Professional Documents
Culture Documents
RNA Synthesis, Processing SBT 211
Uploaded by
TADIWANASHE TINONETSANA0 ratings0% found this document useful (0 votes)
17 views59 pagesDNA is transcribed into RNA through the process of transcription. In eukaryotic cells, this involves RNA polymerases that recognize different promoter sequences. RNA polymerase binds to DNA and unwinds the double helix to copy one of the strands into RNA. The general steps of transcription are initiation, elongation, and termination. In prokaryotes, there is one RNA polymerase that produces mRNA, rRNA, and tRNA, while eukaryotes have three specialized RNA polymerases. Transcription termination can occur through rho-independent or rho-dependent mechanisms and results in the release of the RNA transcript.
Original Description:
Original Title
RNA Synthesis, Processing SBT 211.pptx
Copyright
© © All Rights Reserved
Available Formats
PDF, TXT or read online from Scribd
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentDNA is transcribed into RNA through the process of transcription. In eukaryotic cells, this involves RNA polymerases that recognize different promoter sequences. RNA polymerase binds to DNA and unwinds the double helix to copy one of the strands into RNA. The general steps of transcription are initiation, elongation, and termination. In prokaryotes, there is one RNA polymerase that produces mRNA, rRNA, and tRNA, while eukaryotes have three specialized RNA polymerases. Transcription termination can occur through rho-independent or rho-dependent mechanisms and results in the release of the RNA transcript.
Copyright:
© All Rights Reserved
Available Formats
Download as PDF, TXT or read online from Scribd
0 ratings0% found this document useful (0 votes)
17 views59 pagesRNA Synthesis, Processing SBT 211
Uploaded by
TADIWANASHE TINONETSANADNA is transcribed into RNA through the process of transcription. In eukaryotic cells, this involves RNA polymerases that recognize different promoter sequences. RNA polymerase binds to DNA and unwinds the double helix to copy one of the strands into RNA. The general steps of transcription are initiation, elongation, and termination. In prokaryotes, there is one RNA polymerase that produces mRNA, rRNA, and tRNA, while eukaryotes have three specialized RNA polymerases. Transcription termination can occur through rho-independent or rho-dependent mechanisms and results in the release of the RNA transcript.
Copyright:
© All Rights Reserved
Available Formats
Download as PDF, TXT or read online from Scribd
You are on page 1of 59
SBT 211
RNA Synthesis, Processing,
& Modification
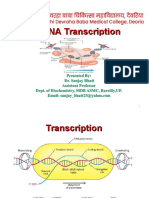
DNA transcription is a process that involves the
transcribing of genetic information from DNA to
RNA. The transcribed DNA message is used to
produce proteins.
The synthesis of an RNA molecule from DNA is a
complex process involving one of the group of RNA
polymerase enzymes and a number of associated
proteins.
The general steps required to synthesize the
primary transcript are initiation, elongation, and
termination.
The RNA molecules synthesized in mammalian
cells are made as precursor molecules that have to
be processed into mature, active RNA.
Types of RNA
All eukaryotic cells have four major classes of RNA:
ribosomal RNA (rRNA), messenger RNA (mRNA),
transfer RNA (tRNA), and small nuclear RNA
(snRNA). The first three are involved in protein
synthesis, and snRNA is involved in mRNA splicing.
Prokaryotic DNA dependent RNA
Polymerase (RNAP)
Bacterial cells have a single RNA polymerase that
transcribes DNA to generate all of the different
types of RNA (mRNA, rRNA, and tRNA).
Using DNA as a template, this enzyme polymerizes
ribonucleoside triphosphates (RNA nucleotides).
The complete RNA polymerase enzyme of E. coli
—the holoenzyme—is composed of a core enzyme
and a sigma factor.
An approximately 400 kDa core complex
consisting of two identical α subunits, similar but
not identical β and β′ subunits, and an ω subunit.
Beta is thought to be the catalytic subunit
RNAP, a metalloenzyme, also contains two zinc
molecules. The core RNA polymerase associates
with a specific protein factor (the sigma [σ] factor)
that helps the core enzyme recognize and bind to
the specific deoxynucleotide sequence of the
promoter region to form the preinitiation complex
(PIC)
Bacteria contain multiple σ factors, each of which
acts as a regulatory protein that modifies the
promoter recognition specificity of the RNA
polymerase. The major σ factor is σ70 , a
designation related to its molecular weight of
70,000 Daltons.
Following the initiation of transcription, the sigma
factor disassociates from the core enzyme.
Mammalian Cells Possess Three Distinct
Nuclear DNA-Dependent RNA Polymerases
In contrast to prokaryotes, eukaryotic cells have
three RNA polymerases.
◦ Polymerase I produces most of the rRNAs,
◦ polymerase II produces mRNA, and
◦ polymerase III produces small RNAs, such as tRNA and
5S rRNA.
All of these RNA polymerases have the same
mechanism of action. However, they recognize
different types of promoters.
They all have two large subunits and a number of
smaller subunits—as many as 14 in the case of
RNA pol III.
Actinomycin blocks the translocation of RNA
polymerase during transcription
Bacterial Transcription
Initiation
RNA polymerase (RNAP) binds to one of several
specificity factors, σ, to form a holoenzyme. In this
form, it can recognize and bind to specific promoter
regions in the DNA.
The DNA region that RNA polymerase associates with
immediately before beginning transcription is known as
the promoter.
The base pair where transcription initiates is called the
transcription-initiation site, or start site.
By convention, the transcription-initiation site in the
DNA sequence of a transcription unit is usually
numbered +1. Base pairs extending in the direction of
transcription (downstream) are assigned positive
numbers and those extending in the opposite direction
(upstream) are assigned negative numbers.
Promoters
RNA polymerase initiates transcription of most genes at
a unique position (a single base) in the template DNA
lying upstream of the coding sequence.
Surrounding a point in prokaryotic promoters about ten
nucleotides (-10) before the rst transcribed base is a
consensus sequence—TATAAT. This sequence is
known as a Pribnow box after one of its discoverers.
(A consensus sequence is the sequence most
commonly found in a given region when many genes
are examined.)
The nucleotides in the Pribnow box are mostly
adenines and thymines, so the region is primarily held
together by only two hydrogen bonds per base pair.
There is another region with similar sequences among
promoters centred near -35 and referred to as the -35
sequence. The consensus sequence at -35 is TTGTCA.
Promoter of the Escherichia coli ribosomal RNA gene, rrnB.
Note the -10 and -35 sequences and the upstream element.
The rst base transcribed (the transcriptional start site) is
noted (+1), as well as the upstream, downstream, and
transcription directions. (Data from W. Ross, et al., 1993.
Science 262:1407.)
The template (anticoding) strand of DNA is complementary
to both the coding strand and the transcribed RNA.
The sigma factor recognizes both the -35 and the
-10 sequences.
This initiation complex is initially referred to as a
closed complex because the DNA has not melted,
which is the next step in transcription initiation.
The DNA is unwound and becomes single-stranded
("open") in the vicinity of the initiation site (defined
as +1). This holoenzyme/unwound-DNA structure
is called the open complex.
After the transcription of 5–10 bases, the sigma
factor is released
About seventeen base pairs of DNA are opened,
and as transcription proceeds, about twelve bases
of RNA form a DNA-RNA duplex at the point of
transcription.
Transcription elongation
Transcription, like DNA replication, always
proceeds in the 5` to 3` direction. That is, a single
base is added de novo and then new RNA
nucleotides are added to the 3-OH free end, as in
DNA replication.
However, unlike DNA polymerase, prokaryotic RNA
polymerase does not seem to proofread as it
proceeds.
Most transcripts originate using (ATP) and, to a
lesser extent, (GTP) (purine nucleoside
triphosphates) at the +1 site. (UTP) and (CTP)
(pyrimidine nucleoside triphosphates) are
disfavoured at the initiation site.
Differences between replication and
transcription
(1) ribonucleotides are used in RNA synthesis
rather than deoxyribonucleotides;
(2) U replaces T as the complementary base pair
for A in RNA;
(3) a primer is not involved in RNA synthesis;
(4) only a very small portion of the genome is
transcribed or copied into RNA, whereas the entire
genome must be copied during DNA replication;
and
(5) there is no proofreading function during RNA
transcription.
RNA polymerase only goes one direction from a
promoter and only one strand of DNA is used as a
template at any one time.
To provide this template strand, the initiation of
transcription involves a short unwinding of the
DNA double helix. This is accomplished in a
two-step fashion.
First, RNA polymerase binds to the promoter to
form the closed complex, which is relatively weak.
Then, the double-stranded DNA goes through a
conformational change to form the much stronger
open complex through opening of the base pairs at
the -10 sequence, as shown next
Elongation is the function of the RNA polymerase
core enzyme. RNA polymerase moves along the
template, locally “unzipping” the DNA double helix.
This allows a transient base pairing between the
incoming nucleotide and newly-synthesized RNA
and the DNA template strand.
As it is made, the RNA transcript forms secondary
structure through intra-strand base pairing. The
average speed of transcription is about 40
nucleotides per second, much slower than DNA
polymerase.
Other protein factors may bind to polymerase and
alter the rate of transcription and some specific
sequences are transcribed more slowly than others
are. Eventually, RNA polymerase must come to the
end of the region to be transcribed.
A cistron is a region of DNA that encodes a single
polypeptide chain. In bacteria, mRNA is usually
generated from an operon as a polycistronic
transcript (one that contains the information to
produce a number of different proteins).
The polycistronic transcript is translated as it is
being transcribed. This transcript is not modified
and trimmed, and it does not contain introns
(regions within the coding sequence of a transcript
that are removed before translation occurs).
Several different proteins are produced during
translation of the polycistronic transcript, one from
each cistron.
Termination of transcription
The elongation reactions continue until the RNA
polymerase encounters a transcription termination
signal. One type of termination signal involves the
formation of a hairpin loop in the transcript,
preceding a number of U residues (rho
independent).
The second type of mechanism for termination
involves the binding of a protein, the rho factor,
which causes release of the RNA transcript from
the template (rho dependent).
The signal for both termination processes is the
sequence of bases in the newly synthesized RNA.
Terminators
Both types of terminators sequenced so far have
one thing in common: they include a sequence and
its inverted form separated by another short
sequence, all together forming an inverted-repeat
sequence.
Inverted repeats can form a stem-loop structure by
pairing complementary bases within the
transcribed messenger RNA.
both rho-dependent and rho-independent
terminators have the stem-loop structure in RNA
just before the last base transcribed.
rho-independent
terminators
Rho-independent terminators have a characteristic
structure, which features (a) A strong G-C rich stem
and loop, (b) a sequence of 4–6 U residues in the
RNA, which are transcribed from a corresponding
stretch of As in the template.
Uracil-adenine base pairs have two hydrogen bonds
and are thus less stable thermodynamically than
guanine-cytosine base pairs.
Perhaps, the uracil-adenine base pairs
spontaneously denature, releasing the transcribed
RNA and the RNA polymerase,
rho-dependent terminators
Rho-dependent terminators do not have the uracil
sequence after the stem-loop structure.
Termination depends on the action of rho, which
appears to bind to the newly forming RNA. In an
ATP-dependent process, rho binds at a rho
utilisation site on the nascent RNA strand and runs
along the mRNA at a speed comparable to the
transcription process itself.
Possibly, when RNA polymerase pauses at the
stem-loop structure, rho catches up to the
polymerase and unwinds the DNA-RNA hybrid,
letting the DNA, RNA, and polymerase fall free. Rho
can do this because it has DNA-RNA helicase
(unwinding) properties.
TRANSCRIPTION OF EUKARYOTIC
GENES
Eukaryotic transcription is more complex than
prokaryotic transcription.
transcription, is primarily localized to the nucleus,
where it is separated from the cytoplasm (in which
translation occurs)
This allows for the temporal regulation of gene
expression through the sequestration of the RNA in
the nucleus, and allows for selective transport of
RNAs to the cytoplasm, where the ribosomes reside
Eukaryotes—unlike prokaryotes, which have only
one RNA polymerase—have three RNA
polymerases.
Pre-Initiation
Promoter chromatin is transformed from a static to
a dynamic state upon gene activation.
Nucleosomes are rapidly removed and reassembled
in the activated state. In vivo, nucleosomes are
removed from promoter DNA for transcription.
Promoter DNA is made transiently available for
interaction with the RNA polymerase II (Pol II)
transcription machinery.
Histone acetyl transferases HAT acetylates the
lysine on histones thereby making histones less
negative which reduces the electrostatic affinity of
histones to DNA
Initiation
All three eukaryotic RNA polymerases (I, II, and III)
recognize a seven-base sequence, TATAAAA,
located at about -25 on the promoter DNA. It is
similar to the -10 sequence in prokaryotes and is
called the TATA box (or Hogness box).
The TATA box is bound by 34 kDa TATA binding
protein (TBP), which in turn binds several other
proteins called TBP-associated factors (TAFs). This
complex of TBP and TAFs is referred to as TFIID.
Binding of TFIID to the TATA box sequence is
thought to represent the first step in the formation
of the transcription complex on the promoter.
i. RNA Polymerase II and TFIIF bind to the
TFIID-TFIIB complex forming a minimal
transcription initiation complex.
ii. TFIIE and TFIIH bind to produce the
complete transcription initiation complex
or preinitiation complex (PIC).
Once the PIC is formed, the rate of
transcription can be enhanced or repressed
by other transcription factors.
Enhancing TFs called activators must bind
to DNA sequence elements called
enhancers.
Among the large number of promoters that have
been sequenced, a few lack the TATA box, yet are
still transcribed.
Transcription initiation in these promoters appears
to be controlled by a CT-rich area, called the
initiator element (Inr), at +1 of the transcript (close
to the transcription start site), coupled with a
downstream promoter element (DPE) at about +28
to +34 of the transcript.
In TATA-less promoters, TFIID requires both these
elements to bind. The initiator element has a
consensus sequence of TCA(G or T)T(T or C), and
the downstream promoter element has the
consensus sequence of (A or G)G(A or T)CGTG.
Sequences farther upstream from the start site
determine how frequently the transcription event
occurs.
Typical of these DNA elements are the GC and
CAAT boxes, so named because of the DNA
sequences involved.
Together, then, the promoter and
promoter-proximal cis-active upstream elements
confer fidelity and frequency of initiation upon a
gene.
A third class of sequence elements can either
increase or decrease the rate of transcription
initiation of eukaryotic genes.
These elements are called either enhancers or
repressors (or silencers), depending on which
effect they have.
Hormone response elements (for steroids, T3,
retinoic acid, peptides, etc) act as—or in
conjunction with—enhancers or silencers
Tissue-specific expression of genes (eg, the
albumin gene in liver, the haemoglobin gene in
reticulocytes) is also mediated by specific DNA
sequences.
Elongation
Transcription produces heterogeneous nuclear
mRNAs hnRNA from the DNA template which can
not leave the nucleus until processed.
hnRNA processing involves addition of a 5`-cap
and a poly(A) tail and splicing to join exons and
remove introns. The product, mRNA, then can
migrate to the cytoplasm, where it will direct
protein synthesis.
Cap may protect initial end of forming RNA strand
from endonuclease activity.
In absence of the poly-A tail, the RNA is rapidly
degraded.
Transcription and capping of
mRNA
At the 5 end of polymerase II transcripts, 7-methyl
guanosine is added in the “wrong” direction, 5` -
5`.
This cap allows the ribosome to recognize the
beginning of a messenger RNA.
The processing of hnRNA molecules is a site for
regulation of gene expression.
There are three different types of methyl caps,
shown in blue: CAP0 refers to the methylated
guanosine (on the nitrogen at the seven position,
N7) added in the 5` to 5` linkage to the mRNA;
CAP1 refers to CAP0 with the addition of a methyl
to the 22` carbon of ribose on the nucleotide (N1)
at the 5` end of the chain; and CAP2 refers to
CAP1 with the addition of another 2` methyl group
to the next nucleotide (N2).
The methyl groups are donated by
S-adenosylmethionine (SAM).
Addition of a Poly(a) tail
Enzymes cleave the transcript (hnRNA) at a point
10–20 nucleotides beyond an AAUAAA sequence,
forming the 3` end.
After this cleavage, a poly(A) tail that can be over
200 nucleotides in length is added to the 3`-end.
there is no corresponding poly(dT) sequence in the
DNA template that corresponds to this tail; it is
added post - transcriptionally.
ATP serves as the precursor for the sequential
addition of the adenine nucleotides. They are
added one at a time, with poly(A) polymerase
catalysing each addition.
Removal of Introns
Eukaryotic pre-mRNA transcripts contain regions
known as exons and introns.
Exons only, appear in the mature mRNA; introns
are removed from the transcript and are not found
in the mature mRNA.
The consensus sequences at the intron/exon
boundaries of the pre-mRNA are AGGU (AGGT in
the DNA).
The sequences vary to some extent on the exon
side of the boundaries, but almost all introns begin
with a 5` GU and end with a 3` AG
Because every 5` GU and 3` AG combination does
not result in a functional splice site, clearly other
features within the exon or intron help to define
the appropriate splice sites.
At least two types of splicing occur, although they
are related:
◦ self-splicing (via ribozymes)
◦ protein-mediated splicing.
During self-splicing, the U-A bond at the left (5`)
side of the intron is transferred to GTP. The U that
is now unbound displaces the G at the right (3`)
side of the intron, reconnecting the RNA with a U-U
connection and releasing the intron
Since all bonds are reversible transfers
(transesterications) rather than new bonds, no
external energy source is required. Self-splicing
introns of this type are called group I introns.
Self-splicing has also been found in genes in the
mitochondria of yeast.
These introns are referred to as group II introns
because they use a different mechanism of splicing
that does not require an external nucleotide.
Instead, the rst bond is transferred within the
intron to an adenosine, forming a lariat structure
In order for the lariat to form, the ribose of the
adenosine must make three phosphodiester bonds
Protein-Mediated Splicing (the
Spliceosome)
Nuclear ribonucleoproteins (snurps U1 to U6) bind to
the intron, causing it to form a loop. The complex is
called a splicesosome. The U1 snurp binds near the
first exon/intron junction, and U2 binds within the
intron in a region containing an adenine nucleotide
residue.
Another group of snurps, U4, U5, and U6, binds to the
complex, and the loop is formed. The phosphate
attached to the G residue at the 5`-end of the intron
forms a 2`–5` linkage with the 2`-hydroxyl group of
the adenine nucleotide residue.
Cleavage occurs at the end of the first exon, between
the AG residues at the 3’ end of the exon and the GU
residues at the 5` end of the intron. The complex
continues to be held in place by the spliceosome. A
second cleavage occurs at the 3`-end of the intron
after the AG sequence.
The exons are joined together. The intron, shaped like
a lariat, is released and degraded to nucleotides.
The splicing machinery for the majority of introns
also includes numerous other polypeptides called
auxiliary and splicing factors; the entire splicing
process requires about 50 polypeptides.
A second, less common, intron, called the
U12-dependent intron, with different consensus
sequences, also exists. It is removed by a similar
splicing process involving different snRNPs (U11,
U12) as well as many components shared with the
major spliceosome.
RNA editing
The term RNA editing describes those molecular
processes in which the information content in an
RNA molecule is altered through a chemical change
in the base makeup.
To date, such changes have been observed in
tRNA, rRNA, mRNA and microRNA molecules of
eukaryotes but not prokaryotes. RNA editing
occurs in the cell nucleus and cytosol, as well as in
mitochondria and plastids
The diversity of RNA editing phenomena includes
nucleoside modifications such as cytidine (C) to
uridine (U) and adenosine (A) to inosine (I)
deaminations, as well as non-templated nucleotide
additions and insertions
You might also like
- Rna and Amino AcidDocument14 pagesRna and Amino AcidIndestructible queenNo ratings yet
- Transcriprion ManuscriptDocument14 pagesTranscriprion ManuscriptAsif AhmedNo ratings yet
- Bala Sir Transcrption PDFDocument9 pagesBala Sir Transcrption PDFAnjali Ak GuptaNo ratings yet
- DNA to RNA: The Process of TranscriptionDocument10 pagesDNA to RNA: The Process of TranscriptionHardik ManekNo ratings yet
- TranscriptionDocument34 pagesTranscriptiondrhydrogenNo ratings yet
- Transcription: Dr. Anirudra Gurung, Department of Botany, St. Joseph's College, DarjeelingDocument9 pagesTranscription: Dr. Anirudra Gurung, Department of Botany, St. Joseph's College, DarjeelingRahul JaiswalNo ratings yet
- Pro TranscriptionDocument20 pagesPro TranscriptionnitralekhaNo ratings yet
- Biochemistry: RNA Synthesis and ProcessingDocument54 pagesBiochemistry: RNA Synthesis and ProcessingAqsa YaminNo ratings yet
- Transcription, Is The Process of Creating An EquivalentDocument7 pagesTranscription, Is The Process of Creating An EquivalentSandarbh GoswamiNo ratings yet
- SYBT TranscriptionDocument69 pagesSYBT TranscriptionMeir SabooNo ratings yet
- Transcription in ProkaryotesDocument18 pagesTranscription in ProkaryotesPrakash100% (1)
- Mrs. Ofelia Solano Saludar: Department of Natural Sciences University of St. La Salle Bacolod CityDocument55 pagesMrs. Ofelia Solano Saludar: Department of Natural Sciences University of St. La Salle Bacolod CitymskikiNo ratings yet
- Chapter 6 SummaryDocument28 pagesChapter 6 SummaryCharlotteNo ratings yet
- Molecular Biology: Unit 3Document64 pagesMolecular Biology: Unit 3John Rajan ThavamNo ratings yet
- G. Gene Expression and Protein SynthesisDocument50 pagesG. Gene Expression and Protein SynthesisMary Rose Bobis VicenteNo ratings yet
- TranscriptionDocument56 pagesTranscriptionVipin100% (8)
- 5 11transcription-2013Document30 pages5 11transcription-2013jernsssNo ratings yet
- Rna Biosynthesis (Transicription)Document33 pagesRna Biosynthesis (Transicription)Alaa AlmajedNo ratings yet
- Transcription: From DNA To RNADocument74 pagesTranscription: From DNA To RNAmd habibur rahmanNo ratings yet
- Transcription ProkaryoticDocument30 pagesTranscription ProkaryoticDibya Jyoti ParidaNo ratings yet
- Gift RNADocument13 pagesGift RNAPeter DindahNo ratings yet
- Chapter 5 Protein SynthesisDocument26 pagesChapter 5 Protein SynthesisT MokshithaNo ratings yet
- Transcription (Semister I MIC)Document117 pagesTranscription (Semister I MIC)Challagandla AnilNo ratings yet
- Dna L12 NotesDocument6 pagesDna L12 NotesellieNo ratings yet
- Transcription: in ProkaryotesDocument12 pagesTranscription: in ProkaryotesMonisha vNo ratings yet
- CENTRAL DOGMA OF MOLECULAR BIOLOGY - RNA TRANSCRIPTION AND TRANSLATIONDocument11 pagesCENTRAL DOGMA OF MOLECULAR BIOLOGY - RNA TRANSCRIPTION AND TRANSLATIONMaiSakurajimaNo ratings yet
- TranscriptionDocument23 pagesTranscriptionNikhil PularruNo ratings yet
- DNA Transcription ProcessDocument61 pagesDNA Transcription Processdeepak mauryaNo ratings yet
- Transcription ProcessDocument4 pagesTranscription ProcessArya ChaphekarNo ratings yet
- Transcriptio N: Submitted By: Yamini Saini Submitted To: Dr. Arshi Hussain Course: Msc. Chemistry - IDocument16 pagesTranscriptio N: Submitted By: Yamini Saini Submitted To: Dr. Arshi Hussain Course: Msc. Chemistry - IYamini SainiNo ratings yet
- The Key Steps of TranscriptionDocument13 pagesThe Key Steps of TranscriptionbiochemNo ratings yet
- Transcription and RNA ProcessingDocument38 pagesTranscription and RNA ProcessingRishi Kumar100% (1)
- DNA transcription - Regulation of gene expression through RNA polymeraseDocument38 pagesDNA transcription - Regulation of gene expression through RNA polymerasetoobashafiNo ratings yet
- TranscriptionDocument23 pagesTranscriptionareen fakhouryNo ratings yet
- Transcription WorkingDocument51 pagesTranscription Workingapi-3858544No ratings yet
- Transcription 2Document66 pagesTranscription 2Safe BoxNo ratings yet
- RNA - Polymerase 1 ClassDocument21 pagesRNA - Polymerase 1 ClassSeemaNo ratings yet
- Regulation of Gene Expression in Eukaryotes: Presented by Quratulain (19-Arid-1403) Asadullah (19-Arid-1379)Document32 pagesRegulation of Gene Expression in Eukaryotes: Presented by Quratulain (19-Arid-1403) Asadullah (19-Arid-1379)Beauty LiciousNo ratings yet
- Lesson 9Document66 pagesLesson 9Irica Mae CiervoNo ratings yet
- Transcription: M.Prasad Naidu MSC Medical Biochemisty, Ph.D.Research ScholarDocument33 pagesTranscription: M.Prasad Naidu MSC Medical Biochemisty, Ph.D.Research ScholarDr. M. Prasad NaiduNo ratings yet
- Prokaryotic TranscriptionDocument14 pagesProkaryotic TranscriptionAjay RawatNo ratings yet
- Chemistry Lec.22 (Nucleic Acids 6)Document6 pagesChemistry Lec.22 (Nucleic Acids 6)Muhammed AbdulsamadNo ratings yet
- From Gene To Protein,: Cellular and Sub-Cellular Protein ExpressionDocument25 pagesFrom Gene To Protein,: Cellular and Sub-Cellular Protein ExpressionPUBG HackerNo ratings yet
- TranscriptionDocument21 pagesTranscriptionYehya JaberNo ratings yet
- Transcription and Regulation of Gene Expression: By: Lyka Marie C. Falcasantos BSN - 1DDocument16 pagesTranscription and Regulation of Gene Expression: By: Lyka Marie C. Falcasantos BSN - 1DRemzAbdullaNo ratings yet
- Transcription: DNA-Directed RNA SynthesisDocument9 pagesTranscription: DNA-Directed RNA SynthesisnitralekhaNo ratings yet
- Kuliah 8. Transcription (Oke)Document43 pagesKuliah 8. Transcription (Oke)Sinta DosiNo ratings yet
- Molecular Biology: TranscriptionDocument101 pagesMolecular Biology: TranscriptionchintyaNo ratings yet
- Transcription: Gene Expression and Protein SynthesisDocument21 pagesTranscription: Gene Expression and Protein SynthesisAli Akand AsifNo ratings yet
- RNA ReplicationDocument23 pagesRNA ReplicationDharaneeshwari Siva-F&NNo ratings yet
- BCH 305 (Chemistry & Metabolism of Nucleic Acids)Document30 pagesBCH 305 (Chemistry & Metabolism of Nucleic Acids)ihebunnaogochukwu24No ratings yet
- Molecular Biology: TranscriptionDocument101 pagesMolecular Biology: TranscriptiontantyNo ratings yet
- Science FairDocument41 pagesScience FairSafanaNo ratings yet
- Gene ExpressionDocument19 pagesGene Expressionkvicto100% (1)
- DNA-Dependent Synthesis of RNA: (Transcription)Document10 pagesDNA-Dependent Synthesis of RNA: (Transcription)Shivam SharmaNo ratings yet
- Transcription in ProkaryotesDocument16 pagesTranscription in ProkaryotesAditya Kanwal100% (1)
- Paper TwoDocument21 pagesPaper TwoSafder AliNo ratings yet
- RNA Structure, Function, and Synthesis RNADocument6 pagesRNA Structure, Function, and Synthesis RNAMohini BajajNo ratings yet
- Transcription in Prokaryotes: Single RNA PolymeraseDocument29 pagesTranscription in Prokaryotes: Single RNA PolymeraseShubhamNo ratings yet
- Microbiology Practical Write Up 1Document5 pagesMicrobiology Practical Write Up 1TADIWANASHE TINONETSANANo ratings yet
- Microbiology Practical Write Up 3Document6 pagesMicrobiology Practical Write Up 3TADIWANASHE TINONETSANANo ratings yet
- Natpharm HR Manager Advert November 2021Document2 pagesNatpharm HR Manager Advert November 2021TADIWANASHE TINONETSANANo ratings yet
- Microbio Prac Write Up 2Document7 pagesMicrobio Prac Write Up 2TADIWANASHE TINONETSANANo ratings yet
- Microbio Prac 4Document8 pagesMicrobio Prac 4TADIWANASHE TINONETSANANo ratings yet
- Microbiology Practical Writeup 2Document8 pagesMicrobiology Practical Writeup 2TADIWANASHE TINONETSANANo ratings yet
- Cell Biology Practical Writeup 3Document9 pagesCell Biology Practical Writeup 3TADIWANASHE TINONETSANANo ratings yet
- Endosymbiotic Theories for Eukaryote OriginDocument18 pagesEndosymbiotic Theories for Eukaryote Originsoham deolankarNo ratings yet
- FungiDocument31 pagesFungiTADIWANASHE TINONETSANANo ratings yet
- Wks 1Document3 pagesWks 1TADIWANASHE TINONETSANANo ratings yet
- One-Way ANOVADocument18 pagesOne-Way ANOVATADIWANASHE TINONETSANANo ratings yet
- Bioprocess Optimization for High Density Endoinulinase Production from Recombinant Aspergillus NigerDocument4 pagesBioprocess Optimization for High Density Endoinulinase Production from Recombinant Aspergillus NigerTADIWANASHE TINONETSANANo ratings yet
- Bioseperation Practical 1Document10 pagesBioseperation Practical 1TADIWANASHE TINONETSANANo ratings yet
- Biochem Prac Write Up 3Document5 pagesBiochem Prac Write Up 3TADIWANASHE TINONETSANANo ratings yet
- DPPs BOOKLET-3 - Inductive Effects and Resonance in Organic ChemistryDocument21 pagesDPPs BOOKLET-3 - Inductive Effects and Resonance in Organic ChemistryJanuaryNo ratings yet
- Toward Industrial Catalysis of Zeolite For Linear Alkylbenzene Synthesis, A Mini ReviewDocument8 pagesToward Industrial Catalysis of Zeolite For Linear Alkylbenzene Synthesis, A Mini ReviewIrpan RomadonaNo ratings yet
- Microbial Control AgentsDocument4 pagesMicrobial Control AgentsJernyNo ratings yet
- Abdominal Fat Deposition in PoultryDocument12 pagesAbdominal Fat Deposition in PoultryCazim CrnkicNo ratings yet
- WARNING: This Document Is Part of Harmonized MS.50031: Flexible Vinyl Plastic For Injection Molding and ExtrusionDocument8 pagesWARNING: This Document Is Part of Harmonized MS.50031: Flexible Vinyl Plastic For Injection Molding and ExtrusionBonifácio Pacheco AmaralNo ratings yet
- BSC Chemistry 2023 24Document72 pagesBSC Chemistry 2023 24rudhrapriya33No ratings yet
- Carboxylic AcidDocument21 pagesCarboxylic AcidMuhammad AjmalNo ratings yet
- Gfps 6219 Brochure Ecofit All You Need in Pe enDocument12 pagesGfps 6219 Brochure Ecofit All You Need in Pe endfdfdNo ratings yet
- TT21 28112019BNN (E)Document40 pagesTT21 28112019BNN (E)Thanh Tâm TrầnNo ratings yet
- KAUL BOTANIQ Rate List for PCD Pharma Franchise on monopoly basisDocument7 pagesKAUL BOTANIQ Rate List for PCD Pharma Franchise on monopoly basisabhicwatNo ratings yet
- Direct Compression ExcipientsDocument10 pagesDirect Compression ExcipientsAzhar DkNo ratings yet
- Food Chemistry: Henrietta Ene-Obong, Nneola Onuoha, Lilian Aburime, Obioma MbahDocument7 pagesFood Chemistry: Henrietta Ene-Obong, Nneola Onuoha, Lilian Aburime, Obioma MbahadNo ratings yet
- Isra 2016Document18 pagesIsra 2016Portgas D. AceNo ratings yet
- Chemical Aspects of Bay of BengalDocument8 pagesChemical Aspects of Bay of BengalSamsun Naher MaishaNo ratings yet
- Chemistry: Investigatory Project FileDocument13 pagesChemistry: Investigatory Project FileSalil AltNo ratings yet
- 2023 H2 Chemical Equilibria Tutorial (QP)Document15 pages2023 H2 Chemical Equilibria Tutorial (QP)nivind88No ratings yet
- Material Safety Data Sheet: F-29, Liquid SanitizerDocument4 pagesMaterial Safety Data Sheet: F-29, Liquid SanitizerNiraNo ratings yet
- Catabolic Activities of Aerobic HeterotrophsDocument26 pagesCatabolic Activities of Aerobic HeterotrophsSh SarkerNo ratings yet
- Chapter 1Document50 pagesChapter 1Matthewzki Ferreras100% (1)
- Homologous Series WorksheetDocument8 pagesHomologous Series WorksheetSonaliNo ratings yet
- Foaming Capacity of Soaps (Ameen Rasool)Document18 pagesFoaming Capacity of Soaps (Ameen Rasool)Ameen RasoolNo ratings yet
- Cosmetics Lectures 1,2,3Document72 pagesCosmetics Lectures 1,2,3معلومة كيميائيةNo ratings yet
- Us 8653174Document21 pagesUs 8653174subramanian.sNo ratings yet
- A Rapid Protocol For Purification of Total RNADocument5 pagesA Rapid Protocol For Purification of Total RNAMS Clinic Anti aging & AestheticNo ratings yet
- Understanding Redox ReactionsDocument64 pagesUnderstanding Redox ReactionsJason Teh59% (66)
- Chemistry For Biologist 1A Parts 1& 2Document17 pagesChemistry For Biologist 1A Parts 1& 2body fayezNo ratings yet
- Leaching Manganese Nodules With Iron-Reducing Agents-A Critical ReviewDocument13 pagesLeaching Manganese Nodules With Iron-Reducing Agents-A Critical ReviewMartinNo ratings yet
- Superhydrophilicity and Strong Salt-Affinity Zwitterionic Polymer GraftedDocument18 pagesSuperhydrophilicity and Strong Salt-Affinity Zwitterionic Polymer GraftedIoana-Alexandra TrofinNo ratings yet
- Results and Discussion of Lipid Solubility, Identification, and AnalysisDocument5 pagesResults and Discussion of Lipid Solubility, Identification, and AnalysisStarrrNo ratings yet
- Waste Management: Thomas H. Christensen, Valentina BisinellaDocument17 pagesWaste Management: Thomas H. Christensen, Valentina BisinellaclaudiacarranzafNo ratings yet