Professional Documents
Culture Documents
CMT WEGS Nejmoa0908094
Uploaded by
MCuk2606Original Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
CMT WEGS Nejmoa0908094
Uploaded by
MCuk2606Copyright:
Available Formats
The n e w e ng l a n d j o u r na l of m e dic i n e
original article
Whole-Genome Sequencing in a Patient
with Charcot–Marie–Tooth Neuropathy
James R. Lupski, M.D., Ph.D., Jeffrey G. Reid, Ph.D., Claudia Gonzaga-Jauregui, B.S.,
David Rio Deiros, B.S., David C.Y. Chen, M.Sc., Lynne Nazareth, Ph.D.,
Matthew Bainbridge, M.Sc., Huyen Dinh, B.S., Chyn Jing, M.Sc.,
David A. Wheeler, Ph.D., Amy L. McGuire, J.D., Ph.D., Feng Zhang, Ph.D.,
Pawel Stankiewicz, M.D., Ph.D., John J. Halperin, M.D., Chengyong Yang, Ph.D.,
Curtis Gehman, Ph.D., Danwei Guo, M.Sc., Rola K. Irikat, B.S., Warren Tom, B.S.,
Nick J. Fantin, B.S., Donna M. Muzny, M.Sc., and Richard A. Gibbs, Ph.D.
A BS T R AC T
BACKGROUND
Whole-genome sequencing may revolutionize medical diagnostics through rapid From the Department of Molecular and
identification of alleles that cause disease. However, even in cases with simple pat- Human Genetics (J.R.L., J.G.R., C.G.-J.,
M.B., F.Z., P.S., D.M.M., R.A.G.), the Hu-
terns of inheritance and unambiguous diagnoses, the relationship between disease man Genome Sequencing Center (J.G.R.,
phenotypes and their corresponding genetic changes can be complicated. Compre- D.R.D., D.C.Y.C., L.N., M.B., H.D., C.J.,
hensive diagnostic assays must therefore identify all possible DNA changes in each D.A.W., D.M.M., R.A.G.), the Center for
Medical Ethics and Health Policy (A.L.M.),
haplotype and determine which are responsible for the underlying disorder. The the Department of Pediatrics (J.R.L.), and
high number of rare, heterogeneous mutations present in all humans and the pau- Texas Children’s Hospital (J.R.L.) — all at
city of known functional variants in more than 90% of annotated genes make this Baylor College of Medicine, Houston;
Atlantic Neuroscience Institute, Summit,
challenge particularly difficult. Thus, the identification of the molecular basis of a NJ, and Mount Sinai School of Medicine,
genetic disease by means of whole-genome sequencing has remained elusive. We New York (J.J.H.); and Life Technologies,
therefore aimed to assess the usefulness of human whole-genome sequencing for Carlsbad, CA (C.Y., C.G., D.G., R.K.I., W.T.,
N.J.F.). Address reprint requests to Dr.
genetic diagnosis in a patient with Charcot–Marie–Tooth disease. Gibbs at 1 Baylor Plaza, Mail Stop BCM226,
Houston, TX 77030, or at agibbs@bcm
METHODS .edu.
We identified a family with a recessive form of Charcot–Marie–Tooth disease for This article (10.1056/NEJMoa0908094) was
which the genetic basis had not been identified. We sequenced the whole genome published on March 10, 2010, at NEJM.org.
of the proband, identified all potential functional variants in genes likely to be related
N Engl J Med 2010;362:1181-91.
to the disease, and genotyped these variants in the affected family members. Copyright © 2010 Massachusetts Medical Society.
RESULTS
We identified and validated compound, heterozygous, causative alleles in SH3TC2
(the SH3 domain and tetratricopeptide repeats 2 gene), involving two mutations, in
the proband and in family members affected by Charcot–Marie–Tooth disease. Sepa-
rate subclinical phenotypes segregated independently with each of the two muta-
tions; heterozygous mutations confer susceptibility to neuropathy, including the
carpal tunnel syndrome.
CONCLUSIONS
As shown in this study of a family with Charcot–Marie–Tooth disease, whole-genome
sequencing can identify clinically relevant variants and provide diagnostic informa-
tion to inform the care of patients.
n engl j med 362;13 nejm.org april 1, 2010 1181
The New England Journal of Medicine
Downloaded from nejm.org on May 17, 2023. For personal use only. No other uses without permission.
Copyright © 2010 Massachusetts Medical Society. All rights reserved.
The n e w e ng l a n d j o u r na l of m e dic i n e
T
he practice of medical genetics re- resulting in foot dorsiflexor weakness, foot drop,
quires gene-specific analyses of DNA se- and secondary steppage gait. Pes cavus (highly
quences and mutations to definitively diag- arched feet) or pes planus (flat feet) occurs in
nose disease, provide prognostic information, and most patients.
guide genetic counseling regarding the risk of re- We applied next-generation-sequencing meth-
currence. Studies of autosomal recessive traits ods to identify the cause of disease in a family
such as cystic fibrosis1 and some dominant traits with inherited neuropathy that had been previous
such as neurofibromatosis type 12 revealed the ly screened, with negative results, for alterations
role of single “disease genes” in conveying traits. of some common Charcot–Marie–Tooth genes,
However, many phenotypes of mendelian diseases including PMP22,11 MPZ, PRX, GDAP1, and EGR2.
(see the Glossary) are genetically heterogeneous:
causative mutations have been identified in more Me thods
than 100 genes for deafness and retinitis pig-
mentosa, for instance. Moreover, specific muta- Study Participants
tions may confer phenotypes that segregate as The study family consisted of four affected sib-
dominant, recessive, or even digenic3 or triallel- lings, four unaffected siblings, and an unaffected
ic4 traits. There is also ample evidence of modi- mother and father, all of whom provided written
fying loci in mendelian disorders.5,6 Thus, even informed consent for participation in the study.
when there are simple patterns of inheritance The study was approved by the institutional re-
in syndromes with a well-characterized patho- view board at Baylor College of Medicine. The di-
logic course, the underlying mutational events, agnosis of Charcot–Marie–Tooth type 1 disease
which need to be resolved for precise molecular in the proband and the three affected siblings was
diagnosis, within individual families may be based on the results of physical examination (dis-
complex. tal muscle weakness and wasting, pes cavus, and
Charcot–Marie–Tooth disease is an inherited absence of deep-tendon reflexes) and electrophys-
peripheral neuropathy with two forms: a demyeli- iological studies.
nating form (type 1) affecting the glia-derived
myelin and an axonal form (type 2) affecting the Neurophysiological Assessments
nerve axon. The two forms can be distinguished Neurophysiological studies consisted of a stan-
by means of electrophysiological or neuropatho- dard battery of nerve-conduction studies, includ-
logical studies. Charcot–Marie–Tooth disease has ing motor responses of the median, ulnar, tibial,
been used as a model disease to describe genetic and peroneal nerves with F-wave latencies; ortho-
heterogeneity, posit the relation of hereditary dromic median-, ulnar-, and sural-nerve sensory
pattern to clinical severity, and investigate the potentials; and bilateral tibial H-reflexes. When
relative importance of principal and modifying these studies revealed demyelinating features,
genes in determining human diseases.7,8 Mutant tests of blink reflexes were generally performed.
alleles underlying Charcot–Marie–Tooth disease Limbs were warmed to a temperature of at least
can segregate in an autosomal dominant, reces- 32°C in all instances. Demyelination was judged
sive, or X-linked manner (Fig. 1). Both single- to be present if conduction velocities were signif
base variants (single-nucleotide polymorphisms icantly slowed and the late-response latencies
[SNPs]) and copy-number variants,10 at 39 sepa- were substantially delayed. Median-nerve mono
rate loci, confer susceptibility to Charcot–Marie– neuropathy at the wrist was judged to be present
Tooth disease. Most of these susceptibility variants when there was prolonged motor terminal latency
cause dominant forms of the disease, although or slowed median-nerve sensory velocity with dis-
mutations in genes at 14 of the loci cause reces- proportionate slowing in the palm-to-wrist seg-
sive disease. ment, or both. The four affected subjects, all of
Adult-onset Charcot–Marie–Tooth disease is whom had diffuse slowing of conduction, were also
highly variable in presentation but is character- thought to have a median-nerve mononeuropathy
ized by distal symmetric polyneuropathy,9 with at the wrist, since the median-nerve motor termi-
slowly progressive distal muscle weakness and nal latency was much more prolonged than the
atrophy (particularly peroneal muscular atrophy) ulnar-nerve motor terminal latency (14.9 vs. 8.1,
1182 n engl j med 362;13 nejm.org april 1, 2010
The New England Journal of Medicine
Downloaded from nejm.org on May 17, 2023. For personal use only. No other uses without permission.
Copyright © 2010 Massachusetts Medical Society. All rights reserved.
Whole-Genome Sequencing and SH3TC2 Mutations
10.2 vs. 7.5, 11.6 vs. 6.2, and 9.2 vs. 6.2 msec) tides. Its accuracy in sequencing 50-base reads is
(Table 1). estimated at approximately 99.94%.12 Multiple se-
quences can be read simultaneously, and when
DNA Sequence Analysis the sequence reads overlap, the overall accuracy
DNA sequencing was performed with the use of increases further, reducing the risk of false posi-
the SOLiD (Sequencing by Oligonucleotide Liga- tive determinations and the need for additional
tion and Detection) system (Applied Biosystems), a data validation. We determined bases from the
next-generation-sequencing platform that involves primary sequencing data, using the standard
ligation-based sequencing and a two-base encod- SOLiD analysis software. (For details, see the Sup-
ing method in which four fluorescent dyes are plementary Appendix, available with the full text
used to tag various combinations of dinucleo of this article at NEJM.org.)
Glossary
Array-based comparative genomic hybridization: A hybridization method for detecting copy-number variations in DNA
samples from a patient as compared with a control sample. The method provides higher resolution than cytogenetic
methods but lower resolution than sequencing methods.
Average depth of coverage: The average number of times each base in the genome was sequenced, as a function of the
distribution and number of sequence reads that map to the reference genome.
Coding single-nucleotide polymorphisms: Single DNA-base changes that occur in the coding regions of genes.
Copy-number variation: DNA changes that involve sequences of more than 100 bp, larger than single-nucleotide changes
or microsatellites, and that vary in their number of copies among individual persons. These variants can be benign
and polymorphic, but some can cause disease.
DNA template: An individual fragment of DNA that is available for sequencing.
Exon capture: Methods for isolating and sequencing gene exons, to the exclusion of the remainder of the genome. The
DNA templates from exons are “captured” with the use of probes complementary to the targeted exon sequences.
After capture, the targeted DNA is eluted and sequenced. The cost of exon capture can be 10 to 50% lower than that of
whole-genome sequencing, although the method is insensitive to copy-number variations and mutations that are out-
side the targeted regions.
Fragment-sequence read: The contiguous nucleotide sequence from one end of a DNA template (as opposed to a mate-
pair read).
Haploinsufficiency: The state that occurs when a diploid organism has only a single functional copy of a gene, which
does not produce enough protein to support normal function.
Mapping: The computational process of identifying the specific region of a reference genome from which an individual
sequenced DNA template originated.
Mappable yield: The number of bases generated by a DNA-sequencing instrument that can be mapped to the reference
genome.
Mate-pair sequencing: A sequencing strategy that permits the inference of structural changes in a genome by sequenc-
ing at both the 5′ and 3′ ends of each DNA template (as opposed to the fragment-sequencing approach).
Mendelian disease: Human disease caused by mutations in a single gene.
Missense mutations: Single DNA-base changes that occur in the coding regions of genes and alter the resulting encoded
amino acid sequence.
Next-generation sequencing: DNA-sequencing methods that involve chemical assays other than the traditional Sanger
dideoxy-chain-termination method. Next-generation-sequencing methods produce much larger quantities of data
at less expense, but the individual raw sequence reads that are generated from individual amplified DNA-template
sequences are shorter and have lower quality.
Nonsense mutations: DNA-base changes that introduce termination codons in the coding sequences of genes, result-
ing in truncated proteins.
Sequence read: The sequence generated from a single DNA template.
Single-base error rate: The total number of mismatched bases found in mapped sequence reads from a sequencing
run, divided by the mappable yield. This rate estimates the probability that any given mappable base is an error.
Two-base encoding: A method used in the SOLiD (Sequencing by Oligonucleotide Ligation and Detection) DNA-sequencing
platform that represents a DNA sequence as a chain of overlapping dimers encoded as single-base “colors.” This
allows for sequencing of the 16 unique sequence dimers with the use of only four unique dye colors and provides a
method for improving the overall accuracy of the sequence reads (reducing the single-base error rate).
n engl j med 362;13 nejm.org april 1, 2010 1183
The New England Journal of Medicine
Downloaded from nejm.org on May 17, 2023. For personal use only. No other uses without permission.
Copyright © 2010 Massachusetts Medical Society. All rights reserved.
The n e w e ng l a n d j o u r na l of m e dic i n e
CMT phenotype
Nerve-conduction studies
and electromyography
Primary myelinopathy Primary axonopathy
(CMT type 1) (CMT type 2)
Dominant
Dominant Recessive X-linked Recessive Dominant
intermediate
10q24–q25
Xq24–q26
Xq26–q28
16p13
17p12
11p15
12p11
19p13
10q21
22q13
10q23
11q22
18q23
19q13
15q13
16q24
19q13
12q12
12q24
16q22
Xp22
Xq13
Xq21
1p35
8p23
1p36
7p14
8p21
1q22
5q32
6q21
8q21
8q24
3q13
1q21
3q21
7q11
GJB1
PRPS1
Unknown
Unknown
Unknown
LMNA
SLC12A6
GAN
Unknown
MPZ
EGR2
LITAF
PMP22
SOX10
YARS
Unknown
ARHGEF10
Unknown
DNM2
SH3TC2
FIG4
GDAP1
NDRG1
Unknown
SBF2
MTMR2
FGD4
CTDP1
PRX
MFN2 and KIF1B
RAB7A
GARS
HSPB1
NEFL
Unknown
HSPB8 and TRPV4
AARS
Figure 1. Charcot–Marie–Tooth (CMT) Disease Phenotypes, Their Genetic Forms of Inheritance, and Their Mapped
Genes and Loci.
AUTHOR: Gibbs (Lupski) RETAKE: 1st
CMT is divided in two major phenotypic types — glial myelinopathy (CMT type 1) 2nd and neuronal axonopathy (CMT
type 2) — according to electrophysiological,
FIGURE: 1 of 2clinical, and nerve-biopsy evaluations.3rd Each type can be inherited in a
dominant, recessive, or X-linked fashion. There are also autosomal dominant intermediate forms of CMT that can
Revised
ARTIST: ts
have features of both axonal and demyelinating neuropathies. Several genes SIZE have been associated with CMT disease
6 col
to date, and other loci have been associated
TYPE: Line and mapped
Combo but 4-CtheirH/T
genes not33p9yet identified. MPZ, GDAP1, and GJB1
are known to be associated with CMT type 1, but select mutations in these genes can also cause CMT type 2; NEFL
AUTHOR, PLEASE NOTE:
is known to be associated with CMT Figuretype 2,has
but select
been mutations
redrawn and typeconvey
has beena reset.
CMT type 1 phenotype. Dominant inter-
mediate forms of CMT have been reported to bePlease associated with MPZ mutations. Specific recessive alleles related
check carefully.
to CMT have also been reported for EGR2 and PMP22. Of the 31 genes in 39 known CMT loci, only 15 genes are cur-
JOB:Current
rently available for clinical testing. 36213 evidence-based clinical guidelines ISSUE:for04-01-10
distal symmetric polyneuropathy
recommend genetic testing consisting of screening for common mutations, including the CMT1A duplication copy-
number variant and point mutations of the X-linked GJB1 gene.9
Array-Based Comparative Genomic Analysis of the copy-number variants was per-
Hybridization formed according to the manufacturer’s instruc-
For array-based comparative genomic hybridiza- tions and software.
tion and analysis of the copy-number variants in
the proband as compared with those in a male Bioinformatic Analysis of SNP Variants
control, we used a 1-million-probe high-resolution Analysis of SNP variants and cross-referencing of
oligonucleotide whole-genome array (Agilent), a them with the Human Gene Mutation Database
2.1-million-oligonucleotide whole-genome array (www.hgmd.cf.ac.uk), the Online Mendelian In-
(NimbleGen), and a 44,000-oligonucleotide array heritance in Man database (www.ncbi.nlm.nih
(Agilent) that was custom-designed to assay genes .gov/omim), and the PolyPhen database (http://
previously implicated in inherited neuropathy. genetics.bwh.harvard.edu/pph/data, based on the
1184 n engl j med 362;13 nejm.org april 1, 2010
The New England Journal of Medicine
Downloaded from nejm.org on May 17, 2023. For personal use only. No other uses without permission.
Copyright © 2010 Massachusetts Medical Society. All rights reserved.
Whole-Genome Sequencing and SH3TC2 Mutations
Table 1. Neurophysiological Findings in the Study Family.*
Subject No. Age Sex Demyelination Axonopathy Median-Nerve Entrapment Diagnosis
yr
I-1 80 M No Motor: peroneal, 2.2 mV Yes
I-2 77 F No Sural: absent; motor: peroneal, Yes Axonal
0.5 mV; tibial, 2.8 mV neuropathy
Maternal grandmother 90 F No Normal Yes; SCV, 43 m/sec MMM
of proband
II-I 58 F No Normal Yes; SCV, 46 m/sec MMM
II-2 57 M No Sural: absent; motor: peroneal, Yes Axonal
0.2 mV; tibial, 1.4 mV; neuropathy
H-reflexes: 38 msec
III-1 37 M No Normal No
III-2 35 M Yes No Yes; median-nerve motor TL, 14.9 msec; CMT
ulnar-nerve motor TL, 8.1 msec
III-3 34 F No No Yes; SCV, 42 m/sec; median-nerve MMM
motor TL, 4.4 msec
III-4 (proband) 32 M Yes No Probably; median-nerve motor TL, CMT
10.2 msec; ulnar-nerve motor TL,
7.5 msec
III-5 31 F No No No
III-6 29 F Yes No Yes; median-nerve motor TL, 11.6 msec; CMT
ulnar-nerve motor TL, 6.2 msec
III-7 26 F No Motor: peroneal, 36 m/sec; Yes; SCV, 36 m/sec; median-nerve MMM
H-reflexes: 35 msec motor TL, 4.8 msec
III-8 25 M Yes No Yes; median-nerve motor TL, 9.2 msec; CMT
ulnar-nerve motor TL, 6.2 msec
* Subject I-1 was a carpenter for more than 50 years. The maternal grandmother of the proband is not included in the pedigree. The ages listed
are the ages at the time at which the subjects were evaluated. The normal value for median-nerve motor terminal latency (TL) is less than
4.0 msec and for sensory conduction velocity (SCV) is 48 m/sec or more. The diagnosis column lists the conclusion based on the aggregate
findings: severe, widespread slowing of conduction was interpreted as evidence of demyelination, and low-potential amplitudes in multiple
nerves with relative preservation of conduction velocities were interpreted as evidence of axonal damage. CMT denotes Charcot–Marie–
Tooth disease, CTS the carpal tunnel syndrome, and MMM mild mononeuropathy of the median nerve.
National Center for Biotechnology Information bers of the study family. To verify the Arg954ter
[NCBI] dbSNP, build 126) were performed with the amino acid mutation (R954X), corresponding to
use of Perl scripts. Alignment of the orthologous a G→A mutation in the genomic DNA in exon 11
SH3TC2 (SH3 domain and tetratricopeptide re- of SH3TC2 on chromosome 5 at nucleotide
peats 2) proteins was performed with the use of the 148,386,628, we also generated a 312-bp PCR
ClustalW program and reference SH3TC2 proteins fragment and incubated it with the restriction
from the following organisms: human (accession enzyme TaqI; the nucleotide mutation results in
number, NP_078853), chimpanzee (XP_527069), elimination of the restriction site for TaqI.
macaque (XP_001104761), dog (XP_546315),
horse (XP_001501607), cow (XP_616288), mouse R e sult s
(NP_766216), rat (XP_225887), opossum (XP_
001380773), and chicken (XP_424256). Nerve-Conduction Studies
In addition to the Charcot–Marie–Tooth type 1
Segregation Analysis phenotype that segregates as a recessive trait, we
Exons 5 and 11 of the SH3TC2 gene were ampli- identified through electrophysiological means an
fied by means of a polymerase-chain-reaction axonal neuropathy in one parent and one grand-
(PCR) assay and directly sequenced in all mem- parent of the proband. Further evidence of a sub-
n engl j med 362;13 nejm.org april 1, 2010 1185
The New England Journal of Medicine
Downloaded from nejm.org on May 17, 2023. For personal use only. No other uses without permission.
Copyright © 2010 Massachusetts Medical Society. All rights reserved.
The n e w e ng l a n d j o u r na l of m e dic i n e
Genome Variation
Table 2. SNPs Identified through Whole-Genome
Sequencing of DNA from the Proband.* The sequencing of DNA samples obtained from
the proband produced a mappable yield of 89.6 Gb
SNP Type No. of SNPs of sequence data, representing an average depth
Nongene 2,255,102 of coverage of approximately 30 times per base.
Gene 1,165,204 The data from sequential machine runs consisted
Intron 1,064,655 of 8.3 Gb of 35-bp fragment sequence reads (one
Promoter 60,075 run), 30.3 Gb of 25-bp mate-pair sequence reads
3′ UTR 16,350 (two runs), and 51.0 Gb of 50-bp mate-pair se-
5′ UTR 3,517 quence reads (one run).
Splice regulatory site 2,089 We identified the differences between the con-
Splice site 112 sensus sequence of the proband and the human
Synonymous 9,337
genome reference sequence. These were used to
produce a list of putative single-base DNA substi-
Stop→stop 17
tutions, small insertions, and deletions and po-
Nonsynonymous 9,069
tential changes in DNA copy number. This list
Stop→gain 121
of variants included 3,420,306 SNPs. A total of
Stop→loss 27
2,255,102 of the SNPs were in extragenic regions
Total 3,420,306 and 1,165,204 SNPs were within gene regions,
including introns, promoters, 3′ and 5′ untrans-
* Stop→stop refers to synonymous substitutions within a
stop codon that maintain the stop codon, stop→gain refers lated regions, and splice sites (Table 2). Of the
to nonsense mutations, and stop→loss refers to nonsynon- intragenic SNPs, 9069 were nonredundant SNPs
ymous substitutions that change a stop codon to any oth- predicted to result in nonsynonymous codon
er codon. SNP denotes single-nucleotide polymorphism,
and UTR untranslated region. changes, and 121 of the 9069 were nonsense
mutations. The approximately 3.4 million SNPs
identified represent about 0.1% of the reference
tle phenotype evidenced by, at a minimum, medi- haploid human genome,15 and both the total
an-nerve mononeuropathy at the wrist was also number of SNPs and the number of novel SNPs
observed among all the proband’s grandparents are similar to those discovered in other diploid
and both parents but had an unclear pattern of genome sequences for individual subjects (Table
inheritance. Its variable presentation (Table 1) in- 3).12,16-21 Of the more than 3.4 million SNPs,
cluded three neurophysiologically defined pheno- 2,858,587 were present in public databases and
types: a normal phenotype with superimposed se- 561,719 were novel (Table 3). Data on the se-
vere median-nerve mononeuropathy at the wrist, quence reads, quality, and mapping have been
thought to be an incidental finding in an 80-year- deposited in the NCBI Sequence Read Archive
old man who had been a carpenter for more (www.ncbi.nlm.nih.gov/Traces/sra/sra.cgi?) (acces-
than 50 years (Subject I-1), a mild median-nerve sion number, SRP001734); variant data have been
mononeuropathy at the wrist (the proband’s ma- deposited in the dbSNP database.
ternal grandmother and mother [Subject II-1]), and We used two approaches to identifying copy-
a more severe median-nerve mononeuropathy at number variation: array-based comparative ge-
the wrist associated with evidence of a more wide- nomic hybridization and mate-pair sequencing.
spread axonal polyneuropathy (Subjects I-2 and We identified 234 copy-number variants ranging
II-2). The latter phenotype is similar to that of in size from 1690 bp to 1,627,813 bp. Of these
patients with hereditary neuropathy with liability 234 variants, 132 were confirmed by at least one
to pressure palsies (Online Mendelian Inheri- other method (Table 1 in the Supplementary Ap-
tance in Man number, 162500), a disorder patho- pendix); 220 of the 234 (94%) overlap with re-
logically characterized by patchy myelin abnor- ported regions of copy-number variants in the
malities and attributed to haploinsufficiency of Database of Genomic Variants (http://projects.tcag
PMP22 (as a consequence of genomic deletion)13; .ca/variation). We found no copy-number variants
duplication of PMP22 causes Charcot–Marie–Tooth affecting genes known to be involved in Char-
type 1A disease, the most common form.14 cot–Marie–Tooth disease or other neuropathies.
1186 n engl j med 362;13 nejm.org april 1, 2010
The New England Journal of Medicine
Downloaded from nejm.org on May 17, 2023. For personal use only. No other uses without permission.
Copyright © 2010 Massachusetts Medical Society. All rights reserved.
Whole-Genome Sequencing and SH3TC2 Mutations
Table 3. Individual Human Genomes Sequenced to Date.*
Average Depth
Genome† Technology Used of Coverage SNPs
Total Known Novel
×10−6
Venter Sanger method 7.5 3.21 2.80 0.74
Watson 454 Sequencing System (Roche) 7.4 3.32 2.71 0.61
Chinese (YH) Genome Analyzer (Illumina) 36 3.07 2.67 0.39
African (NA18507) Genome Analyzer (Illumina) 40.6 3.61 2.72 0.88
African (NA18507) SOLiD system (Applied Biosystems) 17.9 3.86 3.13 0.73
Korean (SJK) Genome Analyzer (Illumina) 28.95 3.43 3.01 0.42
Korean (AK1) Genome Analyzer (Illumina) 27.8 3.45 2.88 0.57
Proband in this study SOLiD system (Applied Biosystems) 29.9 3.42 2.85 0.56
* All genomes listed have a ploidy of 2n. SNP denotes single-nucleotide polymorphism.
† The surname of the individual person or the ethnic group (and HapMap sample name, in parentheses) is given. The
same African (Yoruban) sample NA18507 was sequenced twice, once with the use of the Genome Analyzer and once
with the use of the SOLiD (Sequencing by Oligonucleotide Ligation and Detection) system.
We cross-referenced the nonsynonymous SNPs examination of 3148 putative SNPs, including 54
that we detected by using whole-genome sequenc- coding SNPs. Of these 54, 2 were at the SH3TC2
ing with a database of previously observed muta- locus — 1 missense mutation (identified at 7.7
tions implicated in human disease (the Human average depth coverage) and 1 nonsense muta-
Gene Mutation database) (Table 4, and Table 2 tion (identified at 29.9 average depth coverage)
in the Supplementary Appendix). Of the 174 non- (Fig. 1 in the Supplementary Appendix). Muta-
synonymous database SNPs identified in the tions in this locus have previously been found to
proband, 159 had a clear association with a be associated with Charcot–Marie–Tooth type 4C
heritable trait (i.e., the database entry was not disease, described in families of eastern Euro-
annotated with a question mark). Of these, 21 pean, Turkish or Spanish Gypsy origin.23-25 The
(13%) were described as causing mendelian dis- R954X nonsense mutation has previously been
ease; 16 were heterozygous in the proband, a implicated in Charcot–Marie–Tooth disease; the
finding that is consistent with the expected load missense mutation (A→G, occurring on chromo-
of autosomal recessive mutations. The other five some 5 at nucleotide 148,402,474 and corre-
SNPs might have been erroneously assigned as sponding to the amino acid mutation Tyr169His
disease mutations, which would explain why four [Y169H]) is novel.
of them were homozygous in the proband and
have been found to be homozygous in unaffected Correlation between Genotype
persons. It would also explain why the sequence and Phenotype
for the proband, who did not have adrenoleuko Segregation analyses verified independent mater-
dystrophy, contained a SNP in ABCD1 previously nal and paternal origins of the mutations (Fig. 2).
described as a dominant mutation that causes The nonsense mutation (R954X) appeared in one
the X-linked disorder adrenoleukodystrophy.22 An parent of the proband and in two siblings who
alternative to the interpretation that the five SNPs did not have Charcot–Marie–Tooth type 1 disease.
might have been erroneously assigned as disease The missense mutation (Y169H) was found in
mutations is that these alleles might have reduced one parent and one grandparent, neither of whom
penetrance. had Charcot–Marie–Tooth disease. Only the pro
We examined the putative mutations in 40 band (Subject III-4) and three of his siblings (Sub-
genes known to cause or be linked to neuro- jects III-2, III-6, and III-8) who had inherited both
pathic or related conditions (Table 3 in the Sup- mutant alleles had the Charcot–Marie–Tooth type 1
plementary Appendix). This exercise led to closer phenotype (Fig. 2).
n engl j med 362;13 nejm.org april 1, 2010 1187
The New England Journal of Medicine
Downloaded from nejm.org on May 17, 2023. For personal use only. No other uses without permission.
Copyright © 2010 Massachusetts Medical Society. All rights reserved.
The n e w e ng l a n d j o u r na l of m e dic i n e
density of samples on the sequencing slide in-
Table 4. Disease and Trait Associations of
Nonsynonymous SNPs Identified in the Proband, creased, and mapping technology improved. Over-
According to the Human Gene Mutation Database.* all, the sequence yield increased by a factor of
three, with no appreciable increase in expense.
Disease or Trait Associated with Mutation SNPs
This rapid pace of technological improvement
no. (%) makes it difficult to accurately determine the ex-
Total 159 (100) pense of repeating this experiment, but given that
Behavioral disorder 6 (4) the expense of sequencing reagents for a single
Cancer 33 (21) run on the SOLiD instrument was $25,000 in
Association 7
April 2009, we estimate that the entire effort
would currently cost less than $50,000.
Increased risk 9
The whole-genome sequencing approach used
Reduced risk 3
in this proband enabled us to identify the cause
Susceptibility 14 of his disease as compound heterozygous muta-
Complex disease 48 (30) tions in the SH3TC2 gene and thus to delineate
Mendelian disease 21 (13) the specific biologic basis of disease in his fam-
Metabolic trait 17 (11) ily. The SH3TC2 protein contains both SH3 and
Pharmacogenetic trait 14 (9)
TPR motifs; SH3 motifs mediate the assembly of
protein complexes binding to proline-rich pro-
Other traits 20 (13)
teins, and TPR motifs are involved in protein–
* SNP denotes single-nucleotide polymorphism. protein interactions.
The mouse orthologue of SH3TC2 is specifical
ly expressed in Schwann cells, and the SH3TC2
The subjects with the heterozygous missense protein localizes to the plasma membrane and
mutation (Y169H) (Subjects I-2 and II-2) (Fig. 2) to the perinuclear endocytic recycling compart-
also had the apparently dominant axonal neu- ment, which is consistent with a role in myelina-
ropathy phenotype, as detected by electrophysio- tion or in axon–glia interactions.28 Mice lacking
logical studies. These findings of axonal neuropa- Sh3tc2 have abnormal organization of the node
thy (Table 1) suggest a gain of function (i.e., a of Ranvier.28 Consistent with a role of SH3TC2 in
toxic effect) of this mutation. In contrast, the pre- endocytic processes29 is the finding that SH3TC2
sumed loss-of-function nonsense variant (R954X) mutations result in disruption of the endocytic
was associated with electrophysiological evidence and membrane recycling pathways.30
of the carpal tunnel syndrome, regardless of We observed that both of the SH3TC2 muta-
whether it was the sole mutation present (i.e., tions, when heterozygous, have phenotypic con-
heterozygous genotype) or was accompanied by sequences that can be detected by electrophysi-
the missense variant (Y169H) (i.e., compound ological means. The Y169H missense variant
heterozygous genotype) (Table 1 and Fig. 2). segregates with an axonal neuropathy, whereas
the nonsense R954X mutation is associated with
Discussion subclinical evidence of the carpal tunnel syn-
drome; therefore, haploinsufficiency of SH3TC2
We ascertained the molecular basis of an inher- may cause susceptibility to the carpal tunnel
ited disease by using next-generation-sequencing syndrome. This susceptibility may also result
methods. We chose whole-genome sequencing from mutations in other genes related to Charcot–
over targeted, exon-capture approaches26,27 be- Marie–Tooth disease in addition to PMP22 and
cause we did not know whether the “causative” SH3TC2. Whole-genome sequencing of other
mutations would reside in known coding ele- members of the proband’s family might help
ments, and targeted approaches are ill suited to clarify whether the additional 69 SNPs at the
capturing copy-number variants. The heteroge- SH3TC2 locus and 3146 SNPs at the other 39
neity of our sequence data is emblematic of the neuropathy-associated gene loci examined (in-
current rapid progress of sequencing technology: cluding many rare variants, 466 of which have
over the 6-month course of this study, sequence not previously been described [Table 3 in the
read lengths doubled (from 25 bp to 50 bp), the Supplementary Appendix]) can modify the highly
1188 n engl j med 362;13 nejm.org april 1, 2010
The New England Journal of Medicine
Downloaded from nejm.org on May 17, 2023. For personal use only. No other uses without permission.
Copyright © 2010 Massachusetts Medical Society. All rights reserved.
Whole-Genome Sequencing and SH3TC2 Mutations
A SH3TC2 Genotype and Phenotype
Axonal neuropathy
I
I-1 I-2 CMT1
+/+ +/Y169H Normal
II
II-1 II-2
R954X/+ +/Y169H
III
III-1 III-2 III-3 III-4 III-5 III-6 III-7 III-8
+/+ R954X/ R954X/+ R954X/ +/+ R954X/ R954X/+ R954X/
B Results of R954X Genotyping Y169H Y169H Y169H Y169H
G→A mutant
(R954X)
Wild type
C Sequence Alignment
Y169
Homo sapiens EHLLFDHKYWLNCILVEDTEIQVSVDDKHLETIYLGLLIQEGHFFCRALCSVTPPAEKEG-ECLTL
Pan troglodytes EHLLFDHKYWLNCILVEDTEIQVSVDDKHLETIYLGLLIQEGHFFCRALCSVTPPAEKEG-ECLTL
Macaca mulatta EHLLFDHKYWLNCILVEDTEIQVSVDDKHLETIYLGLLIQEGHFFCRALCSVIPPAEKEG-ECLTL
Canis familiaris EHLLFDHKYWLNCRLVEDTEIQVSVDEKHLETIYLALLIQEGHFFCRAMCSVAQPAEKEG-EYLTL
Equus caballus EHLLFDHKYWLNSRLVEDTEIQVSVDDKHLESIYLGLLIQEGHFFCRAMCSVAQPAEKEG-EYLTL
Bos taurus EHLLFDHKYWLNCRLVEDTEIHVSIDDKHLETIYLGLLIQEGHFFCRAMCSVAQPAEKEG-EYLTL
Mus musculus EHLFFDHTYWLNSRLVDDTEIQVSVDDNHLENIYLGLLLQEGHFFCRAVCSVAQPADKEG-EYLTL
Rattus norvegicus EHLFFDHTYWLNSRLVDDTEIQVSVDDTHLENIYLGLLLQEGHFFCRAMCSVTQPADKEG-EYLTL
Monodelphis domestica EQLLFDHNYWLNFRLVEDTKIQVIVNYEHLEAIYQSLLIQEGH-FCRTVHTVFRSGEKEGGEYLKL
Gallus gallus EQLLFEQEYWLNCALVEDTEIRVSMDENRLATIYLGLLLQEGHFFSRAVPGVCQPG-GEGQEGLQL
Figure 2. Pedigree of the Study Family and Segregation and Conservation of SH3TC2 Mutations.
Panel A shows the pedigree of the proband (arrow) and his family and their SH3TC2 genotypes: plus signs indicate
the wild-type allele; Y169H indicates the A→G mutation on chromosome 5 at nucleotide 148,402,474 and corre-
sponding to the amino acid missense mutation Tyr169His, and R954X indicates
AUTHOR: Gibbs (Lupski) RETAKE:
the1stG→A mutation in the genomic
DNA in exon 11 of SH3TC2 on chromosome 5 at nucleotide 148,386,628, leading to2nd the amino acid nonsense muta-
FIGURE:
tion Arg954ter. (Genomic coordinates for2 the
of 2 mutations in the proband are based on 3rd the human genome reference
sequence, build 36.2.) Squares indicate male subjects, and circles female Revised subjects; slashes indicate deceased sub-
ARTIST: ts
jects. Subjects in generations I and II had three phenotypes. The paternal grandfather SIZE (Subject I-1) was studied 20
6 col
years ago, at 80 years of age, and had normal
TYPE: Lineresults,
Combowith the
4-C soleH/T
exception33p9
of a median-nerve mononeuropathy at
the wrist, thought to be caused by his occupation as a carpenter. The paternal grandmother (Subject I-2, 77 years of
AUTHOR, PLEASE NOTE:
age at the time of evaluation) and theFigure
fatherhas(Subject II-2) had
been redrawn evidence
and type of areset.
has been patchy axonal polyneuropathy, with def-
inite median-nerve mononeuropathy at the wrist.Please check carefully.
The maternal grandmother (evaluated at 90 years of age; data not
shown) and the mother (Subject II-1) had normal findings except for very mild median-nerve mononeuropathy at
JOB: 36213 ISSUE: 04-01-10
the wrist. Two of the proband’s sisters (Subjects III-3 and III-7) had this same phenotype. Two members of this
generation had completely normal findings (Subjects III-1 and III-5). The other four siblings had diffuse, dispropor-
tionate conduction slowing in the distal median nerve, without evidence of conduction block, findings that are sug-
gestive of a superimposed median mononeuropathy at the wrist. Subjects III-2, III-4, III-6, and III-8 had Charcot–
Marie–Tooth type 1 (CMT1) disease. Panel B shows the results of TaqI restriction digestion of the SH3TC2 exon 11
polymerase-chain-reaction product on which the G→A mutation, corresponding to the R954X allele, occurs. This
mutation was present in the proband’s mother and six of the eight siblings, as well as in the maternal grandmother
(not shown). The mutation destroys the restriction site for TaqI; the wild type yields two small bands and the
heterozygous mutant yields three bands, the upper of which is the uncut DNA. Panel C shows sequence alignment
of the SH3TC2 protein among various species. The downward arrowhead indicates the location of the highly con-
served Tyr169 amino acid that, in persons with the novel missense mutation Y169H, is changed to His. Sequences
were obtained from the National Center for Biotechnology Information.
n engl j med 362;13 nejm.org april 1, 2010 1189
The New England Journal of Medicine
Downloaded from nejm.org on May 17, 2023. For personal use only. No other uses without permission.
Copyright © 2010 Massachusetts Medical Society. All rights reserved.
The n e w e ng l a n d j o u r na l of m e dic i n e
penetrant Y169H and R954X mutations and there- consider this approach for illuminating the mo-
by influence the neuropathy phenotype. lecular causes of these diseases. The approach
The whole-genome sequencing approach that may ultimately contribute to the care of patients
we describe here contrasts with other diagnostic and families living with such diseases.
approaches. A clinical-testing panel that screens Our results suggest that haploinsufficiency of
for a copy-number variant that commonly causes SH3TC2 confers predisposition to a mild polyneu-
Charcot–Marie–Tooth disease14 and nucleotide- ropathy with particular susceptibility to the carpal
sequence variants in 15 of the genes known to tunnel syndrome. More generally, they demon-
be mutated in patients with the disease can cost strate the diagnostic power of whole-genome se-
more than $15,000.31 Mutations in two or more quencing in the context of genetically heteroge-
genes related to Charcot–Marie–Tooth disease neous mendelian disease and inform efforts to
have been described as causing a phenotype more decipher the genetic bases of complex traits. As
severe than that of our proband or other patients new, rare alleles at other gene loci are implicated
affected by the disease.32-34 Such groups of mu- in conditions such as diabetes, obesity, heart dis-
tations include a combination of two SNPs at ease, and cancer and as the patterns of interac-
the ACBD1 locus and a copy-number variant af- tion of the alleles with a patient’s phenotype are
fecting PMP22, as well as the combination of a delineated, genetic susceptibility to such dis-
SNP and a copy-number variant at the same lo- eases may become clearer. As a practical matter,
cus.35,36 There is also a report of mutations in the identification of rare, heterogeneous alleles by
two genes related to Charcot–Marie–Tooth dis- means of whole-genome sequencing may be the
ease segregating in the same family as either a only way to definitively determine genetic contri-
recessive trait or a sporadic trait, the latter of butions to the associated clinical phenotypes.
which was attributed to a de novo copy-number Supported in part by grants from the National Human Ge-
variant.37 Given this locus heterogeneity, with nome Research Institute (5 U54 HG003273, to Dr. Gibbs) and the
evidence of a mutational load that has clinical National Institute of Neurological Disorders and Stroke (R01
NS058529, to Dr. Lupski).
consequences, as well as the ease of use and Disclosure forms provided by the authors are available with
accuracy of the whole-genome sequencing meth- the full text of this article at NEJM.org.
ods we applied, clinical and genetics experts We thank Kevin McKernan, Michael Rhodes, Francisco de la
Vega, Quynh Doan, and Fiona Hyland for extensive discussion
struggling to explain poorly understood high- and support and Cristian Coarfa for structural-variation analysis
penetrance genetic diseases can now seriously and insights.
References
1. Rommens JM, Iannuzzi MC, Kerem B, 7. Allan W. Relation of hereditary pat- 12. McKernan KJ, Peckham HE, Costa
et al. Identification of the cystic fibrosis tern to clinical severity as illustrated by GL, et al. Sequence and structural varia-
gene: chromosome walking and jumping. peroneal atrophy. Arch Intern Med 1939; tion in a human genome uncovered by
Science 1989;245:1059-65. 63:1123-31. short-read, massively parallel ligation se-
2. Wallace MR, Marchuk DA, Andersen 8. Haldane JBS. The relative importance quencing using two-base encoding. Ge-
LB, et al. Type 1 neurofibromatosis gene: of principal and modifying genes in deter- nome Res 2009;19:1527-41.
identification of a large transcript disrupt- mining some human diseases. J Genet 13. Del Colle R, Fabrizi GM, Turazzini M,
ed in three NF1 patients. Science 1990; 1941;41:149-57. Cavallaro T, Silvestri M, Rizzuto N. Heredi-
249:181-6. 9. England JD, Gronseth GS, Franklin G, tary neuropathy with liability to pressure
3. Kajiwara K, Berson EL, Dryja TP. Di- et al. Practice parameter: evaluation of dis- palsies: electrophysiological and genetic
genic retinitis pigmentosa due to muta- tal symmetric polyneuropathy: role of lab- study of a family with carpal tunnel syn-
tions at the unlinked peripherin/RDS and oratory and genetic testing (an evidence- drome as only clinical manifestation.
ROM1 loci. Science 1994;264:1604-8. based review): report of the American Neurol Sci 2003;24:57-60.
4. Katsanis N, Ansley SJ, Badano JL, et al. Academy of Neurology, American Associ- 14. Lupski JR, de Oca-Luna RM, Slaugen-
Triallelic inheritance in Bardet-Biedl syn- ation of Neuromuscular and Electrodiag- haupt S, et al. DNA duplication associated
drome, a Mendelian recessive disorder. nostic Medicine, and American Academy with Charcot-Marie-Tooth disease type 1A.
Science 2001;293:2256-9. of Physical Medicine and Rehabilitation. Cell 1991;66:219-32.
5. Dipple KM, McCabe ERB. Phenotypes Neurology 2009;72:185-92. 15. International Human Genome Sequenc
of patients with “simple” Mendelian dis- 10. Lupski JR. Structural variation in the ing Consortium. Finishing the euchromat-
orders are complex traits: thresholds, mod- human genome. N Engl J Med 2007;356: ic sequence of the human genome. Nature
ifiers, and systems dynamics. Am J Hum 1169-71. 2004;431:931-45.
Genet 2000;66:1729-35. 11. Roa BB, Garcia CA, Suter U, et al. 16. Levy S, Sutton G, Ng PC, et al. The
6. Badano JL, Katsanis N. Beyond Men- Charcot–Marie–Tooth disease type 1A: diploid genome sequence of an individual
del: an evolving view of human genetic association with a spontaneous point mu- human. PLoS Biol 2007;5(10):e254.
disease transmission. Nat Rev Genet tation in the PMP22 gene. N Engl J Med 17. Wheeler DA, Srinivasan M, Egholm M,
2002;3:779-89. 1993;329:96-101. et al. The complete genome of an indi-
1190 n engl j med 362;13 nejm.org april 1, 2010
The New England Journal of Medicine
Downloaded from nejm.org on May 17, 2023. For personal use only. No other uses without permission.
Copyright © 2010 Massachusetts Medical Society. All rights reserved.
Whole-Genome Sequencing and SH3TC2 Mutations
vidual by massively parallel DNA sequenc- 25. Gooding R, Colomer J, King R, et al. A 32. Chung KW, Sunwoo IN, Kim SM, et al.
ing. Nature 2008;452:872-6. novel Gypsy founder mutation, p.Arg1109X Two missense mutations of EGR2 R359W
18. Bentley DR, Balasubramanian S, Swerd- in the CMT4C gene, causes variable periph- and GJB1 V136A in a Charcot-Marie-
low HP, et al. Accurate whole human ge- eral neuropathy phenotypes. J Med Genet Tooth disease family. Neurogenetics 2005;
nome sequencing using reversible termi- 2005;42(12):e69. 6:159-63.
nator chemistry. Nature 2008;456:53-9. 26. Albert TJ, Molla MN, Muzny DM, et al. 33. Auer-Grumbach M, Fischer C, Papić L,
19. Wang J, Wang W, Li R, et al. The dip- Direct selection of human genomic loci et al. Two novel mutations in the GDAP1
loid genome sequence of an Asian indi- by microarray hybridization. Nat Methods and PRX genes in early onset Charcot-
vidual. Nature 2008;456:60-5. 2007;4:903-5. Marie-Tooth syndrome. Neuropediatrics
20. Ahn SM, Kim TH, Lee S, et al. The 27. Ng SB, Buckingham KJ, Lee C, et al. 2008;39:33-8.
first Korean genome sequence and analy- Exome sequencing identifies the cause of 34. Hodapp JA, Carter GT, Lipe HP, Michel-
sis: full genome sequencing for a socio- a mendelian disorder. Nat Genet 2010;42: son SJ, Kraft GH, Bird TD. Double trouble
ethnic group. Genome Res 2009;19:1622-9. 30-5. in hereditary neuropathy: concomitant
21. Kim JI, Ju YS, Park H, et al. A highly 28. Arnaud E, Zenker J, de Preux Charles mutations in the PMP-22 gene and another
annotated whole-genome sequence of a Ko- A-S, et al. SH3TC2/KIAA1985 protein is gene produce novel phenotypes. Arch
rean individual. Nature 2009;460:1011-5. required for proper myelination and the Neurol 2006;63:112-7.
22. Dvoráková L, Storkánová G, Unter- integrity of the node of Ranvier in the pe- 35. Roa BB, Garcia CA, Pentao L, et al.
rainer G, et al. Eight novel ABCD1 gene ripheral nervous system. Proc Natl Acad Evidence for a recessive PMP22 point mu-
mutations and three polymorphisms in Sci U S A 2009;106:17528-33. tation in Charcot-Marie-Tooth disease
patients with X-linked adrenoleukodystro- 29. Roberts RC, Peden AA, Buss F, et al. type 1A. Nat Genet 1993;5:189-94.
phy: the first polymorphism causing an Mistargeting of SH3TC2 away from the re- 36. Shy ME, Scavina MT, Clark A, et al.
amino acid exchange. Hum Mutat 2001; cycling endosome causes Charcot-Marie- T118M PMP22 mutation causes partial
18:52-60. Tooth disease type 4C. Hum Mol Genet loss of function and HNPP-like neuropa-
23. Senderek J, Bergmann C, Stendel C, et 2010;19:1009-18. thy. Ann Neurol 2006;59:358-64.
al. Mutations in a gene encoding a novel 30. Lupo V, Galindo M, Martínez-Rubio 37. Verny C, Ravisé N, Leutenegger AL,
SH3/TPR domain protein cause autoso D, et al. Mutations in the SH3TC2 protein et al. Coincidence of two genetic forms of
mal recessive Charcot-Marie-Tooth type 4C causing Charcot-Marie-Tooth disease type Charcot-Marie-Tooth disease in a single
neuropathy. Am J Hum Genet 2003;73: 4C affect its localization in the plasma family. Neurology 2004;63:1527-9.
1106-19. membrane and endocytic pathway. Hum Copyright © 2010 Massachusetts Medical Society.
24. Azzedine H, Ravisé N, Verny C, et al. Mol Genet 2009;18:4603-14.
Spine deformities in Charcot-Marie-Tooth 31. Athena Diagnostics home page. (Ac-
4C caused by SH3TC2 gene mutations. cessed March 5, 2010, at http://www
Neurology 2006;67:602-6. .athenadiagnostics.com.)
receive immediate notification when
a journal article is released early
To be notified when an article is released early
on the Web and to receive the table of contents
of the Journal by e-mail every Wednesday evening,
sign up through our Web site at
NEJM.org.
n engl j med 362;13 nejm.org april 1, 2010 1191
The New England Journal of Medicine
Downloaded from nejm.org on May 17, 2023. For personal use only. No other uses without permission.
Copyright © 2010 Massachusetts Medical Society. All rights reserved.
You might also like
- Auto-Inflammatory Syndromes: Pathophysiology, Diagnosis, and ManagementFrom EverandAuto-Inflammatory Syndromes: Pathophysiology, Diagnosis, and ManagementPetros EfthimiouNo ratings yet
- Genetics of Endocrinology: Amit R. Majithia - David Altshuler - Joel N. HirschhornDocument20 pagesGenetics of Endocrinology: Amit R. Majithia - David Altshuler - Joel N. HirschhornDr Mehul Kumar ChourasiaNo ratings yet
- 2013 Clinical WES Nejmoa1306555Document10 pages2013 Clinical WES Nejmoa1306555MCuk2606No ratings yet
- Fgf13 Causes Synaptic Excitatory-Inhibitory: Disruption of Imbalance and Genetic Epilepsy and Febrile Seizures PlusDocument16 pagesFgf13 Causes Synaptic Excitatory-Inhibitory: Disruption of Imbalance and Genetic Epilepsy and Febrile Seizures Plushidayatul rahmoNo ratings yet
- Variant of TYR and Autoimmunity Susceptibility Loci in Generalized VitiligoDocument12 pagesVariant of TYR and Autoimmunity Susceptibility Loci in Generalized VitiligoIvo AfianiNo ratings yet
- The Disruption of Profiling of Serotonergic Neurons, Results in Autism-Related BehaviorsDocument22 pagesThe Disruption of Profiling of Serotonergic Neurons, Results in Autism-Related BehaviorsAngelinni Taglioni StangeNo ratings yet
- Nej Mo A 1516767Document11 pagesNej Mo A 1516767mayrapp16No ratings yet
- Pleiotropy in Complex Traits: Challenges and StrategiesDocument13 pagesPleiotropy in Complex Traits: Challenges and Strategiesthe invincibleNo ratings yet
- Foundations, Promises and Uncertainties of Personalized MedicineDocument7 pagesFoundations, Promises and Uncertainties of Personalized MedicineVeronica BalanNo ratings yet
- Charcot Marie Tooth DiseaseDocument9 pagesCharcot Marie Tooth DiseaseMihai CebotariNo ratings yet
- From Mendel To The Human Genome Project: The Implications For Nurse EducationDocument6 pagesFrom Mendel To The Human Genome Project: The Implications For Nurse EducationKome ČemuNo ratings yet
- Kotlyar 2020Document16 pagesKotlyar 2020Merab KvintradzeNo ratings yet
- Lehmann Werman 2016Document9 pagesLehmann Werman 2016Felipe MNo ratings yet
- Germline Mutations in Predisposition Genes in Pediatric CancerDocument11 pagesGermline Mutations in Predisposition Genes in Pediatric CancerGuadalupeA.OsorioNo ratings yet
- Fced GenetikDocument33 pagesFced GenetikyosuanugrahapratamaNo ratings yet
- E472 FullDocument17 pagesE472 FullC_DanteNo ratings yet
- A Haplotype Map of The Human GenomeDocument57 pagesA Haplotype Map of The Human GenomeJanina PuenteNo ratings yet
- Part 16 - Genes, The Environment, and DiseaseDocument226 pagesPart 16 - Genes, The Environment, and Diseasenklinh.y2021No ratings yet
- ALSDocument14 pagesALSZé CunhaNo ratings yet
- Sudden Infant Death Syndrome: Case-Control Frequency Differences at Genes Pertinent To Early Autonomic Nervous System Embryologic DevelopmentDocument5 pagesSudden Infant Death Syndrome: Case-Control Frequency Differences at Genes Pertinent To Early Autonomic Nervous System Embryologic DevelopmentJailyn Carrasquel HerreraNo ratings yet
- EditorialDocument2 pagesEditorialsebastoto778No ratings yet
- 1 Biomarkers Ruggeri Et Al 2013Document16 pages1 Biomarkers Ruggeri Et Al 2013sri309No ratings yet
- 33 PDFDocument4 pages33 PDFThelma CastlleNo ratings yet
- Greenberg1996 PDFDocument8 pagesGreenberg1996 PDFAraNo ratings yet
- Genetic Association Studies: Design, Analysis and InterpretationDocument8 pagesGenetic Association Studies: Design, Analysis and InterpretationfaizahNo ratings yet
- Panel1 GenomicMedicineAnUpdatedPrimer PDFDocument11 pagesPanel1 GenomicMedicineAnUpdatedPrimer PDFMariana ÁlvarezNo ratings yet
- Role For Hedgehog Signaling in Cranial-Suture Development and ObesityDocument9 pagesRole For Hedgehog Signaling in Cranial-Suture Development and ObesityDanny Alvarez FocacciNo ratings yet
- Pkan NejmDocument8 pagesPkan Nejmmartin.ruda93No ratings yet
- How To Interpret A GWASDocument10 pagesHow To Interpret A GWAStpatel0986No ratings yet
- Gene Hunting: J. PongraczDocument15 pagesGene Hunting: J. PongracztahirnaeemkhanNo ratings yet
- This Content Downloaded From 120.126.70.24 On Wed, 29 Mar 2023 11:59:10 UTCDocument8 pagesThis Content Downloaded From 120.126.70.24 On Wed, 29 Mar 2023 11:59:10 UTCmarcia suNo ratings yet
- 1 s2.0 S0888754321002251 MainDocument11 pages1 s2.0 S0888754321002251 Mainhilay.koseNo ratings yet
- Nrneurol 2011 65Document2 pagesNrneurol 2011 65lauravshNo ratings yet
- 1 s2.0 S2589004221008932 MainDocument11 pages1 s2.0 S2589004221008932 MainBen DresimNo ratings yet
- Artigo Cientifico Charcot Marie ToothDocument8 pagesArtigo Cientifico Charcot Marie ToothCleverson Da Silva MattosNo ratings yet
- Primer: Down SyndromeDocument20 pagesPrimer: Down SyndromeangelinaNo ratings yet
- Forward (2017)Document13 pagesForward (2017)DanieleRibeiroNo ratings yet
- ElderlyDocument1 pageElderlyapi-26340035No ratings yet
- Genomewide Scan of Hoarding in Sib Pairs in Which Both Sibs Have Gilles de La Tourette SyndromeDocument9 pagesGenomewide Scan of Hoarding in Sib Pairs in Which Both Sibs Have Gilles de La Tourette SyndromeCita Tri KusumaNo ratings yet
- Mutations in The MORC2 Gene Cause AxonalDocument11 pagesMutations in The MORC2 Gene Cause AxonalMILON KUMAR HORENo ratings yet
- Family History of Alzheimer's Disease and Cortical Thickness in Patients With DementiaDocument7 pagesFamily History of Alzheimer's Disease and Cortical Thickness in Patients With DementiaMarcos Josue Costa DiasNo ratings yet
- Gomez Ospina2016Document11 pagesGomez Ospina2016christian roblesNo ratings yet
- Progressive Increase of The Mutated Mitochondria1 DNA Fraction in Kearns-Sayre SyndromeDocument6 pagesProgressive Increase of The Mutated Mitochondria1 DNA Fraction in Kearns-Sayre SyndromeGréta BotyánszkiNo ratings yet
- Zheng 2013Document6 pagesZheng 2013namiraNo ratings yet
- Principles of Pharmacogenetics-Implications For The AnaesthetistDocument11 pagesPrinciples of Pharmacogenetics-Implications For The AnaesthetistLulu SetiyabudiNo ratings yet
- Female Predominance in Intermittent Exotropia: ConclusionsDocument2 pagesFemale Predominance in Intermittent Exotropia: ConclusionsIkmal ShahromNo ratings yet
- 2020 - Autism Genetics. Over 100 Risk Genes and Counting - ForrestDocument1 page2020 - Autism Genetics. Over 100 Risk Genes and Counting - ForrestJesus RiveraNo ratings yet
- Clinical Interpretation and Management of Genetic VariantsDocument14 pagesClinical Interpretation and Management of Genetic VariantsAnastasija ZuravlovaNo ratings yet
- Current Concepts Review: Ph. Debeer, L. de Smet, W. J. M. Van de Ven, G. Fabry, J.-P. FrynsDocument12 pagesCurrent Concepts Review: Ph. Debeer, L. de Smet, W. J. M. Van de Ven, G. Fabry, J.-P. FrynsDr. Pedro Javier Cadena GonzálezNo ratings yet
- Autismo EstudioDocument41 pagesAutismo Estudiomauricio lopezNo ratings yet
- Zheng Hong 郑 红 Department of Medical Genetics & Cell BiologyDocument63 pagesZheng Hong 郑 红 Department of Medical Genetics & Cell BiologyinakiNo ratings yet
- Brochure Genetics-The Basic FactsDocument7 pagesBrochure Genetics-The Basic FactsVale FlociaNo ratings yet
- Genomewide Association Study of Leprosy: Original ArticleDocument10 pagesGenomewide Association Study of Leprosy: Original ArticleFickry AdiansyahNo ratings yet
- 5 Ensuring Equity Diversity and Inclusiveness in Genetic Analysis Will Empower The Future ofDocument4 pages5 Ensuring Equity Diversity and Inclusiveness in Genetic Analysis Will Empower The Future ofabdeali hazariNo ratings yet
- Guimier Et Al. (2015) PMID 26437028 - 0Document14 pagesGuimier Et Al. (2015) PMID 26437028 - 0Filip MilošićNo ratings yet
- Background: The Genetic of AnticipationDocument3 pagesBackground: The Genetic of AnticipationAh LynNo ratings yet
- Melkersson Rosenthal Sindrome Reporte Caso InglesDocument4 pagesMelkersson Rosenthal Sindrome Reporte Caso InglesOscar Cabezas CalderónNo ratings yet
- Genetics 0727Document9 pagesGenetics 0727Implant Surgical GuidesNo ratings yet
- Journal Pre-Proof: Biological PsychiatryDocument38 pagesJournal Pre-Proof: Biological PsychiatryTakelooNo ratings yet
- Genome Sequencing in The ClinicDocument17 pagesGenome Sequencing in The ClinicChristopher NorrieNo ratings yet
- 5 Scherloc SlikaDocument1 page5 Scherloc SlikaMCuk2606No ratings yet
- 3 Scherloc SlikaDocument1 page3 Scherloc SlikaMCuk2606No ratings yet
- 2 Scherloc SlikaDocument1 page2 Scherloc SlikaMCuk2606No ratings yet
- Alfa 1 Antitripsin Manjak Review 2020 10.1056@NEJMra1910234Document13 pagesAlfa 1 Antitripsin Manjak Review 2020 10.1056@NEJMra1910234MCuk2606No ratings yet
- CRISPR-Cas9 Gene Editing for Sickle Cell Disease and β-ThalassemiaDocument9 pagesCRISPR-Cas9 Gene Editing for Sickle Cell Disease and β-ThalassemiadrsourabhsinghNo ratings yet
- Ga1 Outcome Treatment Kevine Strauss 2020Document16 pagesGa1 Outcome Treatment Kevine Strauss 2020MCuk2606No ratings yet
- 647-Article Text-964-1-10-20200103Document4 pages647-Article Text-964-1-10-20200103MCuk2606No ratings yet
- 647-Article Text-964-1-10-20200103Document4 pages647-Article Text-964-1-10-20200103MCuk2606No ratings yet
- Biological Induced CorrosionDocument22 pagesBiological Induced CorrosionHemanth100% (1)
- A New Species of Riverine Crab of The Genus Sundathelphusa Bott, 1969 (Crustacea: Brachyura: Gecarcinucidae) From Northeastern Luzon, PhilippinesDocument10 pagesA New Species of Riverine Crab of The Genus Sundathelphusa Bott, 1969 (Crustacea: Brachyura: Gecarcinucidae) From Northeastern Luzon, PhilippinesNel YagosNo ratings yet
- BIO 328 Full Semester PackageDocument25 pagesBIO 328 Full Semester PackageNerdy Notes Inc.No ratings yet
- Syllabus CompletionDocument1 pageSyllabus CompletiongopimicroNo ratings yet
- Study Guide Cardio Tayang by Gextha 30 Maret 2015Document82 pagesStudy Guide Cardio Tayang by Gextha 30 Maret 2015Adi ParamarthaNo ratings yet
- Pulse Flour From Wheat MillerDocument23 pagesPulse Flour From Wheat MillerCao Trọng HiếuNo ratings yet
- ZIZKA Bromeliaceae ChileDocument22 pagesZIZKA Bromeliaceae ChileJoaquín Eduardo Sepúlveda AstudilloNo ratings yet
- Angiogenesis and Direct Myocardial RevascularizationDocument364 pagesAngiogenesis and Direct Myocardial RevascularizationPerdana SidaurukNo ratings yet
- CAPE Bio Mark SchemeDocument4 pagesCAPE Bio Mark Schemeron97150% (2)
- EASL 2021 Version 4 NewDocument691 pagesEASL 2021 Version 4 NewGupse Köroğlu AdalıNo ratings yet
- Personal Development Lesson 2Document54 pagesPersonal Development Lesson 2Ysay FranciscoNo ratings yet
- RPT SC Year 2 (DLP) 2022-2023Document21 pagesRPT SC Year 2 (DLP) 2022-2023ANNIE MARGARET A/P ANTHONY MoeNo ratings yet
- The Differences Between Asexual and Sexual ReproductionDocument2 pagesThe Differences Between Asexual and Sexual ReproductionAmirul IkhwanNo ratings yet
- Chapter 1Document4 pagesChapter 1Nikoleta RudnikNo ratings yet
- Gender and PsychotherapyDocument12 pagesGender and PsychotherapyEhh EnNaNo ratings yet
- Kimia Daun PandanDocument4 pagesKimia Daun Pandanlutfi_alhayathullahNo ratings yet
- OB Ultrasound Report Template 2Document1 pageOB Ultrasound Report Template 2PriyankaNo ratings yet
- Selina Solutions For Class 9 Biology Chapter 2 Cell The Unit of LifeDocument15 pagesSelina Solutions For Class 9 Biology Chapter 2 Cell The Unit of LifePrashun PriyadarshiNo ratings yet
- 7 Pearson SquareDocument19 pages7 Pearson Squareapi-235944649No ratings yet
- Sri Roth 2000Document11 pagesSri Roth 2000ottoojuniiorNo ratings yet
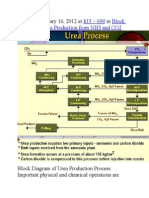
- Published January 16, 2012 at In: 813 × 699 Block Diagram of Urea Production From NH3 and CO2Document9 pagesPublished January 16, 2012 at In: 813 × 699 Block Diagram of Urea Production From NH3 and CO2himanshuchawla654No ratings yet
- AP Biology Practice Exam: Section I: Multiple-Choice QuestionsDocument83 pagesAP Biology Practice Exam: Section I: Multiple-Choice QuestionsKamilla DzhanzakovaNo ratings yet
- Manage Your Personal Energy PDFDocument46 pagesManage Your Personal Energy PDFezhil manickamNo ratings yet
- NCERT Class 3 English Baloon ManDocument10 pagesNCERT Class 3 English Baloon ManAnonymous 9WyPyismNo ratings yet
- Chemical Equilibrium and Le Chatelier's Principle: Chemistry 1Document17 pagesChemical Equilibrium and Le Chatelier's Principle: Chemistry 1azamatNo ratings yet
- Microscopy and StainingDocument7 pagesMicroscopy and StainingDenmark ManlusocNo ratings yet
- Human Genetics Concepts and Applications 11Th Edition Ricki Lewis Test Bank Full Chapter PDFDocument36 pagesHuman Genetics Concepts and Applications 11Th Edition Ricki Lewis Test Bank Full Chapter PDFdavid.cordero230100% (14)
- 6.1 MicrobiologyDocument32 pages6.1 MicrobiologyYsabelle BautistaNo ratings yet
- Prelim QuizDocument3 pagesPrelim QuizXyriel MacoyNo ratings yet
- Approved LaboratoriesDocument22 pagesApproved LaboratoriesPaul CNo ratings yet