ability to utilize PAHs individually or as a mixture containing naphthalene, fluorene, phenanthrene, pyrene,
and biphenyl. Table 2 lists the amount (percent) of PAH remaining after 11 days of incubation. Results from cultures
containing a mixture of PAHs were similar to those of cultures
containing a single PAH (data not shown). None of the tested
isolates was able to utilize pyrene in liquid suspension within
the 11-day incubation time; however, all of the isolates were
able to utilize naphthalene completely or partially (PS-P2 and
PR-N10). In general, the strains isolated on phenanthrene
(PS-P2, PR-P3, and PR-P1) were able to utilize a greater
number of PAHs than the strains isolated on naphthalene
(Table 2).
Sediment slurry transformation studies. Two gram-positive
PAH-degrading isolates, PR-P1 (P. validus) and PR-P3 (A.
oxydans), were inoculated at cell densities ranging from 105 to
109 cells ml1 into an aerobic marine sediment slurry containing a mixture of PAHs (naphthalene, fluorene, phenanthrene,
and pyrene) and biphenyl at a concentration of 50 ppm for
each substrate. The transformation experiment was designed
to follow the disappearance of a mixture of PAHs over time,
and the results would determine whether specifically inoculated rhizosphere bacteria could enhance transformation over
the indigenous microbial community. Uninoculated and inoculated sediment slurries both showed loss of the two- and
three-ringed PAHs within 2 weeks of incubation (data not
shown). However, pyrene transformation was only observed in
treatments inoculated with strain PR-P1 or PR-P3 at 109 cells
ml1 (Fig. 3). The sterile slurry microcosms showed no loss of
pyrene, indicating that pyrene transformation was microbially
mediated. In addition, the indigenous bacteria present in the
nonsterile slurry microcosms were not responsible for the loss
of pyrene seen in the inoculated microcosms. Pyrene transformation was not observed in the microcosms inoculated to a cell
density of 105 or 107 cells ml1. Microcosms inoculated with
the spore-forming P. validus strain PR-P1 showed a more rapid
loss of pyrene than did microcosms inoculated with the non-
We present a study of the culturable PAH-degrading bacteria associated with the rhizosphere of several salt marsh plant
species in contaminated and uncontaminated estuarine sediments. In addition, a pasteurization method was successful in
isolating spore-forming bacteria. Numerous studies have demonstrated the importance of the rhizosphere effect on degradation of organic contaminants. Most of these studies have
examined terrestrial plants and agricultural chemicals (1, 2,
27); few have looked at the influence of plant-associated microorganisms on the fate of PCBs (15, 16) and PAHs (34, 39).
There have been a limited number of studies on PAH degradation involving wetland or salt marsh ecosystems, but none
have studied the diversity of PAH-degrading microorganisms
present (28, 30, 49).
Recently, more studies have focused on PAH degradation in
marine and estuarine ecosystems (3, 11, 12, 13, 17, 20, 21, 46).
No studies have been conducted on the PAH-degrading microorganisms associated with salt marsh plants. In the present
study, we isolated a variety of PAH-degrading microorganisms
from the rhizosphere of the salt marsh grasses S. alterniflora, J.
gerardi, D. spicata, and S. airoides. We found both gram-positive (predominantly nocardioform) and gram-negative (predominantly pseudomonad) bacteria, and a pasteurization tech-
FIG. 3. Loss of pyrene in contaminated sediment slurry inoculated
with either the Paenibacillus sp. strain PR-P1 or the Arthrobacter sp.
strain PR-P3. Data points represent the means of triplicate samples.
Error bars indicate the standard deviation. Where no error bar is
present, the value is less than the height of the symbol.
2690
DAANE ET AL.
nique prior to enrichment resulted in the isolation of sporeforming (exclusively Paenibacillus sp.) bacteria.
The rhizosphere has been reported to be predominantly
associated with gram-negative bacteria (7); however, this view
has been changing with more studies using molecular tools (5).
Indeed, the culturable diversity of PAH-degrading microorganisms in this study would lend evidence to this trend. Many
of the bacterial species and genera that we isolated have been
reported to degrade PAHs previously. Few reports have been
made regarding Paenibacillus and its ability to degrade PAHs.
Pichinoty et al. (36) described Bacillus gordonae sp. nov., later
to be emended as P. validus (22), which was able to utilize
p-hydroxybenzoate, phthalate, isophthalate, protocatechuate,
trimellitate, quinate, phenol, p-cresol, and naphthalene. Recently, Meyer et al. (33) reported the isolation of a PAHdegrading tentative Paenibacillus sp. from tar oil-contaminated
soil.
We wanted to determine the ability of the rhizosphere and
estuarine sediment isolates to degrade a variety of PAHs, individually and in combination, in order to determine their
biodegradation and bioremediation potentials. The pure culture transformation experiments showed that phenanthreneenriched isolates were able to utilize a greater variety of other
PAHs than the naphthalene-enriched isolates, although none
degraded pyrene in minimal media. The microbial slurries
demonstrated a greater transformation activity than did the
minimal-medium dense-cell experiments. The greater activity
in the sediment slurries indicates that the organic or inorganic
constituents or the extant microbial community of the sediment stimulated the activity of the inoculated strains. Siciliano
and Germida (41) reported that inoculants required an unknown soil factor to degrade 2-chlorobenzoic acid in the rhizosphere of Dahurian wild rye. It should be noted that both the
pure culture and microbial slurry experiments were performed
under highly oxygenated conditions and that the limited diffusion of oxygen into organic-rich sediments has been found to
restrict PAH biodegradation in the natural environment (10).
The anoxic conditions found in most sediments, however, may
not be the case in tidal and salt marsh ecosystems where plants
provide oxygen to their deeper roots. For example, Spartina is
known to pump oxygen into the rhizosphere (23). Indeed,
when the plants were collected at low tide, highly oxidized
areas were observed around the root system in comparison to
the bulk nonrhizosphere sediment. Of specific interest is evaluating the role of oxygen cycling by roots of salt marsh plants
in the enhancement of PAH biodegradation by rhizosphereassociated microorganisms.
Many different types of PAH oxygenase genes have been
described (for a review, see reference 50). Recent studies using
known PAH degradation genes as probes indicate that previously undescribed genes are present (3). In our study, 75% of
the pseudomonad isolates hybridized to the classical nah gene
from P. putida NCIB 9816-4, and approximately the same number hybridized to the nag genes cloned from C. testosteroni
GZ42. The absence of homology of the Paenibacillus isolates
to all of the tested gene probes indicates the possibility of novel
PAH degradation genes.
Based on our results (morphologic and phenotypic characterization), there is a wide variation between the PAH-degrading isolates, indicating that the rhizosphere of S. alterniflora
APPL. ENVIRON. MICROBIOL.
contains a diverse population of PAH-degrading bacteria. In
addition, vegetated salt marsh ecosystems may hold promise as
a means of remediation of contaminated coastal environments.
ACKNOWLEDGMENTS
This work was supported by a grant from The State of New Jersey
Commission on Science and Technology. G.J.Z. acknowledges the
support of the NSF under grant MCB-9723921.
We thank John Gallagher of the University of Delaware Halophyte
Biology Laboratory for supplying marsh plants; John Cigolini, Jon
Dennis, Lisa Newman, and Michael Murillo for helpful discussions and
assistance; and Gregory Flanagan and Quinn Im for technical assistance.
REFERENCES
1. Anderson, T. A., E. L. Kruger, and J. R. Coats. 1994. Enhanced degradation
of a mixture of three herbicides in the rhizosphere of a herbicide-tolerant
plant. Chemosphere 28:15511557.
2. Arthur, E. L., B. S. Perkovich, T. A. Anderson, and J. R. Coats. 2000.
Degradation of an atrazine and metolachlor herbicide mixture in pesticidecontaminated soils from two agrochemical dealerships in Iowa. Water Air
Soil Pollut. 119:7590.
3. Berardesco, G., S. Dyhrman, E. Gallagher, and M. P. Shiaris. 1998. Spatial
and temporal variation of phenanthrene-degrading bacteria in intertidal
sediments. Appl. Environ. Microbiol. 64:25602565.
4. Boyle, J. J., and J. R. Shann. 1995. Biodegradation of phenol, 2,4-DCP,
2,4-D, and 2,4,5-T in field-collected rhizosphere and nonrhizosphere soils. J.
Environ. Qual. 24:782785.
5. Cattelan, A. J., P. G. Hartel, and J. J. Fuhrmann. 1998. Bacterial composition in the rhizosphere of nodulating and non-nodulating soybean. Soil Sci.
Soc. Am. J. 62:15491555.
6. Cerniglia, C. E. 1993. Biodegradation of polycyclic aromatic hydrocarbons.
Curr. Opin. Biotechnol. 4:331338.
7. Curl, E. A., and B. Truelove. 1986. The rhizosphere. Springer-Verlag, Berlin,
Germany.
8. Daane, L. L., J. A. E. Molina, E. C. Berry, and M. J. Sadowsky. 1996.
Influence of earthworm activity on gene transfer from Pseudomonas fluorescens to indigenous soil bacteria. Appl. Environ. Microbiol. 62:515521.
9. Davies, J. I., and W. C. Evans. 1964. Oxidative metabolism of naphthalene by
soil pseudomonads. Biochem. J. 91:251261.
10. DeLaune, R. D., W. H. Patrick, and M. E. Casselman. 1981. Effect of
sediment pH and redox conditions on degradation of benzo(a)pyrene. Mar.
Pollut. Bull. 12:251253.
11. Dyksterhouse, S. E., J. P. Gray, R. P. Herwig, J. C. Lara, and J. T. Staley.
1995. Cycloclasticus pugetii gen. nov., sp. nov., an aromatic hydrocarbondegrading bacterium from marine sediments. Int. J. Syst. Bacteriol. 45:116
123.
12. Geiselbrecht, A. D., B. P. Hedlund, M. A. Tichi, and J. T. Staley. 1998.
Isolation of marine polycylic aromatic hydrocarbon (PAH)-degrading Cycloclasticus strains from the Gulf of Mexico and composition of their PAH
degradation ability with that of Puget Sound Cycloclasticus strains. Appl.
Environ. Microbiol. 64:47034710.
13. Geiselbrecht, A. D., R. P. Herwig, J. W. Deming, and J. T. Staley. 1996.
Enumeration and phylogenetic analysis of polycyclic aromatic hydrocarbondegrading marine bacteria from Puget Sound sediments. Appl. Environ.
Microbiol. 62:33443349.
14. Gibson, D. T., R. L. Roberts, M. C. Wells, and V. M. Kobal. 1973. Oxidation
of biphenyl by a Beijerinckia species. Biochem. Biophys. Res. Commun.
50:211219.
15. Gilbert, E. S., and D. E. Crowley. 1997. Plant compounds that induce polychlorinated biphenyl biodegradation by Arthrobacter sp. strain B1B. Appl.
Environ. Microbiol. 63:19331938.
16. Gilbert, E. S., and D. E. Crowley. 1998. Repeated application of carvoneinduced bacteria to enhance biodegradation of polychlorinated biphenyls in
soil. Appl. Microbiol. Biotechnol. 50:489494.
17. Gilewicz, M., Nimatuzahroh, T. Nadalig, H. Budzinski, P. Doumenq, V.
Michotey, and J. C. Bertrand. 1997. Isolation and characterization of a
marine bacterium capable of utilizing 2-methylphenanthrene. Appl. Microbiol. Biotechnol. 48:528533.
18. Goyal, A. K., and G. J. Zylstra. 1996. Molecular cloning of novel genes for
polycyclic aromatic hydrocarbon degradation from Comamonas testosteroni
GZ39. Appl. Environ. Microbiol. 62:230236.
19. Goyal, A. K., and G. J. Zylstra. 1997. Genetics of naphthalene and phenanthrene degradation by Comamonas testosteroni. J. Ind. Microbiol. Biotechnol. 19:401407.
20. Guerin, W. F., and G. E. Jones. 1988. Mineralization of phenanthrene by a
Mycobacterium sp. Appl. Environ. Microbiol. 54:937944.
21. Hedlund, B. P., A. D. Geiselbrecht, T. J. Bair, and J. T. Staley. 1999.
VOL. 67, 2001
22.
23.
24.
25.
26.
27.
28.
29.
30.
31.
32.
33.
34.
35.
36.
PAH-DEGRADING BACTERIA FROM SALT MARSH RHIZOSPHERE
Polycyclic aromatic hydrocarbon degradation by a new marine bacterium,
Neptunomonas naphthovorans gen. nov., sp. nov. Appl. Environ. Microbiol.
65:251259.
Heyndrickx, M., K. Vandemeulebroecke, P. Scheldeman, B. Hoste, K. Kersters, P. De Vos, N. A. Logan, A. M. Aziz, N. Ali, and R. C. W. Berkeley. 1995.
Paenibacillus (formerly Bacillus) gordonae (Pichinoty et al. 1986) Ash et al.
1994 is a later subjective synonym of Paenibacillus (formerly Bacillus) validus
(Nakamura 1984) Ash et al. 1994: emended description of P. validus. Int. J.
Syst. Bacteriol. 45:661669.
Howes, B. L., and J. M. Teal. 1994. Oxygen loss from and its relationship to
salt marsh oxygen balance. Oecologia 97:431438.
Johnson, J. L. 1994. Similarity analysis of rRNAs, p. 683700. In P. Gerhardt,
R. G. E. Murray, W. A. Wood, and N. R. Krieg (ed.), Methods for general
and molecular bacteriology. American Society for Microbiology, Washington, D.C.
Kadlec, R., and R. L. Knight. 1995. Treatment wetlands. Lewis Publishers,
Boca Raton, Fla.
Kim, E., and G. J. Zylstra. 1995. Molecular and biochemical characterization
of two meta-cleavage dioxygenases involved in biphenyl and m-xylene degradation by Beijerinckia sp. strain B1. J. Bacteriol. 177:30953103.
Lappin, H. M., M. P. Greaves, and J. H. Slater. 1985. Degradation of the
herbicide mecoprop [2-(2-methyl-4-chlorphenoxy)propionic acid] by a synergistic microbial community. Appl. Environ. Microbiol. 49:429433.
Lin, Q., I. A. Mendelssohn, C. B. Henry, P. O. Roberts, M. M. Walsh, E. B.
Overton, and R. J. Portier. 1999. Effects of bioremediation agents on oil
degradation in mineral and sandy salt marsh sediments. Environ. Technol.
20:825837.
Lotufo, G. R. 1998. Bioaccumulation of sediment-associated fluoranthene in
benthic copepods: uptake, elimination and biotransformation. Aquat. Toxicol. 44:115.
Machate, T., H. Noll, H. Behrens, and A. Kettrup. 1997. Degradation of
phenanthrene and hydraulic characteristics in a constructed wetland. Water
Res. 31:554560.
Maidak, B. L., J. R. Cole, T. G. Lilburn, C. T. Parker, Jr., P. R. Saxman,
J. M. Stredwick, G. M. Garrity, B. Li, G. J. Olsen, S. Pramanik, T. M.
Schmidt, and J. M. Tiedje. 2000. The RDP (Ribosomal Database Project)
continues. Nucleic Acids Res. 28:173174.
Menzie, C. A., B. B. Potocki, and J. Santodonato. 1992. Exposure to carcinogenic PAHs in the environment. Environ. Sci. Technol. 26:12781284.
Meyer, S., R. Mose, A. Neef, U. Stahl, and P. Ka
mpfer. 1999. Differential
detection of key enzymes of polyaromatic-hydrocarbon-degrading bacteria
using PCR and gene probes. Microbiology 145:17311741.
Nichols, T. D., D. C. Wolf, H. B. Rogers, C. A. Beyrouty, and C. M. Reynolds.
1997. Rhizosphere microbial populations in contaminated soils. Water Air
Soil Pollut. 95:165178.
Paul, E. A., and F. E. Clark. 1989. Soil microbiology and biochemistry.
Academic Press, New York, N.Y.
Pichinoty, F., J. B. Waterbury, M. Mandel, and J. Asselineau. 1986. Bacillus
37.
38.
39.
40.
41.
42.
43.
44.
45.
46.
47.
48.
49.
50.
2691
gordonae sp. nov., une nouvelle espace appartenant au second groupe morphologique, degrandant divers composes aromatiques. Ann. Inst. Pasteur/
Microbiol. (Paris) 137A:6578.
Reilley, K. A., M. K. Banks, and A. P. Schwab. 1996. Dissipation of polycyclic
aromatic hydrocarbons in the rhizosphere. J. Environ. Qual. 25:212219.
Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular cloning: a
laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold
Spring Harbor, N.Y.
Schwab, A. P., and M. K. Banks. 1994. Biologically mediated dissipation of
polyaromatic hydrocarbons in the root zone, p. 132141. In T. A. Anderson
and J. R. Coats (ed.), Bioremediation through rhizosphere technology.
American Chemical Society, Washington, D.C.
Serdar, C. M., and D. T. Gibson. 1989. Studies of nucleotide sequence
homology between naphthalene-utilizing strains of bacteria. Biochem. Biophys. Res. Commun. 164:772779.
Siciliano, S. D., and J. J. Germida. 1999. Enhanced phytoremediation of
chlorobenzoates in rhizosphere soil. Soil Biol. Biochem. 31:299305.
Simon, M. J., T. D. Osslund, R. Saunders, B. D. Ensley, S. Suggs, A.
Harcourt., W.-C. Suen, D. L. Cruden, D. T. Gibson, and G. J. Zylstra. 1993.
Sequences of genes encoding naphthalene dioxygenase in Pseudomonas
putida strains G7 and NCIB 9816-4. Gene 127:3137.
Stanier, R. Y., N. J. Palleroni, and M. Doudoroff. 1966. The aerobic pseudomonads: a taxonomic study. J. Gen. Microbiol. 43:159271.
Sutherland, J. B., F. Rafii, A. A. Khan, and C. E. Cerniglia. 1995. Mechanisms of polycyclic aromatic dydrocarbon degradation, p. 269306. In L. Y.
Young and C. E. Cerniglia (ed.), Microbial transformation and degradation
of toxic organic chemicals. John Wiley and Sons, Inc., New York, N.Y.
Walton, B. T., and T. A. Anderson. 1990. Microbial degradation of trichloroethylene in the rhizosphere: potential application to biological remediation of waste sites. Appl. Environ. Microbiol. 56:10121016.
West, P. A., G. C. Okpokwasili, P. R. Brayton, D. J. Grimes, and R. R.
Colwell. 1984. Numerical taxonomy of phenanthrene-degrading bacteria isolated from the Chesapeake Bay. Appl. Environ. Microbiol. 48:988993.
Wetzel, S. C., M. K. Banks, and A. P. Schwab. 1996. Rhizosphere effects on
the degradation of pyrene and anthracene in soil, p. 254262. In E. L.
Kruger, T. A. Anderson, and J. R. Coats (ed.), Phytoremediation of soil and
water contaminants. American Chemical Society, Washington, D.C.
Wilson, K. 1994. Preparation of genomic DNA from bacteria, p. 2.4.12.4.5.
In F. M. Ausubel, R. Brent, R. E. Kingston, D. D. Moore, J. G. Seidman,
J. A. Smith and K. Struhl (ed.), Current protocols in molecular biology, vol.
1. John Wiley & Sons, New York, N.Y.
Wright, A. L., R. W. Weaver, and J. W. Webb. 1997. Oil bioremediation in
salt marsh mesocosms as influenced by N and P fertilization, flooding, and
season. Water Air Soil Pollut. 95:179191.
Zylstra, G. J., E. Kim, and A. Goyal. 1997. Comparative molecular analysis
of genes for polycyclic aromatic hydrocarbon degradation. Genet. Eng. 19:
257269.
You might also like
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryFrom EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryRating: 3.5 out of 5 stars3.5/5 (231)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)From EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Rating: 4.5 out of 5 stars4.5/5 (119)
- Never Split the Difference: Negotiating As If Your Life Depended On ItFrom EverandNever Split the Difference: Negotiating As If Your Life Depended On ItRating: 4.5 out of 5 stars4.5/5 (838)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaFrom EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaRating: 4.5 out of 5 stars4.5/5 (265)
- The Little Book of Hygge: Danish Secrets to Happy LivingFrom EverandThe Little Book of Hygge: Danish Secrets to Happy LivingRating: 3.5 out of 5 stars3.5/5 (399)
- Grit: The Power of Passion and PerseveranceFrom EverandGrit: The Power of Passion and PerseveranceRating: 4 out of 5 stars4/5 (587)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyFrom EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyRating: 3.5 out of 5 stars3.5/5 (2219)
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeFrom EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeRating: 4 out of 5 stars4/5 (5794)
- Team of Rivals: The Political Genius of Abraham LincolnFrom EverandTeam of Rivals: The Political Genius of Abraham LincolnRating: 4.5 out of 5 stars4.5/5 (234)
- Shoe Dog: A Memoir by the Creator of NikeFrom EverandShoe Dog: A Memoir by the Creator of NikeRating: 4.5 out of 5 stars4.5/5 (537)
- The Emperor of All Maladies: A Biography of CancerFrom EverandThe Emperor of All Maladies: A Biography of CancerRating: 4.5 out of 5 stars4.5/5 (271)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreFrom EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreRating: 4 out of 5 stars4/5 (1090)
- Her Body and Other Parties: StoriesFrom EverandHer Body and Other Parties: StoriesRating: 4 out of 5 stars4/5 (821)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersFrom EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersRating: 4.5 out of 5 stars4.5/5 (344)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceFrom EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceRating: 4 out of 5 stars4/5 (890)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureFrom EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureRating: 4.5 out of 5 stars4.5/5 (474)
- The Unwinding: An Inner History of the New AmericaFrom EverandThe Unwinding: An Inner History of the New AmericaRating: 4 out of 5 stars4/5 (45)
- The Yellow House: A Memoir (2019 National Book Award Winner)From EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Rating: 4 out of 5 stars4/5 (98)
- Apu White SmokeDocument3 pagesApu White SmokeGoutam SahaNo ratings yet
- On Fire: The (Burning) Case for a Green New DealFrom EverandOn Fire: The (Burning) Case for a Green New DealRating: 4 out of 5 stars4/5 (73)
- Measure RPA ROI with KPIsDocument4 pagesMeasure RPA ROI with KPIsAdnan FarooqNo ratings yet
- Fractal ForestsDocument50 pagesFractal ForestsWell Fournier0% (1)
- Niche PartitioningDocument3 pagesNiche PartitioningKhang LqNo ratings yet
- Insulation Resistance TestDocument7 pagesInsulation Resistance Testcarlos vidalNo ratings yet
- GA 33 KV VCB HT Panel - Siddharth Nagar Project. UPDocument17 pagesGA 33 KV VCB HT Panel - Siddharth Nagar Project. UPaayushNo ratings yet
- 02 Stuck Pipe - Free Point & Back Off PDFDocument31 pages02 Stuck Pipe - Free Point & Back Off PDFام فاطمة البطاط100% (2)
- Dictionary of Oil Industry TerminologyDocument79 pagesDictionary of Oil Industry Terminologyniksharris100% (22)
- Classic Text Messages Morning GreetingsDocument2 pagesClassic Text Messages Morning GreetingsDhamukarthikeyanNo ratings yet
- Simulation of BJT Amplifier: Course - Section: ECE20L-E06 Group NumberDocument10 pagesSimulation of BJT Amplifier: Course - Section: ECE20L-E06 Group NumberLuch ÜNo ratings yet
- Rip Bushing PDFDocument38 pagesRip Bushing PDFTravis Wood100% (1)
- BW1114-B2 Bendix Brake CatalogDocument116 pagesBW1114-B2 Bendix Brake Cataloggearhead1100% (1)
- ISSAQ: An Integrated Sensing Systems For Real-Time Indoor Air Quality MonitoringDocument15 pagesISSAQ: An Integrated Sensing Systems For Real-Time Indoor Air Quality MonitoringKemHuyềnNo ratings yet
- Chapter 1 - Notes (Properties of Fluid) PDFDocument23 pagesChapter 1 - Notes (Properties of Fluid) PDFHappy Ocean100% (1)
- Bing WorksheetDocument3 pagesBing WorksheetFrutti MataniNo ratings yet
- Electrochemistry PD lab insightsDocument4 pagesElectrochemistry PD lab insightsEdilberto PerezNo ratings yet
- INCaDocument47 pagesINCaMehdi SoltaniNo ratings yet
- 3M Water Filtration Products - High Flow Series - HF40 and HF40 - S Performance Data SheetDocument2 pages3M Water Filtration Products - High Flow Series - HF40 and HF40 - S Performance Data SheetSergioNo ratings yet
- Valve Type Trim Type CF XTDocument1 pageValve Type Trim Type CF XTAye KyweNo ratings yet
- ICN Question Bank Unit-1, 2 and 3 (Upto GSM Identifier)Document1 pageICN Question Bank Unit-1, 2 and 3 (Upto GSM Identifier)Snehal PatelNo ratings yet
- Nervous System Reaction PaperDocument3 pagesNervous System Reaction PaperJohn Ruel Sanchez IINo ratings yet
- LGBT Workplace Equality Policy and Customer Satisfaction: The Roles of Marketing Capability and Demand InstabilityDocument20 pagesLGBT Workplace Equality Policy and Customer Satisfaction: The Roles of Marketing Capability and Demand InstabilityFatima ZafarNo ratings yet
- M & E Eyerusalem TesfayeDocument12 pagesM & E Eyerusalem Tesfayeeyerusalem tesfayeNo ratings yet
- Closed Coke Slurry System: An Advanced Coke Handling ProcessDocument33 pagesClosed Coke Slurry System: An Advanced Coke Handling ProcessFayaz MohammedNo ratings yet
- GCMS-QF 15 - Calibration (IMTE) Form - MPSDocument7 pagesGCMS-QF 15 - Calibration (IMTE) Form - MPSMobin Thomas AbrahamNo ratings yet
- Arrester Facts 004a - Externally Gapped ArresterDocument2 pagesArrester Facts 004a - Externally Gapped ArresterCbdtxd PcbtrNo ratings yet
- British Airways Case Study SolutionDocument2 pagesBritish Airways Case Study SolutionHassan ZafarNo ratings yet
- A Fracture Mechanics Analysis of The Texture of Fried Potato Crust PDFDocument7 pagesA Fracture Mechanics Analysis of The Texture of Fried Potato Crust PDFRomaric OuetchehouNo ratings yet
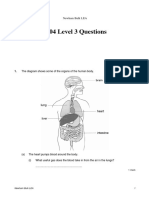
- 2004 Level 3 Questions: Newham Bulk LEADocument18 pages2004 Level 3 Questions: Newham Bulk LEAPatience NgundeNo ratings yet
- Anti Climbers FlyerDocument2 pagesAnti Climbers Flyeredark2009No ratings yet