Professional Documents
Culture Documents
Dissecting Complexity The Hidden Impact of Applica
Uploaded by
NealOriginal Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
Dissecting Complexity The Hidden Impact of Applica
Uploaded by
NealCopyright:
Available Formats
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022.
The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
Dissecting Complexity: The Hidden Impact of Application
Parameters on Bioinformatics Research
Mikaela Cashman1* , Myra B. Cohen2 , Alexis L. Marsh2 , Robert W. Cottingham1
1 Biosciences Division, Oak Ridge National Laboratory, Oak Ridge, TN, USA
2 Dept. of Computer Science, Iowa State University, Ames, IA, USA
* cashmanmm@ornl.gov
Abstract
Biology is a quest; an ongoing inquiry about the nature of life. How do the different
forms of life interact? What makes up an ecosystem? How does a tiny bacterium work?
To answer these questions biologists turn increasingly to sophisticated computational
tools. Many of these tools are highly configurable, allowing customization in support of
a wide range of uses. For example, algorithms can be tuned for precision, efficiency,
type of inquiry, or for specific categories of organisms or their component subsystems.
Ideally, configurability provides useful flexibility. However, the complex landscape of
configurability may be fraught with pitfalls. This paper examines that landscape in
bioinformatics tools. We propose a methodology, SOMATA, to facilitate systematic
exploration of the vast choice of configuration options, and apply it to three different
tools on a range of scientific inquires. We further argue that the tools themselves are
complex ecosystems. If biologists explore these, ask questions, and experiment just as
they do with their biological counterparts, they will benefit by both finding improved
solutions to their problems as well as increasing repeatability and transparency. We end
with a call to the community for an increase in shared responsibility and communication
between tool developers and the biologists that use them in the context of complex
system decomposition.
Introduction 1
Scientific investigation in biology has become increasingly dependent on computational 2
discovery. Modern research workflows rely on the use of various evolving bioinformatics 3
software applications which has led to a sophisticated computational ecosystem that 4
accommodates increasingly diverse and integrated data sets along with their associated 5
experimental probes, as well as comprehensive conceptual models of scientific 6
understanding. As an example, consider an investigation of the mechanisms of 7
promoting growth related to plant and microbe interactions as described in Finkel et 8
al. [1]. They found a trait — such as the inhibition of root growth — can be 9
significantly reversed by interaction with a single operon found in the bacteria 10
This manuscript has been co-authored by UT-Battelle, LLC under Contract No. DE-AC05-
00OR22725 with the U.S. Department of Energy. The United States Government retains and the
publisher, by accepting the article for publication, acknowledges that the United States Government
retains a non-exclusive, paid-up, irrevocable, worldwide license to publish or reproduce the published form
of this manuscript, or allow others to do so, for United States Government purposes. The Department
of Energy will provide public access to these results of federally sponsored research in accordance with
the DOE Public Access Plan (https://energy.gov/downloads/doe-public-access-plan).
December 20, 2022 1/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
Variovorax. The fact that Variovorax is involved is not surprising as it had previously 11
been observed, but that such an important function is provided in a relatively simple 12
way within a complex biological system, by a single operon, is unexpected. This 13
investigation involved the use of multiple software tools to support data driven discovery 14
and model based hypothesis generation [2]. 15
A conceptual approach to reproducing their workflow might start with microbiome 16
sequencing (performed in the wet lab), recovering metagenome assembled genomes or 17
MAGs (using an assembly computational tool), annotation of the MAGs (using an 18
annotation computational tool that performs alignment), followed by a taxonomic 19
classification (using another computational tool), and followed by an analysis of 20
associated biochemical pathways [3] (using a tool such as a flux balance analysis). Thus, 21
this novel result required four different computational tools. As we demonstrate later, 22
each of these tools may be considered almost as complex as the biological subsystems 23
being explored. 24
The above example illustrates a single software workflow. Different research 25
questions will require different workflows with different algorithmic parameters. This 26
has led to the inclusion of many different parameters (or configuration options) in 27
bioinformatics tools. For instance, BLAST (Basic Local Alignment Search Tool ) [4], one 28
of the most commonly used tools to study DNA and protein sequences, has 29
approximately 50 parameters a user can modify and customize. Changing these can 30
impact core functionality such as increasing the string match length, or modifying the 31
output format. Users can simply leave the default settings, or modify a few settings 32
based on their laboratory norms, but this practice may result in the full power of these 33
tools not being leveraged with less than optimal results or missed observations. 34
However, modifying parameters without a full understanding of their purpose can 35
also be problematic. For example, in a letter to the editor of Bioinformatics by Shah et 36
al. [5], they discuss a misunderstanding of a commonly used BLAST parameter 37
(max target seqs) by another researcher. BLAST documentation states the following 38
definition of max target seqs [6]: 39
Number of aligned sequences to keep. Use with report formats that do not 40
have separate definition line and alignment sections such as tabular (all 41
outfmt > 4). Not compatible with num descriptions or num alignments. 42
Ties are broken by order of sequences in the database. 43
Instead of this parameter filtering at the end as this may be commonly interpreted, 44
it actually filters results during the algorithm’s iterative search. Thus it may not be 45
clear to a user how it impacts their results. The authors of the referenced paper stated 46
their explicit reason for limiting max target seqs to a value of 1, under the assumption 47
it filtered out only the best (top) result at the end of the BLAST search. However, they 48
misunderstood the parameter’s purpose. In a response from the NCBI team [7], they 49
acknowledge that Shah discovered a bug (which they fixed) and that they have updated 50
their documentation. 51
In addition, several studies have examined how users interact with common 52
bioinformatics tools. Morrison-Smith et al. noted that users are unsure of how to select 53
the correct parameters and have to trust a tool’s correctness without real confidence [8]. 54
For example participants have said “At the end of the day, you just have faith that the 55
program’s working or not.” Others have examined how changing parameters impacts 56
system performance [9] or correctness [10], however, there is a lack of research on how to 57
train scientists to efficiently leverage the potential of parameters. We argue that if 58
scientific software is viewed as a magic box, the community loses scientific potential. 59
Instead, the bioscience user community needs to peel away the covers and develop 60
techniques that explore the impact these parameters have on their results, managing 61
December 20, 2022 2/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
them using a controlled and scientific approach. Reading documentation alone may not 62
be sufficient. 63
Without a set of systematic techniques, it can be difficult to manage this software 64
complexity. Since scientists use these tools to help form the basis of their conclusions, a 65
lack of understanding of how parameters change the outcome of the results is both a 66
missed opportunity and potentially problematic. In fact, it has already led to 67
publication retractions [11, 12]. Biologists, hence, often develop a laboratory-specific 68
routine protocol of what to change and leave the rest of the options alone. In this article 69
we present a systematic and exploratory protocol (SOMATA) to work with scientific 70
software. We also argue for more interaction between tool developers and end-users to 71
build a shared body of knowledge. This will aid the advancement of computational 72
methods to match the rigor that already exists in the documentation of lab work. 73
In the following sections, we first present an example and approach for working with 74
bioinformatics software that matches the rigor and practice that already exists in the 75
documentation and execution of laboratory work. Then we discuss background on 76
software configurability and the bioinformatics tools used in this work. Next, we 77
propose a systematic protocol (SOMATA) for breaking down the complexity of software 78
tools. We then apply this protocol in a series of experimental studies with one study 79
focused on empirically measuring the effect of configurations, and a second study 80
demonstrating an approach to managing configurability. Finally, we end with a 81
discussion mapping out three key takeaways. 82
Systems View - A Motivating Example 83
A systems view in the biological context is a useful approach to study complex and 84
potentially interacting systems. In this paper we take a systems view of software itself. 85
Instead of using software as a single tool, we demonstrate how users can approach 86
software with the same mindset as the organisms they study; as a complex system that 87
can be optimized based on one’s environment and goals. We demonstrate this idea with 88
an example showing how parameters can change how a researcher perceives and 89
interacts with bioinformatics tools while conducting a common experimental scenario. 90
In this scenario we are trying to understand how different chemical compounds in a 91
growth media change the metabolic pathways utilized in Escherichia coli (E. coli ). For 92
our growth medium we use the minimal media formulation Carbon-D-Glucose. 93
Before running extensive laboratory experiments we might want to exploit a common 94
computational method simulating an organism’s growth using a genome-scale metabolic 95
network [13]. We can run a Flux Balance Analysis (FBA) which calculates the flow of 96
metabolites through the metabolic network in a specific growth media to optimize for 97
growth (measured by flux through the biomass reaction) using a linear programming 98
algorithm. The FBA tool used here (further discussed in experimental study 99
methodology) provides input options such as the FBA model and the media, as well as 100
algorithmic options such as the reaction to maximize, custom flux bounds, expression 101
thresholds, and maximum uptakes of different nutrient sources. One output of this tool 102
is the objective value (OV) which represents the maximum flow through the biomass 103
reaction as a measurement of growth [14]. If we do not change any of the settings (i.e. 104
run the default configuration) E. coli produces an OV of 0.164692 mmol/g CDW hr. 105
For the purpose of this example we will explore the effect of the max oxygen 106
uptake option which is described in the documentation as the “maximum number of 107
moles of oxygen permitted for uptake (default uptake rates varies from 0 to 100 for all 108
nutrients)”. This option has no fixed default value since the maximum is constrained by 109
the media input. We will change this to a value of 0 to see what the behavior is. Under 110
this new configuration, E. coli no longer grows (OV of 0 is returned). Next we increase 111
December 20, 2022 3/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
this value in increments of 10 to observe how the OV changes. Results for values from 0 112
to 100 can be seen in Table 1. We see the OV increases linearly until it reaches the 113
maximum (also the default growth) of 0.164692 when max oxygen is set to 40. If we set 114
this value past 40, there is no added effect. We will explore why this is in a later study.
Table 1. Flux Balance Analysis (FBA) in E. coli in a Carbon-D-Glucose media.
Objective Value (OV) is reported under various values of the max oxygen parameter.
Parameter OV
Default 0.164692
Max O 0 0
Max O 10 0.0536659
Max O 20 0.107332
Max O 30 0.160998
Max O 40 0.164692
Max O 50 0.164692
Max O 60 0.164692
Max O 70 0.164692
Max O 80 0.164692
Max O 90 0.164692
Max O 100 0.164692
115
This example demonstrates changes in the measured result (OV), but there are also 116
changes occurring to the internal model of E. coli — specifically — the flows or fluxes 117
of each reaction in the metabolic network. The resulting flux can be positive meaning 118
the net flow of metabolites occurs from left to right in the reaction equation, negative 119
meaning the net flow occurs right to left, or zero indicating the flux is equal on both 120
sides resulting in a net of zero. For more information on FBA within this tool please 121
refer to [14]. The metabolic model used in this example has a total of 1,591 reactions. 122
In modifying the level of oxygen, for example changing from the default to a value of 10, 123
431 reactions (27.09%) result in different fluxes. Most reactions either stayed the same 124
(72.91%), or increased or decreased in flux but their direction stayed the same (26.34%). 125
However 12 reactions had a significant change as seen in Table 2. Five reactions change 126
from a negative flux to a flux of zero, three from zero to negative, two zero to positive, 127
and two changed from positive to zero. These 12 reactions represent significant changes 128
that occur to metabolism of E. coli when oxygen levels are varied by means of the tool 129
parameter. 130
We can observe these changes to the reactions and corresponding pathways directly 131
in the KEGG pathway maps as seen in Fig 1 which depicts Pyruvate Metabolism. The 132
reaction through EC 1.1.5.4 changes from a negative net flux (left) to a net flux of zero 133
(right). We also see reactions change in the magnitude of their flux. For example ECs 134
2.3.1.54, 2.7.1.40, and 2.7.2.1 change from a higher net negative flux (left) to a lower net 135
flux (right). EC 2.3.1.8 changes from a higher net positive flux (left) to a lower net flux 136
(right). 137
These configuration changes can be considered as a “state change” to the system. 138
Consider the state diagram in Fig 2. In the initial state (before running FBA) we have 139
a static model with no defined fluxs through its reactions. After running the default 140
configuration through, we arrive at a state with a growth of 0.164692 with fluxes 141
through its 1,591 reactions. But if we take a different transition, for example through 142
max oxygen of 10 then we arrive a different state with a growth of 0.0536659 where the 143
flux through 431 reactions differ leading to changes in the pathways. The same occurs if 144
we set max oxygen to 30 with a growth of 0.160998 resulting again in different reactions 145
and paths. Hence, the configuration has a direct impact on the individual reaction fluxes 146
December 20, 2022 4/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
Table 2. Change in individual reaction fluxes after setting the max oxygen parameter to a value of 10 versus the default of
unset.
KBase ID KEGG ID/ Effect on Associated Pathways
EC Number Flux
rn00230 Purine metabolism
R00722
rxn00515 c0 neg to zero rn01100 Metabolic pathways
2.7.4.6
rn01232 Nucleotide metabolism
R00848 rn00564 Glycerophospholipid metabolism
rxn00616 c0 neg to zero
1.1.99.5 rn01110 Biosynthesis of secondary metabolites
R00970
rxn00715 c0 neg to zero rn00240 Pyrimidine metabolism
2.7.1.48
R01257
rxn00935 c0 neg to zero rn00620 Pyruvate metabolism
1.1.99.16
rn00680 Methane metabolism
R06983 rn01100 Metabolic pathways
rxn04792 c0 neg to zero
1.1.1.284 rn01120 Microbial metabolism in diverse environments
rn01200 Carbon metabolism
R00842 rn00564 Glycerophospholipid metabolism
rxn00611 c0 zero to neg
1.1.1.8, 1.1.1.94, 1.1.1.261 rn01110 Biosynthesis of secondary metabolites
rn00240 Pyrimidine metabolism
R02326
rxn01673 c0 zero to neg rn01100 Metabolic pathways
2.7.4.6
rn01232 Nucleotide metabolism
rn00330 Arginine and proline metabolism
R10507 rn00332 Carbapenem biosynthesis
rxn09188 c0 zero to neg
1.5.99.8 rn01100 Metabolic pathways
rn01110 Biosynthesis of secondary metabolites
rn00330 Arginine and proline metabolism
R01251 rn01100 Metabolic pathways
rxn00931 c0 zero to pos
1.5.1.2 rn01110 Biosynthesis of secondary metabolites
rn01230 Biosynthesis of amino acids
ec00630 Glyoxylate and dicarboxylate metabolism
ec01100 Metabolic pathways
rxn08657 c0 1.1.3.15 zero to pos
ec01110 Biosynthesis of secondary metabolites
ec01120 Microbial metabolism in diverse environments
rn00480 Glutathione metabolism
R07140 rn00680 Methane metabolism
rxn04938 c0 pos to zero
1.1.1.284 rn01100 Metabolic pathways
rn01120 Microbial metabolism in diverse environments
ec00630 Glyoxylate and dicarboxylate metabolism
ec01100 Metabolic pathways
rxn08656 c0 1.1.3.15 pos to zero
ec01110 Biosynthesis of secondary metabolites
ec01120 Microbial metabolism in diverse environments
and the pathways through the network which leads to different behavior (e.g. growth). 147
By approaching this software tool with a systems view, we have demonstrated that 148
not only can the ultimate objective change based on the parameters used (e.g. objective 149
value), but the internal biological state (the direction and magnitude of the internal 150
metabolic reactions) can change in significant ways as well. In the rest of this paper we 151
demonstrate via case studies how using a systems mindset allows the scientist to explore 152
and understand dependencies between input parameters and the final scientific result. 153
This improved understanding can build confidence and lead to better science. 154
December 20, 2022 5/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
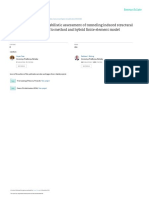
Fig 1. KEGG map for Pyruvate Metabolism under the default configuration (left) and with max oxygen set to 10 (right).
Green represents a positive net flux, red is a negative net flux, and blue means the enzyme is present but the net flux is zero.
The shade of the color represents the magnitude of the flux (a darker color represents a larger value).
n =1
0 MaxOxygen 10
yge
ax0x
M
• OV = 0.0536659
Default • 431 Reaction
Init fluxes change
• KEGG map
• OV = 0.164692
• Media
Default • 1,591
• Gapfilled
Reactions
Model
• KEGG map
MaxOxygen 30
• OV = 0.160998
• 431 Reaction
fluxes change
M • KEEG map
ax
0x
yg
en
=3
0
Fig 2. State change of FBA under different conditions.
Next we present some background on configurability to lay the groundwork for our 155
case studies. 156
December 20, 2022 6/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
Fig 3. The Nucleotide BLAST (BLASTn) Graphical User Interface (GUI) environment from NCBI. The left shows the
default input options, and the right shows the additional algorithmic parameters [18].
Background 157
Software Configurability 158
Many software tools are designed with flexibility to satisfy a variety of use cases by 159
supporting different facets of a user task. For instance, tools may handle different input 160
data formats, various data sizes, and algorithms can be selected or customized 161
depending on the user’s goal. A common example of configurability is a web browser, 162
such as Firefox or Chrome. A user can select menu options that change the security 163
settings, or configure how windows and tabs open and close. Some users may want 164
explicit warnings before exiting a window or tab, and others may prefer to close 165
windows without confirmation. Which levels of security are chosen or the ability to 166
embed JavaScript (or not) are all options that can be customized. Current versions of 167
Firefox have more than 2,000 different options a user can manipulate [15, 16]. 168
Configurability benefits developers by allowing them to create software for a broad 169
audience through building on reusable components, of which the cost of development 170
can be amortized over the full software development lifecycle [17]. It also has been 171
demonstrated to lead to higher quality of individual components since these are tested 172
and used in multiple systems [17]. This type of development stands in contrast to 173
building many unique specialized programs, each with individual code, and only slight 174
variations in behaviors. From a user’s perspective, this would require them to switch 175
between different software tools as their goals change. 176
A configuration of a software system is the set of choices, one for each of the system 177
parameters (or configuration options). For instance, in Firefox a configuration would be 178
defined by the set of choices for all 2,000+ options. Most will have a default value that 179
is assigned when the application starts, hence the user rarely changes more than a few 180
of these on their own; in theory any combination of parameter options can be used 181
together. Hence, the complete configuration space of a configurable system is the set of 182
all possible unique sets of parameter values (or the Cartesian product of the options for 183
the individual values). In the simplest case, where all options are binary (turn the 184
option on/off), the full configuration space would be 2|p| where |p| represents the 185
number of parameters of the system. Returning to the Firefox example, assuming 2,000 186
December 20, 2022 7/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
parameters, this would be 22000 . 187
By design, changing parameters modifies how an application behaves. The possible 188
space of behavior of a configurable software system is usually infeasible to exhaustively 189
enumerate [10, 15, 19]. The range of system behaviors can lead to unexpected failures 190
(both in outcome and performance) when interactions occur between configuration 191
options. For instance, one option might reduce the amount of memory for caching and 192
another might increase some data that is being cached. When the new data is cached it 193
might overflow the internal structures for caching which have now been reduced in size. 194
There has been a large body of research that has identified efficient ways to sample a 195
configurable software system’s configuration space and discover a large portion of the 196
possible behavior [15, 20, 21]. We do not attempt to survey that literature here. 197
In this paper we focus on configurable software in the bioinformatics domain. We 198
consider a software parameter (herein shortened to parameter ) to be an option that can 199
be specified by the user and then used by the program or application when it is running. 200
As noted previously, users have highlighted difficulties understanding parameters, and 201
this has even led to retracted articles [11, 12]. For a concrete biological example of a 202
parameter, consider the graphical user interface of NCBI’s BLASTn implementation 203
(Figure 3). This interface has more than 15 parameters for a user to adjust. In fact, 204
BLASTn actually has over 50 parameters that can be modified thus not all are exposed 205
in the NCBI GUI. Some of the parameters are in the form of a drop-down menu or a set 206
of options such as Max target sequences and gap costs. Others are check boxes or 207
binary parameters such as short queries and Mask lower case letters. Others are 208
open text fields such as Expect threshold which can be set to a numeric value. Any 209
parameter that is not explicitly defined will use the default value as determined by the 210
program. While many laboratories have their own custom set of options they regularly 211
choose, there is no one best configuration that is best for all research questions, hence, 212
understanding ways to navigate and explore these, is an important part of scientific 213
discovery. 214
(1) Assembly
(2) Alignment
OrgA OrgB OrgA
GeneA GeneB GeneC
(3) Annotation
(4) Metabolic Flux Balance
Modeling Analysis
growth @ 0.165 mol/CDG /hr
Fig 4. Methods overview of a common bioinformatics workflow.
December 20, 2022 8/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
Bioinformatics Methods 215
A common bioinformatics workflow, as shown in Figure 4, starts with read files output 216
from a DNA sequencer that are then assembled (1) into reconstructions of the 217
underlying original source genome DNA sequence. These are then aligned (2) against 218
known DNA sequences to find regions of similarity with known genes and their 219
functions. This provides a basis for identifying genes and annotation (3) of the original 220
source genomes. The gene annotations then provide the basis for establishing 221
genome-scale metabolic models (4) and doing flux balance analysis. 222
In this work we consider three different exemplars of bioinformatics techniques 223
commonly used in the workflow described: genome assembly, genome alignment, and 224
metabolic modeling. The following subsections provide brief background on these 225
techniques including a description of the biological functional objective of each. The 226
term functional objective is used here referring to the biological output of the methods 227
relative to the user’s biological query. 228
DNA Assembly 229
The read file output from a DNA sequencer contains a set of short sequences of 230
base-pairs. Each of the short sequences is called a read, and are each an exact copy of a 231
short segment of genomic DNA. Depending on the sequencing technology and methods, 232
each read ideally overlaps with many others. Assembly is the computational method to 233
align and recombine reads to reconstruct the original genome sequence. When this is 234
not fully accomplished due to sequencing error or lack of sufficient coverage, and 235
especially with metagenomic samples containing genomes of many different species, the 236
result is a set of assembled contigs, longer contiguous sequences reflecting the 237
underlying genomes with hopefully high fidelity. A common functional objective of 238
DNA assembly is to obtain a few long contigs, ideally a complete one, per species. For 239
metagenomics there can be a trade off between completeness of individual species 240
genomes and a more comprehensive representation of most species in the sample [22, 23]. 241
Here we will focus on and optimize for the first. 242
DNA Alignment 243
Given a newly discovered or unknown DNA sequence, DNA Alignment (or Sequence 244
Alignment) is a procedure for finding the most similar or homologous sequence in a 245
database of known sequences. The unknown sequence is often referred to as the query 246
sequence. The alignment algorithm compares the query with all other sequences in the 247
database, and calculates the significance of each match. We investigate a common use 248
case, searching for the most similar known sequences in order to infer the likely 249
evolutionary source or biological function of the unknown query sequence. The matches 250
or hits found in the database to the query sequence are returned with a variety of 251
quality scores that indicate how confident the alignment is with respect to each possible 252
match. Two common metrics are the percentage identity (how identical two sequences 253
are) and the expected value (or e-value) which is a measure of how likely the hits would 254
be found in the database by chance. The functional objective of DNA alignment can 255
vary by use case. We define one such objective that covers a range of use cases as the 256
“number of quality hits” where the exact definition of quality may vary by use case, but 257
typically involves a combination of several of the quality metrics such as percentage ID 258
and e-value. 259
December 20, 2022 9/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
Metabolic Modeling 260
Metabolic modeling aims to reconstruct and simulate metabolic pathways of an 261
organism to study its metabolism and gain further insight into its cellular biology. A 262
metabolic network is a graphical representation of the set of chemical reactions where 263
nodes represent chemical compounds and edges represent chemical reactions between 264
those compounds. A path in these networks is a biological pathway. For example, the 265
Kyoto Encyclopedia of Genes and Genomes (KEGG) database contains numerous 266
metabolic networks for various model organisms [24]. One common type of analysis 267
performed on metabolic networks is to estimate the flow of metabolites through the 268
network under specific environmental conditions. A popular type of algorithm to 269
perform this is known as flux balance analysis (FBA) [13]. FBA optimizes reaction 270
fluxes (flows) through an organism’s metabolic reaction network to predict the 271
organism’s growth or other objective function. As one example of a functional objective, 272
a user might provide an FBA algorithm one environmental condition, and want to 273
observe how much the organism grows. This can then be assessed by an objective value 274
such as a measurement of biomass. Other examples of use cases include observing how 275
pathways change under different environmental conditions, identifying what 276
environmental conditions are needed for organism growth, and identifying how much an 277
organism grows in a specific environment. The functional objective of FBA varies by 278
use case. Herein we follow one of the most common; to determine the optimum flux over 279
the metabolic network that maximizes growth. 280
Experimental Protocol: SOMATA 281
We propose a process SOMATA, a core methodology to analyze any complex software 282
systems and systematically break it down. SOMATA involves Selecting tools and data, 283
identifying Objective metrics, Modeling the parameter space, choosing a sample design 284
Approach, Testing, and Analyzing. SOMATA includes techniques from the field of 285
software testing as its foundation. The details of each step are as follows: 286
1. Select tool and data for exploration 287
• The first step is to select the tool or software of interest. In our motivating 288
example the tool of interest was an implementation of Flux Balance Analysis, 289
and the data was a metabolic model of E. coli along with a compatible 290
definition of the Carbon-D-Glucose media. 291
2. Define an outcome metric based on user Objective 292
• The metric to evaluate is an effect or output of the tool. The metric should 293
be related to the user’s objective of interest. This could be a functional 294
output such as a measurement associated with growth, as in our motivating 295
example, or a non-functional output such as runtime. Several metrics may be 296
selected depending on what is important in a particular use case. In our 297
example the objective metric was the OV (a measurement of growth). 298
3. Model the parameter space of the tool 299
• This involves identifying possible parameters of interest, defining their valid 300
choices and range of options, and determining default values. Parameters can 301
be environmental or functional parameters that have a known direct effect on 302
the biology of the system, or they can be algorithmic parameters that are 303
known to impact the underlying algorithmic method. In the example, a 304
December 20, 2022 10/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
single biological parameter — MaxO — was varied since that is intuitively 305
related to growth, the output objective of interest. 306
4. Choose sample design Approach 307
• Next, choose from the constructed model in Step 3 which options to 308
experimentally evaluate. A discrete range or set of options for each 309
parameter must be chosen. Furthermore, the user must decide if they want 310
to test the parameters individually, in combination, or some mixture of the 311
two. In the example, we chose 11 choices for the parameter (values of MaxO 312
between 0 and 100 in steps of 10) and explored these individually. 313
5. Execute the experimental Test evaluation 314
• Given the chosen model and sample design approach, run the selected 315
parameters against the tool with respect to the selected data. Often this 316
process can be automated to allow for large search spaces. Our example runs 317
an FBA analysis for each of the 11 parameter choices. 318
6. Observe results and Analyze effects 319
• Once complete, observe the effect of changing the model parameters given by 320
Step 3, and conduct deeper analysis if desired. Given the results, to gain 321
further insight, it may be beneficial to return to a previous step and alter the 322
sections. For our example we examine the differences in OVs of growth and 323
the impact on various associated pathways. 324
We want to emphasize the highly customizable nature of this process. For instance, 325
in Step 4 there are several sampling design approaches that can be used. We can use 326
techniques from software testing and design of experiments, or simply use a random 327
method of investigation. In Step 5, simulation can replace experimentation. Finally, in 328
Step 6 we could add in additional methods of analysis such as tools from machine 329
learning. We follow this methodology in our Experimental Evaluation. 330
This process does not in itself contain any unprecedented elements individually. 331
However as we argue in the Introduction, the bioinformatics and biology community 332
would benefit from a clear approach to dissecting the complex software systems on 333
which their research and science rely. Therefore, we present this as a formal 334
methodology along with several experimental demonstrations of its application, and the 335
real scientific impact that follow from the examples. 336
Experimental Evaluation 337
In this section we expand on the scenarios in our motivating example, and ask how 338
much of an impact changing parameters have on scientific results. For this we measure 339
the behavioral impact of modifying parameters in several commonly used bioinformatics 340
tools, which can lead to different scientific conclusions. We then ask if a systematic 341
investigation of parameter settings can lead to any insights that can assist a user in 342
navigating the parameter space. This would allow scientists to infer information to help 343
them navigate the parameter space. We formally ask the following research questions: 344
• Experiment 1: What is the impact on a tool’s behavior when 345
changing parameters? 346
• Experiment 2: Can we use an experimental approach to explore the 347
parameter space and infer the internal behavior of the software 348
system? 349
December 20, 2022 11/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
We further split up Experiment 2 into two studies (Experiment 2a and Experiement 2b). 350
We followup our study by sharing several lessons learned during this work that can lead 351
to a set of best practices for users of highly configurable scientific software tools. Please 352
refer to our supplementary website for study details and data1 . 353
Software Subjects (tools) 354
In Experiment 1 we leverage prior work which explored a set of four bioinformatics 355
tools [25]. That work demonstrated the effect of parameters on various objectives, and 356
identified software failures, but did not analyze the impact on the scientific outcome in 357
depth. In this work, we provide an alternative analysis of the data and focus on the 358
extent of the changes in the results, and whether they conform to scientific expectations. 359
We use an exemplar tool for each of the three bioinformatics applications described 360
in the Background. For one of these applications we use two different versions of a tool 361
resulting in four subjects in total. Criteria for choosing the tool subjects include: (1) 362
the tool is frequently used in the community, (2) has a set of modifiable parameters, (3) 363
can be run using automated scripts, and (4) includes a tutorial (or sample data) and 364
documentation. We explicitly use data from others to avoid biasing our findings, and 365
use tool documentation to guide parameter settings and ranges. For several of the 366
subjects we utilize implementations from the U.S. Department of Energy’s System 367
Biology Knowledgebase (KBase) which is a software and data science platform for 368
researchers to analyze, integrate and model plant and microbial physiology and 369
community dynamics [26]. 370
The exemplar DNA alignment subject is the Nucleotide BLAST Basic Local 371
Alignment Search Tool (BLASTn), a popular bioinformatics tool developed by the 372
National Center for Biotechnology Information (NCBI) [4, 27]. We use the command line 373
implementation of nucleotide BLAST (BLASTn), referred to simply as BLAST from 374
here on [28]. The second subject is a DNA assembly tool. We chose the MEGAHIT 375
algorithm [29, 30], and use a version implemented within KBase (version 2.2.8) [31]. 376
The last two subjects perform Flux Balance Analysis (FBA). The first subject is the 377
graphical user interface (GUI) app, Run Flux Balance Analysis [32], from within 378
KBase’s main interface (herein referred to as FBA-GUI). It exposes only a limited set of 379
all the possible parameters available in the underlying algorithm of FBA to be more 380
tractable for new KBase users. Alternatively users may also adopt the the command 381
line interface (CLI) implementation of FBA (herein referred to as FBA-CLI). This FBA 382
subject exposes many more parameters which can be manipulated. We utilize the open 383
source standalone version (toolkit) of the FBA version 1.7.4 in this work [33]. 384
Public KBase narratives demonstrating all experimentation discussed here are 385
provided2 . 386
Experiment 1: Impact of Changing Configurations 387
This first study aims to answer the first research question: how does a tool’s behavior 388
changes when parameter settings are modified. 389
Methodology 390
The methodology here follows the experimental approach previously outlined: SOMATA: 391
Select tool and data, choose Objective metric, Model parameter space, choose sample 392
1 Released upon publication
2 Released upon publication
December 20, 2022 12/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
design Approach, Test, and Analyze. Starting with step 1 we Select the tools to be 393
evaluated, we chose four subjects: BLAST, FBA-GUI, FBA-CLI, and MEGAHIT. 394
In step 2 we define a set of functional outputs as Objective metrics for each tool. 395
For BLAST we use the number of quality hits using a strict definition of 100% identity 396
value and and e-value of 0.0. This is a common measurement used to determine how 397
well the query has performed. For FBA-GUI and FBA-CLI we use an objective metric 398
that represents growth or biomass called the objective value. For MEGAHIT we selected 399
four different metrics. The first two metrics count the number of contigs after a filter is 400
applied. The metric gt1kbp is the number of contigs of length greater than one thousand 401
base pairs, and the metric gt10kbp is the count greater than 10 thousand base pairs. 402
Metric N50 is the N50 score provided by MEGAHIT which is a weighted median 403
statistic on the contig length. The last metric, Max-bp, is the length of the longest 404
contig found. Note that these metrics can be adjusted to suit multiple use-cases. In this 405
work we are simply restricting that choice. 406
Next in step 3, we define a Model of the parameter space for each subject tool by 407
selecting a sample of its parameters to test as seen in Table 3. For each tool we show 408
the number of parameters, and for each parameter we list the type (i.e. Boolean, 409
integer, float, or string) and state the number of choices used. For instance, we have two 410
choices for Boolean options, three for strings and five each for integers or floats. As an 411
example, if we examine BLAST we can see the model has three Boolean, three floats, 412
and one string option for a total of seven options. This results in a combinatorial 413
parameter space of 23 × 53 × 3 or 3, 000 different configurations. Please refer to our 414
supplementary data for the exact parameters tested. 415
In step 4 we design an Approach for sampling the parameter space. For BLAST, 416
MEGAHIT, FBA-GUI we exhaustively test all combinations of their configuration 417
options. For FBA-CLI the total configuration space is 1.28 × 1016 which is too many to 418
exhaustively test. Thus we took a random sample of 125,000 configurations. Note that 419
the configuration space for MEGAHIT of size 260 has been reduced from the full space 420
of 625 due to constraints between parameters. Please refer to our supplementary data 421
for more information. 422
To measure the impact of changing parameters we measure several data points to 423
observe the number of configurations that change the output (for better or for worse). 424
We can also look at the entire configuration space given our input data and research 425
objective, and ask how hard it is to find a better, or even the best result compared to 426
the default value. 427
Table 3. Subjects. For each subject we show the number of total parameters followed
by the number chosen for testing broken down by parameter type. The number in
parenthesis is the number of choices for that type. Constraints on options are not
reflected here, please refer to our supplementary website for more information.
BLAST MEGAHIT FBA-GUI FBA-CLI
Parameters 58 7 17 52
Parameters Selected for Model
Boolean (2) 3 - 3 21
Integer (5) - 4 - -
Float (5) 3 - 5 14
String (3) 1 - - -
Total 7 4 8 35
Config. Space 3,000 260 25,000 1.28 x 1016
December 20, 2022 13/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
Results 428
After executing the experiment (Test in step 5), Table 4 presents the results for our 429
Analysis of the effects in step 6. Each row represents one subject tool and objective 430
metric combination. Table 4a reports the number of configurations tested, the number 431
of unique output metrics, and a summary of the range of output metrics (default, best, 432
worst, and most common). This demonstrates the impact on the raw objective metric 433
demonstrating the effect the configuration space has on the scientific output. Table 4b 434
breaks down the percentage variation to give an idea of the landscape of the outputs. 435
In Table 4a we can see the number of unique objective values is low compared with 436
the total number of configurations tested in most subjects, suggesting random sampling 437
may be very helpful. However in MEGAHIT, for the N50 and gt1kbp metrics, we see 438
about half of the configurations leading to unique objective values. This may suggest we 439
need to create larger samples to fully explore all potential behaviors of those objectives. 440
Looking at the range of the objective metrics (last four columns), we can see in some 441
cases there is a large increase in the result comparing the default output to the best 442
(BLAST, FBA-CLI, MEGAHIT gt1kbp). Other subjects show a smaller range. However 443
all subjects have a substantially lower worst metric, and all except BLAST have an 444
output of zero as their most common output metric. 445
In Table 4b we can see the landscape of the distribution of the scores varies greatly 446
on the subject and objective. For BLAST 72% of the configurations improve over the 447
default, but the best result is only seen in 13.33% of the configurations. In FBA-GUI 448
the default is the best value, but that is only achievable by 18.43% of configurations. In 449
FBA-CLI - the largest configuration space - only 1.58% of the random configurations 450
improve over the default’s output, and 0.05% result in the best output value. In fact 451
most of the time (in 95.14% of configurations) the worst value (0 or no growth) is 452
achieved. For MEGAHIT the odds of improving over the default range from 2.69% (for 453
N50 score) to 66.15% for the number of contigs greater than 1,000 base-pairs. Finding 454
the optimal value for any MEGAHIT metric is low across all subjects ranging from a 455
0.38% to 1.54%. We see that the number of configurations that achieve the same result 456
as the default ranges across subjects, as low as 0.38% for MEGAHIT (measured by 457
maximum base pairs) and as high as 18.43% for FBA-CLI. In all subjects more than 458
80% of the results are different from the default. 459
December 20, 2022 14/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
Table 4. Analysis of objective metric impacts.
(a) Objective Metric Impacts. Model is the tool (and objective metric) used. The No. Configs Tested column shows the number of
configurations tested. The No. Unique column is the number of unique output values from all configurations tested as an indication of
the diversity of output. The next four columns are the raw objective metric output values for the default, best, worst, and most
common output values.
Objective Metric
No. Configs
Model No. Unique Default Best Worst Most Common
Tested
BLAST
3,000 17 2,569 4,351 1,809 2,569
(no. quality hits)
FBA-GUI
25,000 4 0.508 0.508 0 0
(OV)
FBA-CLI
125,000 39 0.508 1.171 0 0
(OV)
MEGAHIT
260 122 284 656 2 0
(gt1kbp)
MEGAHIT
260 47 143 147 0 0
(gt10kbp)
MEGAHIT
260 147 17,227 17,593 0 0
(N50)
MEGAHIT
260 88 64,476 70,112 0 0
(Max-bp)
(b) Percentage Variation Summary. Model is the tool (and objective metric) used. The No. Configs Tested column shows the number
of configurations tested. The remaining columns represent the percentage of configurations tested with respect to the default value. For
example Better represents the percentage of configurations that returned an objective metric better than the default. The final column
is the percentage of configurations that resulted in an error by the software.
No. Configs
Model Default Better Best Worse Worst Most Common Errors
Tested
BLAST
3,000 17.33% 72.00% 13.33% 10.67% 2.00% 17.33% 0.00%
(no. quality hits)
FBA-GUI
25,000 18.43% 0.00% 18.43% 81.57% 67.23% 67.23% 0.06%
(OV)
FBA-CLI
125,000 0.56% 1.58% 0.05% 97.60% 95.14% 95.14% 1.03%
(OV)
MEGAHIT
260 1.15% 66.15% 0.38% 9.62% 0.38% 23.08% 0.00%
(gt1kbp)
MEGAHIT
260 8.46% 17.69% 0.38% 50.77% 7.31% 30.38% 0.00%
(gt10kbp)
MEGAHIT
260 2.31% 2.69% 0.38% 95.00% 0.38% 23.08% 0.00%
(N50)
MEGAHIT
260 0.38% 13.46% 1.54% 63.08% 0.38% 23.08% 0.00%
(Max-bp)
December 20, 2022 15/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
Summary
These results show that the default configuration does not always provide the
best objective value, and that instead of randomly choosing configurations, a
more systematic method of investigation is needed. For example in all subjects
except BLAST, if a user randomly chooses a configuration, most often the
worst value (0) will be obtained. The variation in the value of the result
also demonstrates the importance of reporting the exact configuration
used in any documentation or publications; otherwise the results are
not reproducible.
The ranges in the configuration distributions in MEGAHIT tell us several things.
We can see that the odds of improving over the default N50 score is significantly
less than the other use cases. This signals that MEGAHIT may be optimized for
the N50 score over other use cases. However, if a user is more concerned with
achieving more contigs with sizes greater than 1,000 base-pairs they may need
to consider other configurations. Configurations are not one-size-fits-all. This
demonstrates how essential it is for the user to understand their particular use
case, have an effective metric that is representative of the objective value, and
consider how different parameters may affect it.
460
Experiment 2: Experimental exploration approach 461
Motivated by the results of Experiment 1 demonstrating tool configurations can have a 462
large impact on the scientific conclusion, in this next study we ask how a user might use 463
an experimental approach to explore the parameter space and infer the internal 464
behavior of the system. We do so through two studies. In Experiment 2a, we focus on a 465
single parameter to explore in depth how this parameter affects an objective. In the 466
process we uncover some unexpected behavior. In Experiment 2b we explore the overlap 467
in functionality between multiple parameters. We noticed that some parameters can be 468
changed from either setting a single parameter within the GUI interface, or alternative 469
sources. We wanted to understand the interplay between these choices. Experiment 2a 470
demonstrates how a user can systematically probe a software tool to verify and learn 471
the impact of single parameters. Experiment 2b provides a method to explore the 472
interaction between multiple parameters. 473
Experiment 2a: In-Depth Study of Single Parameter 474
We extend the experiment from our motivating example on two additional subjects to 475
observe if the effect is the same or different. 476
Methodology 477
Following step 1 of our protocol we Select the KBase GUI implementation of FBA as 478
the subject tool and select three subject organisms that come from tutorial or public 479
data in KBase. We tested both Escherichia coli (E. coli ) and Rhodobacter sphaeroides 480
(R. sphaeroides) growing in the standard Carbon-D-Glucose media. We also tested 481
Shewanella oneidensis (S. oneidensis) in the Lactate media. Please refer to our 482
supplementary material for an in-depth description of the experimental design. Per 483
step 2, we use the same Objective metric to evaluate as Experiment 1, growth 484
measured by the objective value, in all cases. For our Model of the parameter space 485
December 20, 2022 16/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
(step 3), we investigate the use of a single parameter the Max Oxygen parameter 486
(maximum number of moles of oxygen permitted for uptake) which according to the 487
specifications can be set between 0 and 100. The default has no explicit set value. We 488
determine the sample design Approach (step 4) by systematically testing a range of 489
values (from 0 to 100 in steps of 10) for max oxygen. In order to test the bounds of the 490
parameter, we also tested values larger than the maximum according to the 491
documentation (100), thus we also tested values of 101, 250, 500, and 1000. 492
We consider a use case where the user may want to identify what setting for the max 493
flux of oxygen results is the highest growth. However they do not want to oversupply 494
the levels of oxygen, so they want to identify the minimal value of the flux of oxygen to 495
achieve the highest OV. 496
Results 497
After executing the experimental Test in step 5, Table 5 presents our results for all 498
three organisms for Analysis in step 6. E. coli and R. sphaeroides both showed similar 499
behavior. The growth for each starts at 0 when max oxygen is set to zero, then 500
increases to the default growth at a max oxygen value of 40 for E. coli and 50 for R. 501
sphaeroides. Any value higher results in no additional growth. 502
S. oneidensis also grows at a linear rate starting from 0 growth at a max oxygen 503
value of 0. However it does not reach the default growth at the maximum value of 100. 504
However, if we go beyond the documentation specified range, the default growth can be 505
achieved by setting max oxygen to 500. Surprised by this behavior we asked the 506
developers of this tool for further information where we learned that the documentation 507
was misleading and that 100 is not a hard maximum value. 508
Table 5. FBA varying MaxO
E. coli S. oneidensis R. sphaeroides
Configuration
(CDG) (Lactate) (CDG)
Default 0.164692 3.271000 0.545390
Max O 0 0 0 0
Max O 10 0.053666 0.113570 0.120263
Max O 20 0.107332 0.227139 0.240526
Max O 30 0.160998 0.340709 0.360790
Max O 40 0.164692 0.454278 0.481053
Max O 50 0.164692 0.567848 0.545390
Max O 60 0.164692 0.681418 0.545390
Max O 70 0.164692 0.794987 0.545390
Max O 80 0.164692 0.908557 0.545390
Max O 90 0.164692 1.007780 0.545390
Max O 100 0.164692 1.099590 0.545390
Max O 101 0.164692 1.108690 0.545390
Max O 250 0.164692 2.123170 0.545390
Max O 500 0.164692 3.271000 0.545390
Max O 1000 0.164692 3.271000 0.545390
December 20, 2022 17/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
Summary
Using a systematic approach we were able to verify the expected behavior that
increasing the maximum allowed oxygen flux would allow for an increase in
organism growth. This increases the user trust in the tool and verifies
our understanding is in line with the tool’s functionality. We were also
able to identify the smallest value of this setting that results in the best scientific
output (40 for E. coli, 50 R. sphaeroides, and 500 for S. oneidensis). This answers
our use case question. Furthermore, we identified some unexpected behavior that
was not in line with the documentation. Tools are not without bugs or mis-
takes in documentation and it is useful for users to be equipped with
alternative methods to be able to identify any unexpected behaviors
on their own data and use case, and clarify them with developers.
509
Experiment 2b: Interaction between parameters and the 510
environment 511
As previously noted, we identified the flux limits can be altered in more than one 512
manner. In this experiment we further investigate how these different methods interplay. 513
One way to control flux is within the media input file via a field for max uptake of each 514
compound with units “mol/per CDW hr”. As a second method, from the GUI the 515
carbon fluxes can be constrained via an application parameter. There are five key 516
nutrients that can be restricted: carbon (C), nitrogen (N), phosphate (P), sulfur (S), 517
and oxygen (O). We will refer to these as MaxC, MaxN, MaxP, MaxS, and MaxO 518
respectively. The description of these options is “Maximum number of moles of 519
[C/N/P/S/O] permitted for uptake (default uptake rates varies from 0 to 100 for all 520
nutrients)”. There is yet a third way to constrain the fluxes. We can also set individual 521
limits on either compounds or reactions using a different (string) parameter (Custom 522
Flux Bounds). We will explore the first two methods. The third method can be 523
explored using a similar approach. 524
This experiment investigates how changing different parameters affects the final 525
scientific result. For example if there is a limit of 10 on MaxO through the application 526
parameter, and a limit of 40 for H2 O specified by the media, what is the resulting limit 527
for the exchange flux of H2 O? Does one supersede another? Or is the minimum or 528
maximum taken as the limit? How is the flux of multiple compounds affected if they all 529
share common nutrients? The impact of such changes may be non-trivial to understand 530
without a systematic exploration. Furthermore, the output of the FBA tool produces 531
multiple values for exchange fluxes including upper bound, max exchange flux, and 532
exchange flux. It may not be obvious to a user what precisely each of these fields mean 533
and where the limits are coming from. Two specific questions we investigate in this 534
study include: 535
1. If the max flux GUI options are left as unset, what source does FBA use to 536
determine the limits on the uptake of the different carbon? 537
2. How can we identify the minimal, required uptake rate for each nutrient? 538
Methodology 539
Following step 1 we Select as the subject tool Flux Balance Analysis with E.coli in the 540
Carbon-D-Glucose media as the base media. The outcome metric (step 2) remains as 541
the Objective value. In order to explore the sources of the flux constraints, the first 542
December 20, 2022 18/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
step was to identify what compounds in the media the important nutrients could be 543
coming from. For carbon, nitrogen, phosphate, and sulfur there is only one source for 544
each: D-Glucose (C6H12O6), NH3 (H4N), Phosphate (HO4P), and Sulfate (O4S) 545
respectively. Oxygen has several sources: Dioxide (O2), Sulfate (O4S), Phosphate 546
(HO4P), H2O, Molybdate (H2MoO4), D-Glucose (C6H12O6). For simplicity, we chose 547
two cases where there was only one source: carbon (MaxC) with source D-Glucose and 548
nitrogen (MaxN) with source NH3. We left the media limits fixed (5 for D-Glucose and 549
100 for NH3), and then varied the max flux limits of carbon and nitrogen respectively. 550
This defines our Model of the parameter space (step 3). 551
For the sample design Approach in step 4 we take a different approach than the 552
previous studies. We take a more iterative exploratory approach by starting with the 553
default settings (unset in this case), altering them by a value of 10, and reassessing 554
based on the results. In this type of a design, we started with an initial model (default 555
as well as increments of 10), experimentally Tested (step 5), then observed and 556
Analysed (step 6) and went back to add to the Model (step 3) and re-evaluate. Since 557
there was no change as the Max uptake of N was increased from 10 to 20 and the max 558
exchange flux was observed to be 1.5551, further exploration was performed by limiting 559
the max uptake of N at values between 1 and 2. 560
Results 561
Results for exploring the MaxC option can be seen in Table 6. The rows upper bound, 562
max exchange flux, exchange flux, and OV are all output values of the FBA object. 563
Based on these results we can extrapolate about how the tool is working.
Table 6. Max Uptake C Exploration - D-Glucose (C6H12O6)
Max Uptake C Default 10 20 30
Upper bound 5 5 5 5 ← media defined limit
Max exchange flux 5 1.667 3.333 5 ← parameter defined
Exchange flux 5 1.667 3.333 5 limit
OV 0.675 0.225 0.450 0.675
564
We can see that the upper bound is consistently at a value of 5, this must mean the 565
upper bound is set by the media file. The field for max exchange flux however varies. 566
Under the default parameter we see it is identical to the upperbound. However when we 567
constraint the amount of MaxC this value varies. It is reduced in MaxC=10 and MaxC=20, 568
but at MaxC=30 we see it is again equal to the default of 5. When E.coli has at least 5 569
units of carbon, it will grow at is maximum amount (0.675). Furthermore, we can see 570
both the uptake and the OV increase linearly. At a MaxC of 10 the uptake is 1/3rd of 571
the uptake under the default, and the OV is also 1/3rd of the OV under the default. At 572
MaxC of 20 the uptake and OV are 2/3rds of the corresponding values under the default. 573
Table 7. Max Uptake N Exploration - NH3 (H4N)
Max Uptake N Default 1 1.5 1.5551 2 10 20
Upper bound 100 100 100 100 100 100 100
Max Exchange flux 12.9392 1 1.5 1.5551 2 10 20
Exchange flux 1.55528 1 1.5 1.5551 1.55528 1.55528 1.55528
OV 0.164692 0.116501 0.164269 0.16469 0.164692 0.164692 0.164692
We repeat this process for nitrogen seen in Table 7. In this case we vary the nitrogen 574
supplying compound (NH3) initially at levels of 10 and 20. In follow-up exploration, we 575
also used levels of 1, 1.5, 1.5551, and 2. At 2 units and above, E.coli grows at is 576
maximum (default) amount. 577
December 20, 2022 19/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
We observe that the exchange flux for NH3 is limited to 1.55528 as this is the 578
amount of uptake for 2 through 20 units of nitrogen as well as the default case. Further, 579
E.coli grows at its maximum (default) amount for these amounts of NH3. If we set a 580
value slightly smaller than that as our MaxN (e.g. 1.5551) we see that it now grows at 581
just 0.000002 less than its maximum potential. Further, at MaxN=1 it grows at 70.74% 582
of its maximum amount while at MaxN=1.5 it grows at 99.74% of its maximal potential. 583
Summary
This experiment reverse engineered the effect of changing max flux options in FBA,
and investigated how the application parameter interacts with the media options.
We were able to extract the meaning of two of the output values that
were unclear from documentation. We found that upper bound is the flux
bound by the media and max exchange flux is bound by the app parameter. We
found that the max exchange flux is correlated with which compounds in the
media contain those nutrients, and that the minimum value between the two is
used as the limiting value. We can also identify the minimal required exchange
flux by looking at the actual exchange flux reported by the output.
This process shows how a user can systematically perturb a scientific
software system in order to reverse engineer pieces of the underlying
algorithm. It was initially unclear from documentation how the interaction
between these options operated, and what the meaning of the different output
fields meant. By using this experimental approach the user was able to discover
the answers on their own. Similar experimental designs could be set up to ask
different types of questions.
584
Discussion 585
We have seen using three exemplar experiments that biologists can benefit from 586
approaching their computational tools the same way they approach their bench 587
experiments. Instead of using tools as magic boxes, we propose SOMATA, an 588
exploratory approach that leads to computational understanding and expertise. By 589
leveraging the configuration options (which were developed for specific use cases), a 590
biologist can better understand the range of a tool’s application and thereby optimize 591
the results for their use case. However, this is also a cautionary tale. Modifying 592
configurations of a tool may change how the tool works in unintended ways, and doing 593
so in an ad-hoc manner can lead to results that are not reproducible or are potentially 594
sub-optimal. 595
We also demonstrated that some parameters, while well documented, provide 596
inaccurate information. While the tool authors document the valid values, these were in 597
fact incorrect limits for the tool. We discovered via experimentation, and confirmed 598
with the developers, that we can use larger values in the application, and that larger 599
values result in (correct) larger values of growth. Situations like this can lead to a 600
confusing landscape for biologists. Hence, we suggest using a FAIR-like (Findable, 601
Accessible, Interoperable and Reusable) approach [34] not only with data collection but 602
also to using and reporting interactions and final settings with all tools, just as FAIR 603
has been proposed to be extended to research tools [35]. 604
The FAIR initiative came about in recognition that science was being impeded by 605
the lack of reproducibility caused by lax data standards. In a similar way, the 606
advancement of research may also be hampered by ad hoc use of software tools that are 607
December 20, 2022 20/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
increasingly becoming an integral part of scientific investigation. As noted in the 608
original article on FAIR [34] the principles should be applied equally to workflows and 609
software tools, and data: “All scholarly digital research objects [36] — from data to 610
analytical pipelines—benefit from application of these principles, since all components 611
of the research process must be available to ensure transparency, reusability and 612
reproducibility.” Therefore, it is incumbent on researchers to have a working 613
understanding of the tools used in the context of their research, and to properly 614
investigate and communicate their findings in a manner adhering to the principles of the 615
scientific method. 616
We now synthesize and discuss some of the takeaways (or key lessons learned) during 617
this study. 618
1. Importance of Configuration Options. The key takeaway from this study is 619
that configuration options can have a direct impact on the output and can 620
effectively change the biological state of the experiment. Thus, it is essential a 621
user understand the impact each option has. Furthermore, if a user does not take 622
advantage of the variety of configuration options, they may be missing out on 623
functionality. On the flip side, we have also seen that having a limited 624
understanding (or simplified/incorrect documentation) of different parameters 625
may cause problems, hence the burden should not be solely on the biologist. 626
Configurations matter, and the creation, documentation, and experimentation 627
with them should be a community effort. We envision community resources where 628
information from both experimentalists and tool developers interact. Rather than 629
simply FAIR we suggest we also need to adopt a SHARE (Simple, Helpful, 630
Accessible, Readable with Explanations) principle. First, we need methods that 631
are easy to implement and understand so that biologists use them. Second, our 632
systems should provide more hints (or help) to aid users in understanding the 633
various configuration options. Third, the differences (or impact of) changing 634
configuration options should be annotated with readable explanations. None of 635
this can be achieved in isolation, hence we think this is a community effort. 636
2. Mapping Configurations to Behavior May not be Obvious. If we return 637
to the documented max target sequence parameter in BLAST (discussed in the 638
Introduction), we saw that a user could easily interpret the use of setting that 639
option to represent an output filter after the search returns some number of hits; 640
hence setting it to a small number could provide just the best results. But in fact, 641
this parameter is used internally to filter results early in the algorithm, hence it 642
may cause the search to perform sub-optimally. Why this parameter exists is out 643
of the scope of this paper, but it was likely added to improve performance (i.e. 644
make the search run faster). When that objective is required, then a less than 645
optimal search may be beneficial. Had the author of that paper used our approach 646
to explore parameter options they would have seen inconsistent results when 647
setting this parameter (in fact it would only ‘by chance’ return only a single hit). 648
The lesson here is that relying on simplified (or only) documentation could be 649
deceptive. 650
3. We Need Better Tools. While we propose the use of exploration in this paper, 651
we also see the need for next generation tools which provide explanations and 652
expose configuration behavior. As science continues to advance, computational 653
tools are increasingly vital to discovery. In order to maximize their benefit, we 654
should strive to ensure they perform in an explainable, transparent, and intuitive 655
way. 656
December 20, 2022 21/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
Conclusion 657
In biological research, as the variety and resolution of data grows to increasingly reflect 658
the complexity of biology, so too are the associated computational tools becoming more 659
comprehensive mirrors of the same complexity. Just as scientists apply methods to 660
investigate the complexity through laboratory protocols, and shifting approaches not 661
just through reductionism, but also taking an increasingly more comprehensive systems 662
approach, we propose through SOMATA here taking a systems view to bioinformatics 663
research and tool development that mimics how biologists explore organisms in the 664
laboratory, and therefore provides a common basis for communication and interaction 665
that would help the research community overall to more productively develop and use 666
those tools, and thereby advance their research efforts. 667
We have demonstrated that different results can be obtained using different 668
algorithm and tool parameters and that this is a potential minefield for the novice user. 669
Rather than limit exploration, we argue that the community of bioinformatics 670
researchers should provide open, explainable tools and that this information should be 671
shared and crowd sourced to provide a more repeatable environment. While the FAIR 672
principles are of value here, we see a need for a more comprehensive way to view 673
configurability in software tools. 674
A systems approach to biology implies the need for this same kind of interchange 675
within the research community especially as the computational tools are refined to be 676
more precise reflections of biological elements and the experimental processes 677
investigating them. Having a common framework for decomposing these increasingly 678
complex tools that mimics a systems view of biology is the kind of standard that we 679
believe would substantially advance the overall scientific research effort toward the 680
future. 681
Supporting information 682
Table 8. BLAST configuration model
Parameter Name Category Type Range Tested
dust Query filtering options String yes, “20 64 1”, no
soft masking Query filtering options Boolean TRUE, FALSE
lcase masking Query filtering options Flag TRUE, FALSE
xdrop ungap Extension options Real 0, 0.1, 0.5, 20, 100
xdrop gap Extension options Real 0, 0.1, 0.5, 30, 100
xdrop gap final Extension options Real 0, 0.1, 0.5, 10, 100
ungapped Extension options Flag TRUE, FALSE
Table 9. MEGAHIT configuration model
Parameter Name Type Range Allowed Range Tested
min-count Numeric [2,10] 2, 4, 6, 8, 10
k-min Numeric odd in range [1,127] 15, 21, 59, 99, 141
k-max Numeric odd in range [1,255] 15, 21, 59, 99, 141
k-step Numeric even in range [2,28] 2, 6, 12, 20, 28
December 20, 2022 22/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
Table 10. FBA-GUI configuration model
Parameter Name Type Range Tested
flux variability analysis Boolean TRUE, FALSE
minimize flux Boolean TRUE, FALSE
simulate all single KOs Boolean TRUE, FALSE
max Carbon uptake Float 0, 25, 50, 75, 100
max Nitrogen uptake Float 0, 25, 50, 75, 100
max Phosphate uptake Float 0, 25, 50, 75, 100
max Sulfur uptake Float 0, 25, 50, 75, 100
max Oxygen uptake Float 0, 25, 50, 75, 100
Table 11. FBA-CLI configuration model
Parameter Name Type Range Tested
minimize flux Boolean TRUE, FALSE
simulate all single Kos Boolean TRUE, FALSE
find min media Boolean TRUE, FALSE
allRevMFA Boolean TRUE, FALSE
useVarDrainFlux Boolean TRUE, FALSE
rxnKOSen Boolean TRUE, FALSE
decompDrain Boolean TRUE, FALSE
decompRxn Boolean TRUE, FALSE
tightBounds Boolean TRUE, FALSE
phenoFit Boolean TRUE, FALSE
forceUse Boolean TRUE, FALSE
maxActiveRxn Boolean TRUE, FALSE
newPipeline Boolean TRUE, FALSE
prefMFA Boolean TRUE, FALSE
rxnUseVar Boolean TRUE, FALSE
recMILP Boolean TRUE, FALSE
useDataFields Boolean TRUE, FALSE
simpVar Boolean TRUE, FALSE
LPFile Boolean TRUE, FALSE
varKey Boolean TRUE, FALSE
optimMetabo Boolean TRUE, FALSE
maxDrain Float 0, 250, 500 ,750, 1000
minDrain Float 0, -250, -500 , -750, -1000
deltaGSlack Float 0, 5, 10, 15, 20
maxDeltaG Float 0, 2500, 5000, 7500, 10000
maxFlux Float 0, 250, 500 ,750, 1000
minFluxMulti Float 0, 1, 2, 3, 4
minFlux Float 0, -250, -500 , -750, -1000
conObjec Float 0, 0.05, 0.1, 0.15, 0.2
minTargetFlux Float 0, .005, .01, .015, .02
Max O uptake Float 0, 25, 50, 75, 100
Max N uptake Float 0, 25, 50, 75, 100
Max P uptake Float 0, 25, 50, 75, 100
Max S uptake Float 0, 25, 50, 75, 100
Max C uptake Float 0, 25, 50, 75, 100
December 20, 2022 23/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
Acknowledgments 683
This work has been supported in part by the National Science Foundation grant
#CCF-1909688. The opinions and views presented are that of the authors and not of
the NSF. This research has also been supported by the Plant-Microbe Interfaces
Scientific Focus Area in the Genomic Science Program of the Office of Biological and
Environmental Research (BER) at the U.S. Department of Energy Office of Science
under Contract number DE-AC05-00OR22725. This research used resources of the Oak
Ridge Leadership Computing Facility, which is a DOE Office of Science User Facility
supported under Contract DE-AC05-00OR22725.
References
1. Finkel OM, Salas-González I, Castrillo G, Conway JM, Law TF, Teixeira PJPL,
et al. A single bacterial genus maintains root growth in a complex microbiome.
Nature. 2020;587(7832):103–108.
2. Voit EO. Perspective: Dimensions of the scientific method. PLOS Computational
Biology. 2019;15(9):1–14. doi:10.1371/journal.pcbi.1007279.
3. Sun SL, Yang WL, Fang WW, Zhao YX, Guo L, Dai YJ, et al. The Plant
Growth-Promoting Rhizobacterium Variovorax boronicumulans CGMCC 4969
Regulates the Level of Indole-3-Acetic Acid Synthesized from
Indole-3-Acetonitrile. Applied and Environmental Microbiology.
2018;84(16):e00298–18. doi:10.1128/AEM.00298-18.
4. Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment
search tool. Journal of molecular biology. 1990;215(3):403–410.
5. Shah N, Nute MG, Warnow T, Pop M. Misunderstood parameter of NCBI
BLAST impacts the correctness of bioinformatics workflows. Bioinformatics.
2018;35(9):1613–1614. doi:10.1093/bioinformatics/bty833.
6. Bethesda (MD): National Center for Biotechnology Information (US). BLAST
Command Line Applications User Manual; 2008. Available from:
https://www.ncbi.nlm.nih.gov/books/NBK279684/table/appendices.T.
options_common_to_all_blast/t.
7. Madden T, Busby B, Ye J. Reply to the paper: Misunderstood parameters of
NCBI BLAST impacts the correctness of bioinformatics workflows.
Bioinformatics. 2019;35(15):2699–2700.
8. Morrison-Smith S, Boucher C, Bunt A, Ruiz J. Elucidating the role and use of
bioinformatics software in life science research. In: Proceedings of the British
HCI Conference. ACM; 2015. p. 230–238.
9. Jamshidi P, Siegmund N, Velez M, Kästner C, Patel A, Agarwal Y. Transfer
Learning for Performance Modeling of Configurable Systems: An Exploratory
Analysis. In: IEEE/ACM International Conference on Automated Software
Engineering (ASE). IEEE; 2017. p. 497–508.
10. Cohen MB, Dwyer MB, Shi J. Constructing Interaction Test Suites for
Highly-Configurable Systems in the Presence of Constraints: A Greedy Approach.
IEEE Transactions on Software Engineering. 2008;34(5):633–650.
December 20, 2022 24/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
11. Miller G. Scientific publishing. A scientist’s nightmare: software problem leads to
five retractions. Science (New York, NY). 2006;314(5807):1856.
12. Ichikawa T, Suzuki Y, Czaja I, Schommer C, Leßnick A, Schell J, et al.
Retraction Note to: Identification and role of adenylyl cyclase in auxin signalling
in higher plants. Nature. 1998;396. doi:10.1038/24659.
13. Orth JD, Thiele I, Palsson BØ. What is flux balance analysis? Nature
Biotechnology. 2010;28(3):1546–1696.
14. FAQ: Metabolic Modeling; 2022. Available from: https:
//docs.kbase.us/workflows/metabolic-models/faq-metabolic-modeling.
15. Jin D, Qu X, Cohen MB, Robinson B. Configurations Everywhere: Implications
for Testing and Debugging in Practice. In: International Conference on Software
Engineering, Software in Practice Track. ICSE. ACM; 2014. p. 215–225.
16. Sayagh M, Hassan AE. ConfigMiner: Identifying the Appropriate Configuration
Options for Config-related User Questions by Mining Online Forums. IEEE
Transactions on Software Engineering. 2020; p. 1–1.
doi:10.1109/TSE.2020.2973997.
17. Clements P, Northrop L. Software Product Lines: Practices and Patterns.
Addison-Wesley Professional; 2001.
18. Johnson M, Zaretskaya I, Raytselis Y, Merezhuk Y, McGinnis S, Madden TL.
NCBI BLAST: a better web interface. Nucleic Acids Research.
2008;36(2):W5–W9. doi:10.1093/nar/gkn201.
19. Yilmaz C, Cohen MB, Porter AA. Covering Arrays for Efficient Fault
Characterization in Complex Configuration Spaces. IEEE Trans Software Eng.
2006;32(1):20–34. doi:10.1109/TSE.2006.8.
20. Zhang S, Ernst MD. Which Configuration Option Should I Change? In:
Proceedings of the 36th International Conference on Software Engineering. ICSE
2014. New York, NY, USA: Association for Computing Machinery; 2014. p.
152–163. Available from: https://doi.org/10.1145/2568225.2568251.
21. Qu X, Cohen MB, Rothermel G. Configuration-aware Regression Testing: An
Empirical Study of Sampling and Prioritization. In: International Symposium on
Software Testing and Analysis. ISSTA. ACM; 2008. p. 75–86.
22. Wooley JC, Godzik A, Friedberg I. A primer on metagenomics. PLoS
computational biology. 2010;6(2):e1000667.
23. Vollmers J, Wiegand S, Kaster AK. Comparing and evaluating metagenome
assembly tools from a microbiologist’s perspective-not only size matters! PloS
one. 2017;12(1):e0169662.
24. Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic
acids research. 2000;28(1):27–30.
25. Cashman M, Cohen MB, Ranjan P, Cottingham RW. Navigating the Maze: The
Impact of Configurability in Bioinformatics Software. In: Proceedings of the 33rd
ACM/IEEE International Conference on Automated Software Engineering. ASE
2018. New York, NY, USA: ACM; 2018. p. 757–767.
December 20, 2022 25/26
bioRxiv preprint doi: https://doi.org/10.1101/2022.12.20.521257; this version posted December 21, 2022. The copyright holder for this preprint
(which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made
available under aCC-BY 4.0 International license.
26. Arkin AP, Cottingham RW, Henry CS, Harris NL, Stevens RL, Maslov S, et al.
KBase: the United States department of energy systems biology knowledgebase.
Nature biotechnology. 2018;36(7):566.
27. Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, et al.
Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs. Nucleic acids research. 1997;25(17):3389–3402.
28. Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, et al.
BLAST+: architecture and applications. BMC bioinformatics. 2009;10(1):421.
29. Li D, Liu CM, Luo R, Sadakane K, Lam TW. MEGAHIT: an ultra-fast
single-node solution for large and complex metagenomics assembly via succinct de
Bruijn graph. Bioinformatics. 2015;31(10).
30. Li D, Luo R, Liu CM, Leung CM, Ting HF, Sadakane K, et al. MEGAHIT v1. 0:
a fast and scalable metagenome assembler driven by advanced methodologies and
community practices. Methods. 2016;102:3–11.
31. KBase MEGAHIT SDK Repository; 2017. Available from:
https://github.com/kbaseapps/kb_megahit.
32. Henry CS, DeJongh M, Best AA, Frybarger PM, Linsay B, Sevens RL.
High-throughput generation, optimization and analysis of genome-scale metabolic
models. Nature Biotechnology. 2010;28(9):977–982.
33. Henry CS. MFAToolkit GitHub Repository; 2017. Available from:
https://github.com/cshenry/fba_tools/tree/master/MFAToolkit.
34. Wilkinson MD, Dumontier M, Aalbersberg IJ, Appleton G, Axton M, Baak A,
et al. The FAIR Guiding Principles for scientific data management and
stewardship. Scientific data. 2016;3(1):1–9.
35. Lamprecht AL, Garcia L, Kuzak M, Martinez C, Arcila R, Martin Del Pico E,
et al. Towards FAIR principles for research software. Data Science.
2020;3(1):37–59.
36. Bechhofer S, De Roure D, Gamble M, Goble C, Buchan I. Research objects:
Towards exchange and reuse of digital knowledge. Nature Precedings. 2010; p.
1–1.
December 20, 2022 26/26
You might also like
- Systems Biomedicine: Concepts and PerspectivesFrom EverandSystems Biomedicine: Concepts and PerspectivesEdison T. LiuNo ratings yet
- Following The Organism To Map Synthetic GenomicsDocument4 pagesFollowing The Organism To Map Synthetic GenomicsGRAÇAS PERFUMARIANo ratings yet
- Biofabrication: Micro- and Nano-fabrication, Printing, Patterning and AssembliesFrom EverandBiofabrication: Micro- and Nano-fabrication, Printing, Patterning and AssembliesNo ratings yet
- Synthetic Biology: Steven A. Benner and A. Michael SismourDocument11 pagesSynthetic Biology: Steven A. Benner and A. Michael SismourGanesh NagarajanNo ratings yet
- Ligand Efficiency Indices for Drug Discovery: Towards an Atlas-Guided ParadigmFrom EverandLigand Efficiency Indices for Drug Discovery: Towards an Atlas-Guided ParadigmNo ratings yet
- Bioinformatics Approaches and Applications in PlanDocument13 pagesBioinformatics Approaches and Applications in PlantariqshrishNo ratings yet
- Creating Biological Nanomaterials Using Synthetic Biology: Science and Technology of Advanced MaterialsDocument12 pagesCreating Biological Nanomaterials Using Synthetic Biology: Science and Technology of Advanced MaterialsTrungtin35No ratings yet
- Import. Biotec. II Developments in The Tools and Methodologies of Synthetic Biology 2014Document23 pagesImport. Biotec. II Developments in The Tools and Methodologies of Synthetic Biology 2014Biggarage PamplonaNo ratings yet
- Plant As A Bio RobotDocument4 pagesPlant As A Bio RobotRajesh KesapragadaNo ratings yet
- Capturing Scientific Knowledge in Computable FormDocument17 pagesCapturing Scientific Knowledge in Computable Formjoe trumpNo ratings yet
- Bio in For MaticsDocument4 pagesBio in For MaticsHarfebi FryonandaNo ratings yet
- Versioning Biological Cells For Trustworthy Cell EngineeringDocument12 pagesVersioning Biological Cells For Trustworthy Cell EngineeringAnahí TessaNo ratings yet
- Contoh Essay Biologi MolekulerDocument4 pagesContoh Essay Biologi MolekulerImam Taufiq RamadhanNo ratings yet
- Controlling Microbial Co-Culture Based On Substrate Pulsing Can Lead To Stability Through Differential Fitness AdvantagesDocument27 pagesControlling Microbial Co-Culture Based On Substrate Pulsing Can Lead To Stability Through Differential Fitness AdvantagesHamid MatariNo ratings yet
- xenobots2Document14 pagesxenobots2yaswanthjetti21No ratings yet
- Engineering Life Through Synthetic Biology: OpinionDocument10 pagesEngineering Life Through Synthetic Biology: OpinionMiriam RangelNo ratings yet
- DNAextraction BacteriaDocument3 pagesDNAextraction BacteriaKenneth TuladNo ratings yet
- E PDFDocument3 pagesE PDFYulis diyahNo ratings yet
- PIIS0092867422008467Document2 pagesPIIS0092867422008467Nick PowellNo ratings yet
- Thacker - Bioinformatics and Bio-LogicsDocument31 pagesThacker - Bioinformatics and Bio-Logicsantonio damataNo ratings yet
- Diffusion of Synthetic Biology A Challenge To BiosafetyDocument6 pagesDiffusion of Synthetic Biology A Challenge To BiosafetyJonathan Hermilo Velázquez GarcíaNo ratings yet
- Unit IV: Quantitative Biology and BioinformaticsDocument27 pagesUnit IV: Quantitative Biology and BioinformaticskofirNo ratings yet
- Saccharomyces Genome DatabaseDocument6 pagesSaccharomyces Genome DatabasePartha DasNo ratings yet
- Biosecurity Challenges PDFDocument5 pagesBiosecurity Challenges PDFValentina BetancourtNo ratings yet
- Bioprepardeness BrochureDocument4 pagesBioprepardeness BrochureGilbert MethewNo ratings yet
- Adami C Digital Genetics Unravelling The Genetic BDocument11 pagesAdami C Digital Genetics Unravelling The Genetic BPedro PazNo ratings yet
- Btab 649Document8 pagesBtab 649Yann FleatwoodNo ratings yet
- Womelsdorf Et Al - A Kiosk Stations For The Assessment of Multiple Cognitive Domains and Enrichments of Monkeys - Biorxiv 2021Document31 pagesWomelsdorf Et Al - A Kiosk Stations For The Assessment of Multiple Cognitive Domains and Enrichments of Monkeys - Biorxiv 2021KiluaZhuNo ratings yet
- Drug Metabolism and Pharmacokinetics: Hiroshi Kimura, Yasuyuki Sakai, Teruo FujiiDocument6 pagesDrug Metabolism and Pharmacokinetics: Hiroshi Kimura, Yasuyuki Sakai, Teruo FujiigpaivNo ratings yet
- Computational Validation and Analysis of Semi-Quantitative Data Using In-Silico ApproachesDocument5 pagesComputational Validation and Analysis of Semi-Quantitative Data Using In-Silico ApproachesIJAR JOURNALNo ratings yet
- Integrative Work Ows For Metagenomic Analysis: Frontiers in Cell and Developmental Biology November 2014Document12 pagesIntegrative Work Ows For Metagenomic Analysis: Frontiers in Cell and Developmental Biology November 2014airsrchNo ratings yet
- Microarrays: Biotechnology's Discovery Platform For Functional GenomicsDocument6 pagesMicroarrays: Biotechnology's Discovery Platform For Functional GenomicsJohn sornozaNo ratings yet
- Microbiome Data Science: Understanding Our Microbial PlanetDocument3 pagesMicrobiome Data Science: Understanding Our Microbial PlanetMilton Paul Lopez RamosNo ratings yet
- Programming and Systems Biology: HapterDocument9 pagesProgramming and Systems Biology: HapterkofirNo ratings yet
- New Phytologist - 2023 - Antonelli - Why Plant Diversity and Distribution MatterDocument6 pagesNew Phytologist - 2023 - Antonelli - Why Plant Diversity and Distribution MatteribitaNo ratings yet
- MADHU Industrty 4.0Document17 pagesMADHU Industrty 4.0Nayanashree NNo ratings yet
- Fungi - Guru: Comparative Genomic and Transcriptomic Resource For The Fungi KingdomDocument8 pagesFungi - Guru: Comparative Genomic and Transcriptomic Resource For The Fungi KingdomJulian Rolando Avendaño PeñaNo ratings yet
- Pre Urp 논문 2 CODA Integrating Multi-level Context-Oriented Directed Associations for Analysis of Drug Effects-복사Document12 pagesPre Urp 논문 2 CODA Integrating Multi-level Context-Oriented Directed Associations for Analysis of Drug Effects-복사hryswnbbtvNo ratings yet
- Multispe QDocument17 pagesMultispe QChathuranga De SilvaNo ratings yet
- GenomicsbookchapterDocument38 pagesGenomicsbookchapterGhinaa ZahrahNo ratings yet
- Bio Materials Applications For Nano MedicineDocument470 pagesBio Materials Applications For Nano MedicineJosé RamírezNo ratings yet
- Advanced Materials - 2019 - Steier - Emerging Trends in Information Driven Engineering of Complex Biological SystemsDocument22 pagesAdvanced Materials - 2019 - Steier - Emerging Trends in Information Driven Engineering of Complex Biological SystemsramaNo ratings yet
- Engineering Microbial Consortia Through Spatial OrganizationDocument9 pagesEngineering Microbial Consortia Through Spatial OrganizationRoberto José Haro SevillaNo ratings yet
- Biological Data Analytics and ComputingDocument8 pagesBiological Data Analytics and ComputingChristian RamaculaNo ratings yet
- Genomics For BeginnerDocument9 pagesGenomics For BeginnerludhangNo ratings yet
- Workshop 2020Document12 pagesWorkshop 2020AJMRNo ratings yet
- An Introduction To Bioprocess EngineeringDocument11 pagesAn Introduction To Bioprocess Engineeringalison grijalbaNo ratings yet
- 2021 04 04 438427v1 FullDocument40 pages2021 04 04 438427v1 FullFelipe CourtNo ratings yet
- CSC 121 Computers and Scientific Thinking Fall 2005Document15 pagesCSC 121 Computers and Scientific Thinking Fall 2005ShivamNo ratings yet
- Integrating Ecology Into BiotechnologyDocument6 pagesIntegrating Ecology Into BiotechnologyJuan Pablo Martinez MonteroNo ratings yet
- Deep RL For BiologyDocument33 pagesDeep RL For BiologyMarcello BarylliNo ratings yet
- Towards A Systematic Characterization of Protein Complex Function: A Natural Language Processing and Machine-Learning FrameworkDocument39 pagesTowards A Systematic Characterization of Protein Complex Function: A Natural Language Processing and Machine-Learning FrameworkOrion OriNo ratings yet
- Ijar 34947Document9 pagesIjar 34947Baino Olpugad GeraldNo ratings yet
- Spatial and Temporal Tools For Building A Human Cell AtlasDocument4 pagesSpatial and Temporal Tools For Building A Human Cell AtlasShailesh KeskarNo ratings yet
- Protocell Communication Through The Eyes of Synthetic Organic ChemistsDocument9 pagesProtocell Communication Through The Eyes of Synthetic Organic Chemistsimport.xenoninterNo ratings yet
- Hhs Public Access: Integrative Genomic Analysis by Interoperation of Bioinformatics Tools in GenomespaceDocument8 pagesHhs Public Access: Integrative Genomic Analysis by Interoperation of Bioinformatics Tools in GenomespaceranjanmanishblueNo ratings yet
- 17373.selected Works in Bioinformatics by Xuhua Xia PDFDocument190 pages17373.selected Works in Bioinformatics by Xuhua Xia PDFJesus M. RuizNo ratings yet
- Bioinformatics: Applications NoteDocument2 pagesBioinformatics: Applications Notekaran3036No ratings yet
- RP 6Document13 pagesRP 64NM21CS114 PRATHAM S SHETTYNo ratings yet
- Open Source Hardware To Culture and Image Bacterial Colony Forming Units (CFU)Document11 pagesOpen Source Hardware To Culture and Image Bacterial Colony Forming Units (CFU)Santosh PandeyNo ratings yet
- Liaudatetal 2023Document19 pagesLiaudatetal 2023NealNo ratings yet
- Weak convergence of FEM for semilinear parabolic SPDEsDocument32 pagesWeak convergence of FEM for semilinear parabolic SPDEsNealNo ratings yet
- Linking The Effect of Localised Pitting CorrosionDocument12 pagesLinking The Effect of Localised Pitting CorrosionNealNo ratings yet
- KW Yg 2021 1Document19 pagesKW Yg 2021 1NealNo ratings yet
- Integrated Finite Element Neural Network I FENN For Non Local Continuum Damage Mechanics Response To PublisherDocument57 pagesIntegrated Finite Element Neural Network I FENN For Non Local Continuum Damage Mechanics Response To PublisherNealNo ratings yet
- 1 s2.0 S004578252200799X MainDocument29 pages1 s2.0 S004578252200799X MainNealNo ratings yet
- Finite Element Analysis of Static Response of Functionally Graded Porous Sandwich BeamsDocument9 pagesFinite Element Analysis of Static Response of Functionally Graded Porous Sandwich BeamsNealNo ratings yet
- A Benchmark For Automatic Medical Consultation SysDocument8 pagesA Benchmark For Automatic Medical Consultation SysNealNo ratings yet
- EASY-FIA A Readably Usable Standalone Tool For HigDocument14 pagesEASY-FIA A Readably Usable Standalone Tool For HigNealNo ratings yet
- piRNA and miRNA Can Suppress The Expression of MulDocument18 pagespiRNA and miRNA Can Suppress The Expression of MulNealNo ratings yet
- Comprehensive Evaluation of Peptide de Novo SequenDocument12 pagesComprehensive Evaluation of Peptide de Novo SequenNealNo ratings yet
- Multimodal Learning of Noncoding Variant Effects UDocument8 pagesMultimodal Learning of Noncoding Variant Effects UNealNo ratings yet
- Singh Chapter2 CRCDocument15 pagesSingh Chapter2 CRCNealNo ratings yet
- 3D probabilistic assessment of tunneling induced structural damageDocument37 pages3D probabilistic assessment of tunneling induced structural damageNealNo ratings yet
- SCIBER A Simple Method For Removing Batch EffectsDocument8 pagesSCIBER A Simple Method For Removing Batch EffectsNealNo ratings yet
- Exercise Slows Neural Stem Cell Aging and Ad RiskDocument1 pageExercise Slows Neural Stem Cell Aging and Ad RiskNealNo ratings yet
- EDIFY Eating Disorders Delineating Illness and RecDocument9 pagesEDIFY Eating Disorders Delineating Illness and RecNealNo ratings yet
- snpSTARRseqDocument13 pagessnpSTARRseqNealNo ratings yet
- Integrative Bioinformatics Approaches Indicate A PDocument41 pagesIntegrative Bioinformatics Approaches Indicate A PNealNo ratings yet
- An RNA Virome Analysis of The Pink-Winged GrasshopDocument11 pagesAn RNA Virome Analysis of The Pink-Winged GrasshopNealNo ratings yet
- Comprehensive Analysis of The Expression and CliniDocument19 pagesComprehensive Analysis of The Expression and CliniNealNo ratings yet
- voyAGEr Free Web Interface For The Analysis of AgeDocument38 pagesvoyAGEr Free Web Interface For The Analysis of AgeNealNo ratings yet
- Pan-Cancer Landscape of NEIL3 in Tumor MicroenviroDocument30 pagesPan-Cancer Landscape of NEIL3 in Tumor MicroenviroNealNo ratings yet
- TaME-seq2 Tagmentation-Assisted Multiplex PCR EnriDocument26 pagesTaME-seq2 Tagmentation-Assisted Multiplex PCR EnriNealNo ratings yet
- Explainable AI For Bioinformatics Methods Tools AnDocument23 pagesExplainable AI For Bioinformatics Methods Tools AnNealNo ratings yet
- Development and Trends in Artificial IntelligenceDocument10 pagesDevelopment and Trends in Artificial IntelligenceNealNo ratings yet
- InvestigatingDocument2 pagesInvestigatingNealNo ratings yet
- ASTER Accurately Estimating The Number of Cell TypDocument3 pagesASTER Accurately Estimating The Number of Cell TypNealNo ratings yet
- Attentive Variational Information Bottleneck For TDocument9 pagesAttentive Variational Information Bottleneck For TNealNo ratings yet
- Application of Neutralization Titrations for Acid-Base AnalysisDocument21 pagesApplication of Neutralization Titrations for Acid-Base AnalysisAdrian NavarraNo ratings yet
- DeviceDefaults-7 1 5 32017-1Document2 pagesDeviceDefaults-7 1 5 32017-1juanzziNo ratings yet
- AMETANK REPORT: Roof design calculationsDocument41 pagesAMETANK REPORT: Roof design calculationsHasan arif KısaalioğluNo ratings yet
- Spesifikasi Teknis Pengadaan Peralatan Laboratorium: Direktorat Jenderal Perikanan BudidayaDocument6 pagesSpesifikasi Teknis Pengadaan Peralatan Laboratorium: Direktorat Jenderal Perikanan BudidayaSanabil CitraNo ratings yet
- CS134 Web Site Design QuizDocument6 pagesCS134 Web Site Design QuizMoulinaDasNo ratings yet
- AntennasDocument5 pagesAntennasMiguel Ferrando Rocher100% (1)
- 124 KW 166 HP at 2000 RPM: Net HorsepowerDocument12 pages124 KW 166 HP at 2000 RPM: Net HorsepowerAwanNo ratings yet
- MK300 - Catalogue Relay bảo vệ dòng rò (Earth Leakage Relay) MK300 của MikroDocument2 pagesMK300 - Catalogue Relay bảo vệ dòng rò (Earth Leakage Relay) MK300 của MikroconmavtvnNo ratings yet
- Chilled Water Fittings - CoDocument209 pagesChilled Water Fittings - CoHarish MenonNo ratings yet
- CSV Aug09 PDFDocument43 pagesCSV Aug09 PDFtreda23No ratings yet
- Configurable Controller User's ManualDocument22 pagesConfigurable Controller User's ManualFabio Nelson CubidesNo ratings yet
- Caustic Soda 1Document21 pagesCaustic Soda 1arpit garg100% (1)
- Manual BlowerDocument28 pagesManual Blowermircea_raceanuNo ratings yet
- SCSI Hard Drives: Installation CardDocument6 pagesSCSI Hard Drives: Installation CardHiền Nguyễn VănNo ratings yet
- Ds Eurobalise s21 enDocument6 pagesDs Eurobalise s21 enMcDanieldsNo ratings yet
- Scooptram ST 1520 PDFDocument3 pagesScooptram ST 1520 PDFmarcos abalNo ratings yet
- Pentatonics and Blues Solo Mastery Module 2 Natural MinorDocument6 pagesPentatonics and Blues Solo Mastery Module 2 Natural MinorhogarNo ratings yet
- Lampiran SPSSDocument18 pagesLampiran SPSSnovitaNo ratings yet
- Gigabyte Ga60xt ManualDocument93 pagesGigabyte Ga60xt ManualroscribNo ratings yet
- WorksheetDocument4 pagesWorksheetMarrin MarquesNo ratings yet
- Lecture Notes 1 - Chapter 1Document5 pagesLecture Notes 1 - Chapter 1sohailahmed714319No ratings yet
- Calculation Formula For The CapacitorDocument9 pagesCalculation Formula For The CapacitorMedyouNo ratings yet
- Heat Transfer Checklist for PE ExamDocument2 pagesHeat Transfer Checklist for PE ExamAfzalul Karim NirvickNo ratings yet
- LETHALITY TABLEDocument44 pagesLETHALITY TABLEdiagnoz7auto7carsvanNo ratings yet
- Bluetooth Low Energy DevicesDocument24 pagesBluetooth Low Energy DevicesadonismarcoNo ratings yet
- SCC Bucket Elevators For A Variety of Applications: Catalog No. 201Document16 pagesSCC Bucket Elevators For A Variety of Applications: Catalog No. 201sudheer4079100% (2)
- Arduino Circuits and Projects Guide - ElektorDocument15 pagesArduino Circuits and Projects Guide - ElektordeckerNo ratings yet
- Engineering and Material Standard For Double Pipe Heat ExchangersDocument13 pagesEngineering and Material Standard For Double Pipe Heat ExchangersRezaNo ratings yet
- ps08 sp12 PDFDocument8 pagesps08 sp12 PDFQ_TNo ratings yet
- 01 - 04 - Measurement of Skid Resistance PDFDocument2 pages01 - 04 - Measurement of Skid Resistance PDFFlavioMuhaleNo ratings yet
- Excel Essentials: A Step-by-Step Guide with Pictures for Absolute Beginners to Master the Basics and Start Using Excel with ConfidenceFrom EverandExcel Essentials: A Step-by-Step Guide with Pictures for Absolute Beginners to Master the Basics and Start Using Excel with ConfidenceNo ratings yet
- Learn Python Programming for Beginners: Best Step-by-Step Guide for Coding with Python, Great for Kids and Adults. Includes Practical Exercises on Data Analysis, Machine Learning and More.From EverandLearn Python Programming for Beginners: Best Step-by-Step Guide for Coding with Python, Great for Kids and Adults. Includes Practical Exercises on Data Analysis, Machine Learning and More.Rating: 5 out of 5 stars5/5 (34)
- Clean Code: A Handbook of Agile Software CraftsmanshipFrom EverandClean Code: A Handbook of Agile Software CraftsmanshipRating: 5 out of 5 stars5/5 (13)
- Machine Learning: The Ultimate Beginner's Guide to Learn Machine Learning, Artificial Intelligence & Neural Networks Step by StepFrom EverandMachine Learning: The Ultimate Beginner's Guide to Learn Machine Learning, Artificial Intelligence & Neural Networks Step by StepRating: 4.5 out of 5 stars4.5/5 (19)
- Python Programming For Beginners: Learn The Basics Of Python Programming (Python Crash Course, Programming for Dummies)From EverandPython Programming For Beginners: Learn The Basics Of Python Programming (Python Crash Course, Programming for Dummies)Rating: 5 out of 5 stars5/5 (1)
- Linux: The Ultimate Beginner's Guide to Learn Linux Operating System, Command Line and Linux Programming Step by StepFrom EverandLinux: The Ultimate Beginner's Guide to Learn Linux Operating System, Command Line and Linux Programming Step by StepRating: 4.5 out of 5 stars4.5/5 (9)
- ITIL 4: Digital and IT strategy: Reference and study guideFrom EverandITIL 4: Digital and IT strategy: Reference and study guideRating: 5 out of 5 stars5/5 (1)
- Nine Algorithms That Changed the Future: The Ingenious Ideas That Drive Today's ComputersFrom EverandNine Algorithms That Changed the Future: The Ingenious Ideas That Drive Today's ComputersRating: 5 out of 5 stars5/5 (7)
- Art of Clean Code: How to Write Codes for HumanFrom EverandArt of Clean Code: How to Write Codes for HumanRating: 3.5 out of 5 stars3.5/5 (7)
- Introducing Python: Modern Computing in Simple Packages, 2nd EditionFrom EverandIntroducing Python: Modern Computing in Simple Packages, 2nd EditionRating: 4 out of 5 stars4/5 (7)
- The JavaScript Workshop: Learn to develop interactive web applications with clean and maintainable JavaScript codeFrom EverandThe JavaScript Workshop: Learn to develop interactive web applications with clean and maintainable JavaScript codeRating: 5 out of 5 stars5/5 (3)
- What Algorithms Want: Imagination in the Age of ComputingFrom EverandWhat Algorithms Want: Imagination in the Age of ComputingRating: 3.5 out of 5 stars3.5/5 (41)
- Monitored: Business and Surveillance in a Time of Big DataFrom EverandMonitored: Business and Surveillance in a Time of Big DataRating: 4 out of 5 stars4/5 (1)
- Excel VBA: A Comprehensive, Step-By-Step Guide on Excel VBA Programming Tips and Tricks for Effective StrategiesFrom EverandExcel VBA: A Comprehensive, Step-By-Step Guide on Excel VBA Programming Tips and Tricks for Effective StrategiesNo ratings yet
- Agile Metrics in Action: How to measure and improve team performanceFrom EverandAgile Metrics in Action: How to measure and improve team performanceNo ratings yet
- Software Engineering at Google: Lessons Learned from Programming Over TimeFrom EverandSoftware Engineering at Google: Lessons Learned from Programming Over TimeRating: 4 out of 5 stars4/5 (11)
- Python Programming: Your Advanced Guide To Learn Python in 7 DaysFrom EverandPython Programming: Your Advanced Guide To Learn Python in 7 DaysNo ratings yet