Professional Documents
Culture Documents
Kamaljeet Singh (19103052) : DF Datasets Iris
Uploaded by
Jagrit SharmaOriginal Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
Kamaljeet Singh (19103052) : DF Datasets Iris
Uploaded by
Jagrit SharmaCopyright:
Available Formats
Kamaljeet Singh (19103052)
Loading the dataset
In [4]: df<-datasets::iris
Checking the dataset
In [6]: head(df,10)
Sepal.Length Sepal.Width Petal.Length Petal.Width Species
5.1 3.5 1.4 0.2 setosa
4.9 3.0 1.4 0.2 setosa
4.7 3.2 1.3 0.2 setosa
4.6 3.1 1.5 0.2 setosa
5.0 3.6 1.4 0.2 setosa
5.4 3.9 1.7 0.4 setosa
4.6 3.4 1.4 0.3 setosa
5.0 3.4 1.5 0.2 setosa
4.4 2.9 1.4 0.2 setosa
4.9 3.1 1.5 0.1 setosa
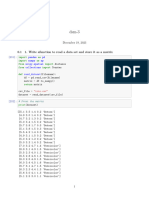
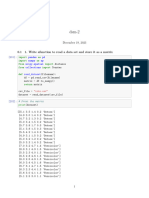
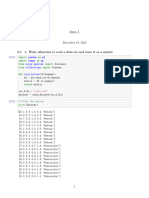
Function to split the dataset into train and test set
Ratio--> 0.3 means 30% rows used as test set
In [7]: train_test_split<-function(df,test_ratio){
set.seed(1)
split<- sample(c(rep(0, (1-test_ratio) * nrow(df)), rep(1, test_ratio * nrow(df
))))
return(split)
Function to calculate Euclidean Distance
In [8]: euclidean_dist<-function(train_row,test_row){
dist<-sum((test_row[1:4]-train_row[1:4])**2)
return(dist)
Function to calculate the euclidean distance of one test instance wrt all the training instances
In [9]: calc_distances<-function(train_df,test_row){
distances<-c(euclidean_dist(train_df[1,],test_row),euclidean_dist(train_df[2,],te
st_row))
classes<-c(as.numeric(train_df[1,][5]),as.numeric(train_df[2,][5]))
n<-nrow(train_df)
for(i in 3:n){
curr_dist=euclidean_dist(train_df[i,],test_row)
curr_class=as.numeric(train_df[i,][5])
distances<-append(distances,curr_dist)
classes<-append(classes,curr_class)
distances=cbind(data.frame(distances), data.frame(classes))
return(distances)
Function to sort the distances and then selecting the majority class with minimum distance
In [11]: predict_class<-function(train_df,test_row,k){
distances<-calc_distances(train_df,test_row)
distances<-distances[order(distances[,1]),]
dist<-table(distances[1:k,2])
class=names(dist)[which.max(dist)]
return(class)
Function to perform knn for all test instances
In [12]: knn<-function(train_df,test_df,k){
n<-nrow(test_df)
predictions<-list()
for(i in 1:n){
curr_pred<-predict_class(train_df,test_df[i,],k)
predictions[length(predictions)+1]<-curr_pred
return(predictions)
Function to calculate accuracy
In [13]: calc_accuracy<-function(test_classes,pred){
correct<-sum(test_classes==pred)
return(correct/length(test_classes))
Creating the train and test sets form the performed split
In [14]: split<-train_test_split(df,0.3)
train_df<-df[split==0,]
test_df<-df[split==1,]
Checking the train and test sets
In [15]: head(train_df,5)
Sepal.Length Sepal.Width Petal.Length Petal.Width Species
1 5.1 3.5 1.4 0.2 setosa
3 4.7 3.2 1.3 0.2 setosa
4 4.6 3.1 1.5 0.2 setosa
5 5.0 3.6 1.4 0.2 setosa
6 5.4 3.9 1.7 0.4 setosa
In [16]: head(test_df,5)
Sepal.Length Sepal.Width Petal.Length Petal.Width Species
2 4.9 3.0 1.4 0.2 setosa
8 5.0 3.4 1.5 0.2 setosa
15 5.8 4.0 1.2 0.2 setosa
17 5.4 3.9 1.3 0.4 setosa
18 5.1 3.5 1.4 0.3 setosa
Funtion to change the classes from numeric to string in the predicted values
In [17]: correct_classes<-function(pred){
n<-length(pred)
for(i in 1:n){
if(pred[i]=='1')
pred[i]='setosa'
else if(pred[i]=='2')
pred[i]='versicolor'
else
pred[i]='virginica'
return(pred)
Performing KNN with k-value as 5
In [18]: pred<-knn(train_df,test_df,5)
pred=correct_classes(pred)
In [19]: pred
1. 'setosa'
2. 'setosa'
3. 'setosa'
4. 'setosa'
5. 'setosa'
6. 'setosa'
7. 'setosa'
8. 'setosa'
9. 'setosa'
10. 'setosa'
11. 'setosa'
12. 'setosa'
13. 'setosa'
14. 'setosa'
15. 'versicolor'
16. 'versicolor'
17. 'versicolor'
18. 'versicolor'
19. 'versicolor'
20. 'versicolor'
21. 'versicolor'
22. 'versicolor'
23. 'versicolor'
24. 'virginica'
25. 'versicolor'
26. 'virginica'
27. 'versicolor'
28. 'versicolor'
29. 'versicolor'
30. 'versicolor'
31. 'versicolor'
32. 'versicolor'
33. 'virginica'
34. 'versicolor'
35. 'virginica'
36. 'virginica'
37. 'virginica'
38. 'virginica'
39. 'virginica'
40. 'virginica'
41. 'virginica'
42. 'virginica'
43. 'virginica'
44. 'virginica'
45. 'virginica'
In [20]: calc_accuracy(test_df$Species,pred)
0.933333333333333
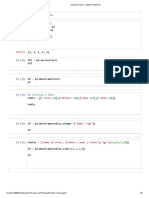
Perform knn for odd k values from 3 to 31 and then comparing accuracies
In [ ]: k_values<-seq(3,31,by=2)
accuracies<-list()
for(i in k_values){
preds=knn(train_df,test_df,i)
preds=correct_classes(preds)
accuracy=calc_accuracy(test_df$Species,preds)
accuracies<-append(accuracies,accuracy)
accuracies
In [ ]:
You might also like
- Data Cleaning and Pre Processing 2Document27 pagesData Cleaning and Pre Processing 2NipuniNo ratings yet
- AP19110010321 MLDocument4 pagesAP19110010321 MLSaijanith PNo ratings yet
- R ProgramsDocument30 pagesR ProgramsRanga SwamyNo ratings yet
- ML 1Document4 pagesML 1yefigoh133No ratings yet
- Import As Import As: "Iris - CSV"Document4 pagesImport As Import As: "Iris - CSV"avinashNo ratings yet
- 2 - 9 - KNN CodeDocument6 pages2 - 9 - KNN CodeYahya SabriNo ratings yet
- Decision TreeDocument1 pageDecision Treesid-harth sid-harth100% (1)
- Week11 N9Document5 pagesWeek11 N920131A05N9 SRUTHIK THOKALANo ratings yet
- Saurabh Verma 9919102005Document11 pagesSaurabh Verma 9919102005Yogendra pratap Singh100% (1)
- Assignment No - 10Document3 pagesAssignment No - 10Sid ChabukswarNo ratings yet
- 22MCA1008 - Varun ML LAB ASSIGNMENTSDocument41 pages22MCA1008 - Varun ML LAB ASSIGNMENTSS Varun (RA1931241020133)100% (1)
- Outliers, Hypothesis and Natural Language ProcessingDocument7 pagesOutliers, Hypothesis and Natural Language Processingsubhajitbasak001100% (1)
- Assignment 10Document9 pagesAssignment 10SHREYANSHU KODILKARNo ratings yet
- Matplotlib Arya ManasDocument44 pagesMatplotlib Arya ManasFarook Muhammad AliNo ratings yet
- # Common Datatype: Print Type Print Type Print Type Print Type Print TypeDocument4 pages# Common Datatype: Print Type Print Type Print Type Print Type Print TypeAYUSH KUMARNo ratings yet
- Week 6.2Document9 pagesWeek 6.2I.A.P. JABH RIDETNo ratings yet
- ТА5 Узунов А.Д. АИ-193Document32 pagesТА5 Узунов А.Д. АИ-193АИ-193 Узунов АртемNo ratings yet
- Module 2 Iris Data SetDocument1 pageModule 2 Iris Data SetRachell Ann UsonNo ratings yet
- Pandas Day 1Document3 pagesPandas Day 1Ali KhokharNo ratings yet
- 03 - K Means Clustering On Iris DatasetsDocument4 pages03 - K Means Clustering On Iris DatasetsJohn WickNo ratings yet
- 1 2 3 4 5 6 7 8 9 10 Sepallength Sepalwidth Petallength Petalwidth SpeciesDocument4 pages1 2 3 4 5 6 7 8 9 10 Sepallength Sepalwidth Petallength Petalwidth SpeciesLINH VU TRANGNo ratings yet
- Set2.py - Jupyter NotebookDocument3 pagesSet2.py - Jupyter Notebooktabish.khan.daNo ratings yet
- Pandas - Jupyter NotebookDocument32 pagesPandas - Jupyter NotebookLahari BanalaNo ratings yet
- Using R For Data Preprocessing, Exploratory Analysis, VisualizationDocument7 pagesUsing R For Data Preprocessing, Exploratory Analysis, VisualizationNikita DesaiNo ratings yet
- DSM 3Document6 pagesDSM 3no0r32200No ratings yet
- Lab - 5 (CB - En.u4ece22115)Document5 pagesLab - 5 (CB - En.u4ece22115)Daejuswaram GopinathNo ratings yet
- Sklearn - Datasets Pandas PD Sklearn - Model - Selection Graphviz Sklearn - Datasets Sklearn Sklearn Sklearn - Metrics Sklearn - Metrics Sklearn - Preprocessing Sklearn - Tree SklearnDocument4 pagesSklearn - Datasets Pandas PD Sklearn - Model - Selection Graphviz Sklearn - Datasets Sklearn Sklearn Sklearn - Metrics Sklearn - Metrics Sklearn - Preprocessing Sklearn - Tree SklearnNguyen Trung ThinhNo ratings yet
- Clustering - Jupyter NotebookDocument11 pagesClustering - Jupyter Notebookreema dsouzaNo ratings yet
- Lab7.ipynb - ColaboratoryDocument5 pagesLab7.ipynb - ColaboratoryPRAGASM PROG100% (1)
- SC Assignment Q2Document7 pagesSC Assignment Q2RAGHUVAMSHI -2019BCSIIITKNo ratings yet
- Pertemuan 8 (BAB 7)Document7 pagesPertemuan 8 (BAB 7)Rifa'i RaufNo ratings yet
- MH3511Document8 pagesMH3511Alex0% (1)
- MINI PROJECT - MergedDocument8 pagesMINI PROJECT - Mergedbhanucse16No ratings yet
- Part A Assignment 10Document3 pagesPart A Assignment 10B49 Pravin TeliNo ratings yet
- DSM 2Document7 pagesDSM 2no0r32200No ratings yet
- Frequency 1Document56 pagesFrequency 1YUviNo ratings yet
- PCA and FA CommandsDocument13 pagesPCA and FA CommandsNguyễn OanhNo ratings yet
- R Introduction by Deepayan SarkarDocument23 pagesR Introduction by Deepayan SarkarSunil AravaNo ratings yet
- From 10-14Document2 pagesFrom 10-14moinNo ratings yet
- Grid Search For SVMDocument9 pagesGrid Search For SVMkPrasad8No ratings yet
- Solutions To Pandas Basic QuestionsDocument1 pageSolutions To Pandas Basic QuestionsJason ShaxNo ratings yet
- Lab Wk1soln PDFDocument14 pagesLab Wk1soln PDFPeper12345No ratings yet
- 2 Pandas Basics Practice 2 PDFDocument1 page2 Pandas Basics Practice 2 PDFShubham TagalpallewarNo ratings yet
- Mathallcodes 1Document32 pagesMathallcodes 1NAGNo ratings yet
- Assignment-7 (CSL5402) Name:-Kommireddy - Manikanta Sai Roll No: - 1906036 Branch: - CSE-1Document7 pagesAssignment-7 (CSL5402) Name:-Kommireddy - Manikanta Sai Roll No: - 1906036 Branch: - CSE-1Manikanta Sai100% (1)
- R Project 1Document36 pagesR Project 1AlvinBurhaniNo ratings yet
- Assignment-7 (CSL5402) Name:-Kommireddy - Manikanta Sai Roll No: - 1906036 Branch: - CSE-1Document7 pagesAssignment-7 (CSL5402) Name:-Kommireddy - Manikanta Sai Roll No: - 1906036 Branch: - CSE-1Manikanta Sai100% (1)
- Experiment 3 J.626Document13 pagesExperiment 3 J.626ITA426 SOHAM LENDENo ratings yet
- R PracticeDocument38 pagesR PracticeyoshitaNo ratings yet
- R Intro STAT5000Document17 pagesR Intro STAT5000vinay kumarNo ratings yet
- R Programming: 122AD0029 - T.MANISHDocument21 pagesR Programming: 122AD0029 - T.MANISHManish McNo ratings yet
- 3a Data Frame - Jupyter NotebookDocument5 pages3a Data Frame - Jupyter Notebookvenkatesh mNo ratings yet
- Lecture 22Document64 pagesLecture 22Tev WallaceNo ratings yet
- Aiml Lab Manual 2023Document17 pagesAiml Lab Manual 2023shamilie17No ratings yet
- R QuestionsDocument5 pagesR QuestionsNatiqNo ratings yet
- Experiment - 2 Misc.Document4 pagesExperiment - 2 Misc.Siffatjot SinghNo ratings yet
- DSM 1Document6 pagesDSM 1no0r32200No ratings yet
- BDA pr2Document2 pagesBDA pr2Dev PathakNo ratings yet
- Logistic RegressionDocument4 pagesLogistic RegressionParamita HalderNo ratings yet
- Advanced Computer Networks - Laboratory - 4 Q)Document7 pagesAdvanced Computer Networks - Laboratory - 4 Q)Jagrit SharmaNo ratings yet
- ACN Lab - 3Document4 pagesACN Lab - 3Jagrit SharmaNo ratings yet
- ACN - Lab - 2Document4 pagesACN - Lab - 2Jagrit SharmaNo ratings yet
- Congrats Samsung Research Bangalore 16871Document1 pageCongrats Samsung Research Bangalore 16871Jagrit SharmaNo ratings yet
- Plant PropagationDocument2 pagesPlant PropagationMarie Grace SuarezNo ratings yet
- 1 - Mycology - BEE - Highest RSDocument117 pages1 - Mycology - BEE - Highest RSHoàng ĐứcAnhNo ratings yet
- Bisexual & Male Flower PDFDocument10 pagesBisexual & Male Flower PDFIcsNo ratings yet
- Determination of Seed Viability of Eight Wild SaudDocument8 pagesDetermination of Seed Viability of Eight Wild SaudtinNo ratings yet
- Smart Test Series: 1. Circle The Correct Answer. (12x1 12)Document2 pagesSmart Test Series: 1. Circle The Correct Answer. (12x1 12)Adnan mirzaNo ratings yet
- 4 Pandza 002Document28 pages4 Pandza 002Mademoiselle QuatreNo ratings yet
- Principles in Crop ProductionDocument76 pagesPrinciples in Crop ProductionBobby AnoraNo ratings yet
- Chapter-4 Growing Plants Question Bank With Answer KeyDocument21 pagesChapter-4 Growing Plants Question Bank With Answer KeySUHANEERIYA100% (1)
- Tree Names TamilDocument20 pagesTree Names Tamilsubikshababu0% (1)
- Evolving Ideas On The Origin and Evolution of FlowersDocument11 pagesEvolving Ideas On The Origin and Evolution of FlowersLuisa GonzálezNo ratings yet
- Pest of Rice CropDocument66 pagesPest of Rice CropGeet Bishnoi100% (1)
- Instant Download Blueprint Reading For Welders 9th Edition Bennett Test Bank PDF Full ChapterDocument32 pagesInstant Download Blueprint Reading For Welders 9th Edition Bennett Test Bank PDF Full ChapterGabrielPattersonatcd100% (9)
- Topic 1:: Types of Textile Fibers - List of Textile Fibers by Its SourcesDocument9 pagesTopic 1:: Types of Textile Fibers - List of Textile Fibers by Its SourcesKAWSER RAFINo ratings yet
- Trees: Drought Tolerant Species ListDocument3 pagesTrees: Drought Tolerant Species ListAbdul WajidNo ratings yet
- Banana Disease 2 PDFDocument4 pagesBanana Disease 2 PDFAlimohammad YavariNo ratings yet
- GrowNote Sorghum North 04 PhysiologyDocument5 pagesGrowNote Sorghum North 04 PhysiologyUMARNo ratings yet
- Grade 7 Comparing Flowering PlantsDocument19 pagesGrade 7 Comparing Flowering PlantsGadgetGlitchKillNo ratings yet
- Cyperus RotundusDocument2 pagesCyperus RotundusaiktiplarNo ratings yet
- Philippine Wood: By: John Ralph A. MagbanuaDocument19 pagesPhilippine Wood: By: John Ralph A. MagbanuaJohn Ralph A. Magbanua100% (1)
- FernDocument16 pagesFernx456456456xNo ratings yet
- Pictorial Guide For Tree MaintenanceDocument1 pagePictorial Guide For Tree Maintenancecity kathyNo ratings yet
- Fruits ClassificationDocument22 pagesFruits Classificationdudeperfecta36No ratings yet
- Wood IdentificationDocument3 pagesWood IdentificationLyu JHNo ratings yet
- Matthew DebaccoDocument91 pagesMatthew DebaccokatyweymNo ratings yet
- FTS - 10 (Code-A) - 22-04-2020 PDFDocument16 pagesFTS - 10 (Code-A) - 22-04-2020 PDFkavyareddyNo ratings yet
- Hama Penyakit Utama Tanaman Bawang Merah (Allium Ascalonicum) Dan Tindakan Pengendalian Di Brebes, Jawa Tengah (Allium Ascalonicum)Document6 pagesHama Penyakit Utama Tanaman Bawang Merah (Allium Ascalonicum) Dan Tindakan Pengendalian Di Brebes, Jawa Tengah (Allium Ascalonicum)Muhammad UmarNo ratings yet
- LAB7 IntroDocument3 pagesLAB7 IntroAngela Joyce MartinNo ratings yet
- Condiciones Óptimas para Almacenamiento de PolenDocument8 pagesCondiciones Óptimas para Almacenamiento de PolenBilly SteelNo ratings yet
- Study of Phytochemical Composition On Kaempferia Parviflora Wall Ex BakerDocument9 pagesStudy of Phytochemical Composition On Kaempferia Parviflora Wall Ex BakerKumaresh PalNo ratings yet
- Vydehi School of Excellence: EnglishDocument86 pagesVydehi School of Excellence: Englishsivasankar nNo ratings yet