Professional Documents
Culture Documents
Creating Standard Curves With Genomic DNA or Plasmid DNA Templates For Use in Quantitative PCR
Uploaded by
rickOriginal Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
Creating Standard Curves With Genomic DNA or Plasmid DNA Templates For Use in Quantitative PCR
Uploaded by
rickCopyright:
Available Formats
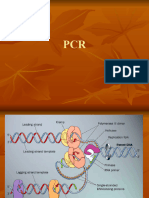
Creating Standard Curves with Genomic DNA or
Plasmid DNA Templates for Use in Quantitative PCR
Overview Genomic DNA (gDNA) and plasmids containing cloned target sequences are
commonly used as standards in quantitative PCR. This tutorial reviews calculations that
can be used for determining the mass of gDNA and plasmid templates that correspond
to copy numbers of target nucleic acid sequences.
Important Notes
• It is generally not possible to use DNA as a standard for absolute quantification
of RNA because there is no control for the efficiency of the reverse transcription
step.
• Once prepared, it is recommended to dilute standards into small aliquots, store at
–80 °C, and thaw only once before use.
• Accurate pipetting is essential because the standards must be diluted over
several orders of magnitude.
• Plasmid DNA must be a single, pure species. For example, plasmid DNA
prepared from E. coli often is contaminated with RNA, which increases the A260
measurement and inflates the copy number determined for the plasmid.
• Because of potential errors with pipetting and/or OD values, it is important to
verify the absolute quantities of an absolute standard by some independent
method. For example, one might use real-time PCR to compare several genomic
DNA samples of known target quantity with a plasmid standard curve (containing
the same target) to verify accuracy of the standards.
• Because plasmid (and to a lesser extent gDNA) sequences are highly abundant,
they can be sources of PCR contamination. Extreme caution must be exercised
when working with these DNAs to prevent their exposure to stock PCR reagents,
solvents used for the dilution of PCR reagents and laboratory equipment and
surfaces.
Example: Creating a gDNA Standard Curve
Prepare a standard curve in which a gene of interest is present at 300,000 copies,
30,000 copies, 3,000 copies, 300 copies and 30 copies. This example uses the human
RNaseP gene, a gene that exists as a single copy per haploid genome (or 2 copies per
human cell).
Step 1
Identify the genome size of the organism of interest.
The size of the human genome as determined by Celera Genomics is approximately 3
billion bp (haploid).
For Reference Only Page 1 of 9
If the genome size of the organism of interest is not readily available, go to
http://www.cbs.dtu.dk/databases/DOGS/index.html
and select the appropriate database. Below is an excerpt from the
“Abbreviated database (By name)”
Size of human
genome
(haploid)
The estimate of 3,400,000,000 bp, or 3.4e9 bp (haploid), is consistent with Celera’s
estimate of 3,000,000,000 bp or 3.0e9 bp (haploid).
Step 2
Identify the mass of DNA per genome
Calculate the mass of the genome by inserting the genome-size value in the formula
below (see page 8 for derivation of this formula).
m = n 1.096e-21 g where: n = genome size (bp)
bp m = mass
e-21 = ×10-21
The mass of the human genome (haploid) is calculated as follows.
m = 3.0e9 bp 1.096e-21 g = 3.3e−12 g
bp
The calculation below converts the mass to picogram units.
3.3e-12 g 1e12 pg = 3.3 pg
g
Step 3:
Divide the mass of the genome by the copy number of the gene of interest per haploid
genome.
The RNase P gene is a target that exists as a single copy gene per haploid genome (or
2 copies per human cell).
3.3 pg/genome ÷ 1 copy RNase P/genome = 3.3 pg genome = 3.3 pg
genome 1 copy 1 copy RNase P
For Reference Only Page 2 of 9
Therefore, 3.3 pg of human gDNA contains one copy of the RNase P gene.
Step 4
Calculate the mass of gDNA containing the copy #s of interest, that is 300,000 to 30
copies.
Copy # of interest × mass of haploid genome = mass of gDNA needed
Copy # Mass of gDNA
needed (pg)
300,000 990,000
30,000 × 3.3 pg 99,000
3,000 9,900
300 990
30 99
Step 5
Calculate the concentrations of gDNA needed to achieve the copy#s of interest. Divide
the mass needed (calculated in Step 4) by the volume to be pipetted into each reaction.
In this example, 5µL of gDNA solution will be pipetted into each PCR reaction.
Calculate the concentration of gDNA needed to achieve the required masses of gDNA.
Copy # Mass of gDNA Final concentration
needed (pg) (pg/µl) of gDNA
300,000 990,000 198,000
30,000 99,000 ÷ 5 µL 19,800
3,000 9,900 1,980
300 990 198
30 99 19.8
Step 6
Prepare a serial dilution of the gDNA.
For the dilutions we will use the formula,
C1V1 = C2V2
The stock concentration of human gDNA was determined by spectrophotometric
analysis to be 1.2 µg/µl. Therefore, in this example, C1 = 1.2 µg/µL or 1,200,000 pg/µL.
Each dilution prepared has a final volume (V2) of 100µL.
For Reference Only Page 3 of 9
Dilution #1
1,200,000 pg V1 = 198,000 pg 100 µL
µL µL
V1 = 16.5 µL
Volume of diluent = 100 µL – 16.5 µL = 83.5 µL
To achieve the final volume of 100 µL, add 16.5 µL of stock gDNA to 83.5 µL of diluent.
Note: The diluent can be sterile 1X TE (1mM Tris, 0.1mM EDTA, pH8.0) or sterile,
nuclease-free H2O.1
Dilutions 2 to 5 were calculated using the same types of calculations (C1V1 = C2V2) as
presented above for Dilution #1.
The following table presents the calculated volumes of gDNA and diluent for all 5
dilutions.
Dilution Source Initial Volume Volume Final Final Resulting
# of concentration of of Volume concentration copy #
gDNA (pg/µL) gDNA diluent (µL) of dilution RNase P
for (µL) (µL) (pg/µl) gene/ 5µl
dilution
C1 V1 V2 C2
1 stock 1,200,000 16.5 83.5 100 198,000 300,000
2 Dilution 198,000 10 90 100 19,800 30,000
1
3 Dilution 19,800 10 90 100 1,980 3,000
2
4 Dilution 1,980 10 90 100 198 300
3
5 Dilution 198 10 90 100 19.8 30
4
1
It is the users responsibility to determine whether the use of background nucleic acid will impact assay
performance (ex. PCR efficiency). Background nucleic acid is DNA or RNA that can be spiked into the
standard so as to mimic the biological unknown samples.
For Reference Only Page 4 of 9
Example: Creating a Standard Curve with a Plasmid DNA Template2
Background
Prepare a standard curve in which the cloned ß-actin sequence is present at 300,000
copies, 30,000 copies, 3,000 copies, 300 copies and 30 copies. The plasmid size is
15,000 bp. The stock of plasmid DNA was determined to be 2.0 µg/µL by
spectrophotometric analysis. The PCR reactions are set-up such that 5µL of plasmid
DNA are pipetted into each PCR reaction.
Step 1
Calculate the mass of a single plasmid molecule.
Insert the plasmid size value into the formula below (see page 8 for derivation of this
formula):
m = n 1. 096e-21 g where: n = plasmid size (bp)
bp m = mass
e-21 = ×10-21
Note: Use the size of the entire plasmid (plasmid + insert) in the calculation above
instead of the size of the insert alone.
Mass of one plasmid molecule= 15,000 bp 1.096e-21 g = 1.64e-17 g
bp
Step 2
Calculate the mass of plasmid containing the copy #s of interest, that is 300,000 to 30
copies.
Copy # of interest × mass of single plasmid = mass of plasmid DNA needed
For example, mass of plasmid DNA containing 300,000 copies of B-actin sequence is
as follows.
1.64e-17 g 300,000 copies = 4.92e-12 g
copy
2
It is the users responsibility to determine whether linearization of the plasmid standard with a restriction
endonuclease will impact assay performance (ex. PCR efficiency).
For Reference Only Page 5 of 9
The following table presents the calculated plasmid masses needed to achieve the copy
numbers of interest.
Copy # Mass of plasmid
DNA (g)
300,000 4.92e-12
30,000 × 1.64e-17 g 4.92e-13
3,000 4.92e-14
300 4.92e-15
30 4.92e-16
Step 3
Calculate the concentrations of plasmid DNA needed to achieve the copy#s of interest.
Divide the mass needed (calculated in Step 2) by the volume to be pipetted into each
reaction.
In this example, 5µL of plasmid DNA solution is pipetted into each PCR reaction.
Calculate the concentration of gDNA needed to achieve the required masses of gDNA.
Copy # Mass of plasmid Final concentration of
DNA needed (g) plasmid DNA (g/µL)
300,000 4.92e-12 9.84e-13
30,000 4.92e-13 ÷ 5 µL 9.84e-14
3,000 4.92e-14 9.84e-15
300 4.92e-15 9.84e-16
30 4.92e-16 9.84e-17
Step 4
Prepare a serial dilution of the plasmid DNA.
Cloned sequences are highly concentrated in purified plasmid DNA stocks. A series of
serial dilutions must be performed to achieve a working stock of plasmid DNA for
quantitative PCR applications. The table on page 7 shows that the first 3 dilutions (each
1:100) were prepared so that the plasmid would be at a workable concentration, that is
2e-12 grams/µL or 1.32e5 copies/µL.
Once the plasmid is at a workable concentration, use the following formula to calculate
the volume needed to prepare the 300,000 copy standard dilution (Dilution #4).
C1V1 = C2V2
For Reference Only Page 6 of 9
Dilution #4
(see table below for C1 ,C2,V1 and V2 values)
2e-12 g V1 = 9.84e-13 g 100 µL
µL µL
V1 = 49.2 µL
Volume of diluent = 100 µL – 49.2 µL = 50.8 µL
To achieve the final volume of 100 µL, add 49.2 µL of stock gDNA to 50.8 µL of diluent.
Note: The diluent can be sterile 1X TE (1mM Tris, 0.1mM EDTA, pH8.0) or sterile,
nuclease-free H2O. 3
Dilutions 5 to 8 were calculated using the same types of calculations as Dilution #4
above.
Source of Initial Volume Volume of Final Final conc. Resulting
plasmid conc. of diluent Volume in copy # of
Dilution #
DNA for (grams/µL) plasmid (µL) (µL) (g/µl) ß-actin
dilution DNA sequence /
(µL) 5 µl
C1 V1 V2 C2
1 stock 2e-06 10 µl 990 µl 1000 µl 2e-08 N/A
2 Dilution 1 2e-08 10 µl 990 µl 1000 µl 2e-10 N/A
3 Dilution 2 2e-10 10 µl 990 µl 1000 µl 2e-12 N/A
4 Dilution 3 2e-12 49.2 µl 50.8 µl 100 µl 9.84e-13 300,000
5 Dilution 4 9.84e-13 10 µl 90 µl 100 µl 9.84e-14 30,000
6 Dilution 5 9.84e-14 10 µl 90 µl 100 µl 9.84e-15 3,000
7 Dilution 6 9.84e-15 10 µl 90 µl 100 µl 9.84e-16 300
8 Dilution 7 9.84e-16 10 µl 90 µl 100 µl 9.84e-17 30
In the example above, dilutions 4 to 8 would be used for the quantitative PCR
application.
3
It is the users responsibility to determine whether the use of background nucleic acid will impact assay
performance (ex. PCR efficiency). Background nucleic acid is DNA or RNA that can be spiked into the
standard so as to mimic the biological unknown samples.
For Reference Only Page 7 of 9
Derivation of DNA Mass Formula
m = n 1.096e-21 g
bp
The formula above was derived as follows
m = n 1 mole 660 g = n 1.096e-21 g
6.023e23 molecules (bp) mole bp
where:
n = DNA size (bp)
m = mass
Avogadros number = 6.023e23 molecules / 1 mole
Average MW of a double-stranded DNA molecule = 660 g/mole
For Reference Only Page 8 of 9
2003. Applied Biosystems. All rights reserved.
For Research Use Only. Not for use in diagnostic procedures.
The PCR process is covered by patents owned by Roche Molecular Systems, Inc. and
F. Hoffmann-La Roche Ltd.
Applied Biosystems is a registered trademark and AB (Design), Applera, Celera and
Celera Genomics are trademarks of Applera Corporation or its subsidiaries in the US
and/or certain other countries.
For Reference Only Page 9 of 9
You might also like
- Lab 21A and 21BDocument8 pagesLab 21A and 21BLateesha ThomasNo ratings yet
- Mushrooms Russia and HistoryDocument579 pagesMushrooms Russia and Historyzeevnaomi100% (4)
- MKBS221. PRAC 5 REPORT Last OneDocument11 pagesMKBS221. PRAC 5 REPORT Last OneNOMKHULEKO ALICENo ratings yet
- Generation of External Standards For Real Time PCR (Relative Quantitication)Document4 pagesGeneration of External Standards For Real Time PCR (Relative Quantitication)Morteza HaghiNo ratings yet
- PCR, RT-PCR, Nested-Pcr, Multiplex PCR, Quantitative PCRDocument116 pagesPCR, RT-PCR, Nested-Pcr, Multiplex PCR, Quantitative PCRYunizardiNo ratings yet
- Principle of The PCR: This Technique Is Invented in 1984 by Kary Mulis. The Purpose of A PCRDocument11 pagesPrinciple of The PCR: This Technique Is Invented in 1984 by Kary Mulis. The Purpose of A PCRantesar reheemNo ratings yet
- # G /genome 3.1x10 1.66x10 G 5.1x10 G /genomeDocument2 pages# G /genome 3.1x10 1.66x10 G 5.1x10 G /genomerickNo ratings yet
- Lab Manual 3Document7 pagesLab Manual 3syazaismailNo ratings yet
- PCR and Agarose Gel ElectrophoresisDocument6 pagesPCR and Agarose Gel ElectrophoresisÍrisNo ratings yet
- PCR Labwork 2 ENGDocument4 pagesPCR Labwork 2 ENGmigas1996No ratings yet
- Supporting Information: Supplementary Note 1 Encoding: Data To DNA SequencesDocument25 pagesSupporting Information: Supplementary Note 1 Encoding: Data To DNA Sequencesweixing wangNo ratings yet
- Quiz #3: Biochemical Engineering Fall 2003Document5 pagesQuiz #3: Biochemical Engineering Fall 2003Princess Janine CatralNo ratings yet
- PCRDocument35 pagesPCRk5zo7jNo ratings yet
- Cold Spring Harb Protoc-2018-Green-pdb - Prot095117 PCRDocument9 pagesCold Spring Harb Protoc-2018-Green-pdb - Prot095117 PCRJosman SalgadoNo ratings yet
- The Polymerase Chain Reaction and Sequence-Tagged SitesDocument7 pagesThe Polymerase Chain Reaction and Sequence-Tagged SitesPisi MicaNo ratings yet
- Ex No.-03 Polymerase Chain ReactionDocument4 pagesEx No.-03 Polymerase Chain ReactionSujit ShandilyaNo ratings yet
- DNA Quantification Using SpectrophotometryDocument19 pagesDNA Quantification Using Spectrophotometryella hullezaNo ratings yet
- The Polymerase Chain Reaction (PCR)Document50 pagesThe Polymerase Chain Reaction (PCR)Deana NamirembeNo ratings yet
- Creating Standard and Copy Number CalculationDocument2 pagesCreating Standard and Copy Number CalculationDeepak Singh LourembamNo ratings yet
- Supplementary For ValidationDocument28 pagesSupplementary For Validation201 101No ratings yet
- Prot m3045 Probe enDocument4 pagesProt m3045 Probe enLuis Arístides Torres SánchezNo ratings yet
- Electrophoresis Ch. 3 - 1Document8 pagesElectrophoresis Ch. 3 - 1Ajit PunjNo ratings yet
- TYPES OF PCR - Dr. De-1 PDFDocument33 pagesTYPES OF PCR - Dr. De-1 PDFJAICHANDRUNo ratings yet
- Polymerase Chain Reaction (PCR) : Evros Polydorou April 2018Document11 pagesPolymerase Chain Reaction (PCR) : Evros Polydorou April 2018Evros PolidorouNo ratings yet
- Genomic Dna ManualDocument7 pagesGenomic Dna ManualZafran KhanNo ratings yet
- Lab MathDocument2 pagesLab MathaishwaryadashNo ratings yet
- Quantitative RT-PCR Protocol (Sybr Green I)Document8 pagesQuantitative RT-PCR Protocol (Sybr Green I)u77No ratings yet
- Early History of Microbiological TechniqueDocument5 pagesEarly History of Microbiological TechniqueFadare Shadrach ONo ratings yet
- RT-PCR Two-Steps ProtocolDocument13 pagesRT-PCR Two-Steps ProtocolFrancisco MartinezNo ratings yet
- Primer Extension System - AMV Reverse Transcriptase ProtocolDocument13 pagesPrimer Extension System - AMV Reverse Transcriptase ProtocoldvNo ratings yet
- 1 KbladderDocument4 pages1 KbladderTan E-ChuanNo ratings yet
- Polymerase Chain Reaction: Salwa Hassan TeamaDocument56 pagesPolymerase Chain Reaction: Salwa Hassan TeamaSamiksha SharmaNo ratings yet
- 1 KB DNA Ladder: 5' DNA Terminus Labeling Protocol (Phosphate Exchange Reaction)Document4 pages1 KB DNA Ladder: 5' DNA Terminus Labeling Protocol (Phosphate Exchange Reaction)Abdessamad EladakNo ratings yet
- Analiza PCR - ThermocyclerDocument7 pagesAnaliza PCR - Thermocyclerionescu_ion_3No ratings yet
- Micro20 - Polymerase Chain ReactionDocument4 pagesMicro20 - Polymerase Chain Reactionaman jaiswalNo ratings yet
- Experiment Record 1Document7 pagesExperiment Record 1JeremyLeNo ratings yet
- Protein Concentration Measurement MH 2017Document2 pagesProtein Concentration Measurement MH 2017kiệt nguyễn lêNo ratings yet
- Biotechnology Plasmid LabDocument12 pagesBiotechnology Plasmid LabCristian DumitrescuNo ratings yet
- Evaluation of Omega Mag-Bind TotalPure NGS Beads MWeitzman April2018Document8 pagesEvaluation of Omega Mag-Bind TotalPure NGS Beads MWeitzman April2018Shaheen AlamNo ratings yet
- Estimate Age of DNADocument3 pagesEstimate Age of DNAashueinNo ratings yet
- MedGen 06week Lab PCRDocument5 pagesMedGen 06week Lab PCRМөнхгэрэл ГанбатNo ratings yet
- NMR End-Group AnalysisDocument5 pagesNMR End-Group AnalysisHaesoo LeeNo ratings yet
- Measuring Trypsin Inhibitor in Soy MealDocument19 pagesMeasuring Trypsin Inhibitor in Soy Mealgabytza_chNo ratings yet
- Scribe4 PCR 11Document4 pagesScribe4 PCR 11ulfanniNo ratings yet
- Chapter 6 Nucleic Acid AmplificationDocument7 pagesChapter 6 Nucleic Acid AmplificationIsraa Al-AlemNo ratings yet
- Genomics Lectures 9 To 14-2023 PDFDocument65 pagesGenomics Lectures 9 To 14-2023 PDFAhire Ganesh Ravindra bs20b004No ratings yet
- Pset 3Document2 pagesPset 3RajNo ratings yet
- PCR and Agarose Gel ElectrophoresisDocument5 pagesPCR and Agarose Gel ElectrophoresisHusna AdilaNo ratings yet
- Gene Specific RTPCRDocument4 pagesGene Specific RTPCRQuimico Inmunologia GeneticaNo ratings yet
- Genei: Student PCR Teaching Kit ManualDocument13 pagesGenei: Student PCR Teaching Kit ManualSoma GhoshNo ratings yet
- Lab 2 ds180 Genotyping LabDocument7 pagesLab 2 ds180 Genotyping Labapi-342081300No ratings yet
- MMBIO Exam1 Review - 241Document23 pagesMMBIO Exam1 Review - 241Riley SarabiaNo ratings yet
- Primer E. ColiDocument4 pagesPrimer E. Coliromario1313No ratings yet
- Usefulness ofDocument42 pagesUsefulness ofAdarshBijapurNo ratings yet
- Chap. 5 Molecular Genetic Techniques: TopicsDocument15 pagesChap. 5 Molecular Genetic Techniques: TopicsPAGINo ratings yet
- 7.02 Fall 2014 Genetics Day 1 In-Lab QuestionsDocument5 pages7.02 Fall 2014 Genetics Day 1 In-Lab QuestionsakiridinoNo ratings yet
- MD Simulation Tutorial in GromacsDocument9 pagesMD Simulation Tutorial in GromacsSimanta PaulNo ratings yet
- 1kb Plus LadderDocument4 pages1kb Plus LadderPedro SoaresNo ratings yet
- Cms 040377Document6 pagesCms 040377Ani IoanaNo ratings yet
- BigDye v3.1Document2 pagesBigDye v3.1pdf2006No ratings yet
- Standard and Super-Resolution Bioimaging Data Analysis: A PrimerFrom EverandStandard and Super-Resolution Bioimaging Data Analysis: A PrimerNo ratings yet
- Light Sources PDFDocument39 pagesLight Sources PDFdiego.pavanello7081No ratings yet
- UVGI Disinfection TheoryDocument35 pagesUVGI Disinfection TheoryrickNo ratings yet
- Biophysical Modeling: Induction of Clustered DNA Lesions by Ionizing Radiation - Insights FromDocument22 pagesBiophysical Modeling: Induction of Clustered DNA Lesions by Ionizing Radiation - Insights FromrickNo ratings yet
- RochetteDocument9 pagesRochetterickNo ratings yet
- 1 The Length Scales of DNADocument7 pages1 The Length Scales of DNArickNo ratings yet
- Acta Dermato-Venereologica: Phototherapy With Narrowband Vs Broadband UVBDocument12 pagesActa Dermato-Venereologica: Phototherapy With Narrowband Vs Broadband UVBrickNo ratings yet
- UV-induced Skin Damage: M. Ichihashi, M. Ueda, A. Budiyanto, T. Bito, M. Oka, M. Fukunaga, K. Tsuru, T. HorikawaDocument19 pagesUV-induced Skin Damage: M. Ichihashi, M. Ueda, A. Budiyanto, T. Bito, M. Oka, M. Fukunaga, K. Tsuru, T. HorikawarickNo ratings yet
- Carcinogenesis 1997 Kvam 2379 84Document6 pagesCarcinogenesis 1997 Kvam 2379 84rickNo ratings yet
- Intercalating and Antitumour Activity of 4-Oxopyrimido (4, 5 :4,5) Thieno (2,3-b) Quinoline-4 (3H) - OneDocument8 pagesIntercalating and Antitumour Activity of 4-Oxopyrimido (4, 5 :4,5) Thieno (2,3-b) Quinoline-4 (3H) - OnerickNo ratings yet
- RBE of KV CBCT Radiation Determined by Monte Carlo DNA Damage SimulationsDocument13 pagesRBE of KV CBCT Radiation Determined by Monte Carlo DNA Damage SimulationsrickNo ratings yet
- A Role For Ultraviolet A in Solar: MutagenesisDocument5 pagesA Role For Ultraviolet A in Solar: MutagenesisrickNo ratings yet
- DIPLOIDnumbersDocument8 pagesDIPLOIDnumbersrickNo ratings yet
- Photochemical & Photobiological Sciences: PerspectiveDocument13 pagesPhotochemical & Photobiological Sciences: PerspectiverickNo ratings yet
- Setlowaction CPDDocument5 pagesSetlowaction CPDrickNo ratings yet
- Enhanced Photodynamic Effect of Cobalt (III) Dipyridophenazine Complex On Thyrotropin Receptor Expressing HEK293 CellsDocument22 pagesEnhanced Photodynamic Effect of Cobalt (III) Dipyridophenazine Complex On Thyrotropin Receptor Expressing HEK293 CellsrickNo ratings yet
- Sensors 13 07902Document14 pagesSensors 13 07902rickNo ratings yet
- Jacques Tissue OpticsDocument60 pagesJacques Tissue OpticsrickNo ratings yet
- PHPP 12009Document3 pagesPHPP 12009rickNo ratings yet
- Solar Simulation: Oriel Product TrainingDocument32 pagesSolar Simulation: Oriel Product TrainingrickNo ratings yet
- Sensors 13 07902Document14 pagesSensors 13 07902rickNo ratings yet
- Carcinogenesis 1997 Kvam 2379 84Document6 pagesCarcinogenesis 1997 Kvam 2379 84rickNo ratings yet
- Jacques Tissue OpticsDocument60 pagesJacques Tissue OpticsrickNo ratings yet
- PHPP 12009Document3 pagesPHPP 12009rickNo ratings yet
- Carcinogenesis 1997 Kvam 2379 84Document6 pagesCarcinogenesis 1997 Kvam 2379 84rickNo ratings yet
- Sunbeds Nine of Ten Over The LimitDocument7 pagesSunbeds Nine of Ten Over The LimitrickNo ratings yet
- GingsengDocument13 pagesGingsengrickNo ratings yet
- Visualization of Plasmid Delivery To Keratinocytes in Mouse and Human EpidermisDocument9 pagesVisualization of Plasmid Delivery To Keratinocytes in Mouse and Human EpidermisrickNo ratings yet
- Copula SpreadsDocument96 pagesCopula SpreadsrickNo ratings yet
- The Campanile (Vol 90, Ed 2), Published Oct 22, 2007Document24 pagesThe Campanile (Vol 90, Ed 2), Published Oct 22, 2007The CampanileNo ratings yet
- Manila Jocky Club Vs CADocument20 pagesManila Jocky Club Vs CAryusuki takahashiNo ratings yet
- Growth PredictionDocument101 pagesGrowth PredictionVartika TripathiNo ratings yet
- Using Games in English Language Learning Mrs Josephine Rama: Jurong Primary SchoolDocument16 pagesUsing Games in English Language Learning Mrs Josephine Rama: Jurong Primary SchoolChristine NguyễnNo ratings yet
- Applications of Digital - Image-Correlation Techniques To Experimental MechanicsDocument13 pagesApplications of Digital - Image-Correlation Techniques To Experimental MechanicsGustavo Felicio PerruciNo ratings yet
- Disseminated Tuberculosis in An AIDS/HIV-Infected Patient: AbstractDocument3 pagesDisseminated Tuberculosis in An AIDS/HIV-Infected Patient: AbstractAmelia Fitria DewiNo ratings yet
- Faith Confession SheetDocument5 pagesFaith Confession SheetArjel Jamias100% (1)
- University of Mumbai: Bachelor of Management Studies (Finance) Semester VIDocument73 pagesUniversity of Mumbai: Bachelor of Management Studies (Finance) Semester VIPranay ShettyNo ratings yet
- Types of Communicative StrategyDocument46 pagesTypes of Communicative StrategyMyra Bolinas100% (1)
- Simplified Analysis of 'Out, Out' - Robert FrostDocument3 pagesSimplified Analysis of 'Out, Out' - Robert FrostSANDREA RUTHNo ratings yet
- Quarter 4 Week 4 Parallel Lines Cut by A TransversalDocument26 pagesQuarter 4 Week 4 Parallel Lines Cut by A TransversalZaldy Roman MendozaNo ratings yet
- The Mind-Body ProblemDocument6 pagesThe Mind-Body ProblemCarlos Mendez PerezNo ratings yet
- Entrepreneurial Culture - Chapter 13Document11 pagesEntrepreneurial Culture - Chapter 13bnp guptaNo ratings yet
- The Creative Bureaucracy 2Document45 pagesThe Creative Bureaucracy 2IndirannaNo ratings yet
- Problems of Alternative Dispute Resolution Mechanisms and Proposals For Improvement: A Study in BangladeshDocument12 pagesProblems of Alternative Dispute Resolution Mechanisms and Proposals For Improvement: A Study in BangladeshssfsdsdNo ratings yet
- The Meaning of LifeDocument1 pageThe Meaning of LifeJayas SharmaNo ratings yet
- Unit 9Document3 pagesUnit 9LexNo ratings yet
- AP Macroeconomics: About The Advanced Placement Program (AP)Document2 pagesAP Macroeconomics: About The Advanced Placement Program (AP)Adam NowickiNo ratings yet
- Case Digest 1-4.46Document4 pagesCase Digest 1-4.46jobelle barcellanoNo ratings yet
- P4 Light NotesDocument6 pagesP4 Light NotesJohn John Appleseed100% (2)
- Moorish Architecture - KJDocument7 pagesMoorish Architecture - KJKairavi JaniNo ratings yet
- Full Download Test Bank For Microbiology The Human Experience First Edition First Edition PDF Full ChapterDocument36 pagesFull Download Test Bank For Microbiology The Human Experience First Edition First Edition PDF Full Chapterscalp.downcast.c7wgo100% (20)
- Key Influence Factors For Ocean Freight Forwarders Selecting Container Shipping Lines Using The Revised Dematel ApproachDocument12 pagesKey Influence Factors For Ocean Freight Forwarders Selecting Container Shipping Lines Using The Revised Dematel ApproachTanisha AgarwalNo ratings yet
- Case Presentation 1Document18 pagesCase Presentation 1api-390677852No ratings yet
- Spiderman EvolutionDocument14 pagesSpiderman EvolutionLuis IvanNo ratings yet
- Types of Drills PDFDocument8 pagesTypes of Drills PDFSummer nightsNo ratings yet
- MultidisciplinaryDocument20 pagesMultidisciplinaryrabiaNo ratings yet
- Application of Inventory Management in Construction IndustryDocument3 pagesApplication of Inventory Management in Construction IndustryEditor IJRITCCNo ratings yet
- A Minor Project Report Submitted To Rajiv Gandhi Prodhyogik Vishwavidyalaya, Bhopal Towards Partial Fulfillment For The Award of The Degree ofDocument53 pagesA Minor Project Report Submitted To Rajiv Gandhi Prodhyogik Vishwavidyalaya, Bhopal Towards Partial Fulfillment For The Award of The Degree ofwahidjaanNo ratings yet