Professional Documents
Culture Documents
Co4 Ha
Uploaded by
kunchalaharikachinmay0 ratings0% found this document useful (0 votes)
2 views1 pageOriginal Title
CO4-HA
Copyright
© © All Rights Reserved
Available Formats
DOCX, PDF, TXT or read online from Scribd
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
© All Rights Reserved
Available Formats
Download as DOCX, PDF, TXT or read online from Scribd
0 ratings0% found this document useful (0 votes)
2 views1 pageCo4 Ha
Uploaded by
kunchalaharikachinmayCopyright:
© All Rights Reserved
Available Formats
Download as DOCX, PDF, TXT or read online from Scribd
You are on page 1of 1
CO4 -HOME ASSIGNMENT
1. Explain substitution matrix and example?
2. pairwise sequence alignment in pattern matching problem.
3. Illustrate nucleotide BLAST (BLASTN) Web Interface.
4. Compare BLASTN and BLASTP.
5. Compare PAM and BLOSUM
6. Compare local and global sequence alignment.
7. Difference between BLAST and FASTA.
8. Analyze dot-matrix methods, dynamic programming, and word methods of sequence
alignment.
9. protein structure prediction in biological data and BALSA pairwise sequence alignment Tool
10. drug discovery and development processes and tools.
11. Difference between PAM and BLOSUM heuristic methods.
12. scoring matrices and dot plot method in sequence alignment.
13. Discuss smith-waterman and needle man dynamic programming algorithm
14. Analyze the algorithm for this form of local alignment and example.
15. List out the pattern matching applications in bioinformatics.
You might also like
- Fasta and BlastDocument3 pagesFasta and BlastSajeel75 HanifNo ratings yet
- Protein ModellingDocument53 pagesProtein ModellingiluvgermieNo ratings yet
- Bioinformatics Is The Inter-Disciplinary Branch of Biology Which Merges Computer Science, Mathematics and Engineering To Study The Biological DataDocument26 pagesBioinformatics Is The Inter-Disciplinary Branch of Biology Which Merges Computer Science, Mathematics and Engineering To Study The Biological DataMalik FhaimNo ratings yet
- Bioinformatics in PAM AND BLOSUMDocument17 pagesBioinformatics in PAM AND BLOSUMgladson100% (15)
- Fasta Sequence DatabaseDocument17 pagesFasta Sequence DatabaseKlevis XhyraNo ratings yet
- Bioinformatics AnswersDocument13 pagesBioinformatics AnswersPratibha Patil100% (1)
- (Methods in Molecular Biology, 2231) Kazutaka Katoh - Multiple Sequence Alignment - Methods and Protocols-Humana (2020)Document322 pages(Methods in Molecular Biology, 2231) Kazutaka Katoh - Multiple Sequence Alignment - Methods and Protocols-Humana (2020)Diego AlmeidaNo ratings yet
- Bioinformatics for Beginners: Genes, Genomes, Molecular Evolution, Databases and Analytical ToolsFrom EverandBioinformatics for Beginners: Genes, Genomes, Molecular Evolution, Databases and Analytical ToolsRating: 4.5 out of 5 stars4.5/5 (3)
- Algorithms On String Trees and SequencesDocument326 pagesAlgorithms On String Trees and SequencesJason ShumNo ratings yet
- Pieper NucleicAcidsRes 2004Document6 pagesPieper NucleicAcidsRes 2004ctak2022No ratings yet
- Bioinformatics & Computational Biology SyllabusDocument2 pagesBioinformatics & Computational Biology SyllabusKamlesh SahuNo ratings yet
- Blast Analisis IIDocument15 pagesBlast Analisis IIUlises Ortiz GutierrezNo ratings yet
- Protein Structure Similarity: Mlesnick@stanford - EduDocument8 pagesProtein Structure Similarity: Mlesnick@stanford - EdualsreshtyNo ratings yet
- Blast NsuiteDocument19 pagesBlast Nsuitegeerthu99No ratings yet
- Introduction To Structural DatabasesDocument10 pagesIntroduction To Structural Databasessumit mahajanNo ratings yet
- Multiple Sequence AlignmentDocument6 pagesMultiple Sequence AlignmentGeovani Gregorio Rincon NavasNo ratings yet
- BLASTDocument4 pagesBLASTdia320576No ratings yet
- Lab Report 05Document20 pagesLab Report 05DewNo ratings yet
- Metamotifs - A Generative Model For Building Families of Nucleotide Position Weight MatricesDocument16 pagesMetamotifs - A Generative Model For Building Families of Nucleotide Position Weight Matricesanon_174376942No ratings yet
- Cavidad PrediccionDocument9 pagesCavidad PrediccionLuis MartinezNo ratings yet
- Davis NucleicAcidsRes 2006Document10 pagesDavis NucleicAcidsRes 2006huouinkyoumaNo ratings yet
- Delta Blast PDFDocument14 pagesDelta Blast PDFHark SranNo ratings yet
- BIOINFORMATICSDocument4 pagesBIOINFORMATICSCecilia MukototsiNo ratings yet
- Cyclic Peptide Structure Prediction and Design Using AlphaFoldDocument25 pagesCyclic Peptide Structure Prediction and Design Using AlphaFoldxinyuwu0528No ratings yet
- Mol - Modelling of BCL-2 FinalDocument37 pagesMol - Modelling of BCL-2 FinalVannakumar SubramanianNo ratings yet
- BMC Bioinformatics: RSEARCH: Finding Homologs of Single Structured RNA SequencesDocument16 pagesBMC Bioinformatics: RSEARCH: Finding Homologs of Single Structured RNA SequencesTheoNo ratings yet
- Nucl. Acids Res. 2005 Ginalski 1874 91Document18 pagesNucl. Acids Res. 2005 Ginalski 1874 91Rizki PerdanaNo ratings yet
- Automated Design of The Surface Positions of Protein HelicesDocument9 pagesAutomated Design of The Surface Positions of Protein Helicesaxva1663No ratings yet
- Sequence AnalysisDocument6 pagesSequence Analysisharrison9No ratings yet
- Protein3dstructureprediction 161109084221Document30 pagesProtein3dstructureprediction 161109084221Jhanvi SNo ratings yet
- UntitledDocument13 pagesUntitledDewNo ratings yet
- Protein Structure Prediction ThesisDocument8 pagesProtein Structure Prediction ThesisWriteMyThesisPaperManchester100% (2)
- COMPASS: A Tool For Comparison of Multiple Protein Alignments With Assessment of Statistical SignificanceDocument20 pagesCOMPASS: A Tool For Comparison of Multiple Protein Alignments With Assessment of Statistical SignificanceВалерий БжезинскийNo ratings yet
- Thesis On Homology ModelingDocument6 pagesThesis On Homology ModelingWriteMyPaperFastUK100% (2)
- Bioinformatics Notes - 17Bt54: Module - 4Document48 pagesBioinformatics Notes - 17Bt54: Module - 4Samuel PrinceNo ratings yet
- Efficient Protein Interaction Hotspot Identification in ProteinsDocument4 pagesEfficient Protein Interaction Hotspot Identification in ProteinsIJERDNo ratings yet
- NIH Public Access: Author ManuscriptDocument19 pagesNIH Public Access: Author Manuscriptantonio.rosato1No ratings yet
- Merin 1Document10 pagesMerin 1merineb002No ratings yet
- Eswar MethodsMolBiol 2008Document25 pagesEswar MethodsMolBiol 2008huouinkyoumaNo ratings yet
- Protein SequenceDocument36 pagesProtein SequenceHarshitNo ratings yet
- Lab Report 03Document18 pagesLab Report 03DewNo ratings yet
- 2013 J Theor Biol 328 77-88Document12 pages2013 J Theor Biol 328 77-88mbrylinskiNo ratings yet
- Homolgy ModelingDocument19 pagesHomolgy ModelingJainendra JainNo ratings yet
- Homology Modeling, Also Known As Comparative Modeling ofDocument19 pagesHomology Modeling, Also Known As Comparative Modeling ofManoharNo ratings yet
- Bioinfo MCQSDocument22 pagesBioinfo MCQSSamana Zaynab100% (1)
- Re Comb 2003Document16 pagesRe Comb 2003mustafaNo ratings yet
- BFG Chapter3 PairwiseAlignment v04Document86 pagesBFG Chapter3 PairwiseAlignment v04Divya AggarwalNo ratings yet
- Binc Syllabus For Paper-Iii Binc Bioinformatics Syllabus - BasicDocument7 pagesBinc Syllabus For Paper-Iii Binc Bioinformatics Syllabus - BasicMuhammed Irfan Ali KNo ratings yet
- Scansite 20 Proteome-Wide Prediction of Cell SignaDocument8 pagesScansite 20 Proteome-Wide Prediction of Cell SignaAarushi sharmaNo ratings yet
- Tertiary Structure Prediction Methods: Any Given Protein SequenceDocument29 pagesTertiary Structure Prediction Methods: Any Given Protein Sequencesurrender003No ratings yet
- Bioinfo Final PracticalDocument66 pagesBioinfo Final PracticalPolu ChattopadhyayNo ratings yet
- 2016 Article 141Document11 pages2016 Article 141Gopal GNo ratings yet
- Intro To Bioinformatics Semester 6 BotanyDocument15 pagesIntro To Bioinformatics Semester 6 BotanyMalabika BhowmickNo ratings yet
- ArticleDocument11 pagesArticleSahil8No ratings yet
- BlastDocument8 pagesBlastreza zeinNo ratings yet
- Towards The Self-Annotating Web: Philipp Cimiano, Siegfried Handschuh, Steffen StaabDocument10 pagesTowards The Self-Annotating Web: Philipp Cimiano, Siegfried Handschuh, Steffen StaabTrần ThiNo ratings yet
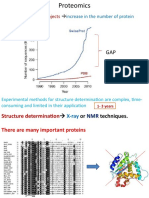
- Genome Sequencing Projects: Increase in The Number of Protein SequencesDocument27 pagesGenome Sequencing Projects: Increase in The Number of Protein SequencesBiju ThomasNo ratings yet
- Hat 1Document2 pagesHat 1kunchalaharikachinmayNo ratings yet
- Applied Physics - CO2 - ALM HA 2 13-12-2023Document1 pageApplied Physics - CO2 - ALM HA 2 13-12-2023kunchalaharikachinmayNo ratings yet
- Ha 2Document1 pageHa 2kunchalaharikachinmayNo ratings yet
- Applied Physics ALM HA 4Document1 pageApplied Physics ALM HA 4kunchalaharikachinmayNo ratings yet
- Handwriting 20240327 095334 Via 10015 IoDocument3 pagesHandwriting 20240327 095334 Via 10015 IokunchalaharikachinmayNo ratings yet
- 21uc2204 - Corporate Readiness SkillsDocument13 pages21uc2204 - Corporate Readiness SkillskunchalaharikachinmayNo ratings yet
- Ha 4Document1 pageHa 4kunchalaharikachinmayNo ratings yet