Professional Documents
Culture Documents
7 - Example Student Final Poster
Uploaded by
api-3318643680 ratings0% found this document useful (0 votes)
18 views1 pageOriginal Title
7 - example student final poster
Copyright
© © All Rights Reserved
Available Formats
PDF, TXT or read online from Scribd
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
© All Rights Reserved
Available Formats
Download as PDF, TXT or read online from Scribd
0 ratings0% found this document useful (0 votes)
18 views1 page7 - Example Student Final Poster
Uploaded by
api-331864368Copyright:
© All Rights Reserved
Available Formats
Download as PDF, TXT or read online from Scribd
You are on page 1of 1
Effect of Human Maintenance on Soil Microbiome Antibiotic
Resistance in Dry Environments
Manish Bhatta, Olivia He, Arman Ostadazim, Michael Shi, and Anna Yang
1Oxford College of Emory University, Oxford, GA
Background Methods Results Continued
Soil microbiomes contain diverse antibiotic resistance genes. Sample Collection and Processing
The antibiotic resistance genes inside many soil microbiome • Soil samples were collected from various sampling locations (Table 1) (Figure 1). 4) Dry, human maintained colonies had higher antibiotic
communities are crucial in the field of biology, as they can • Each sample was shaken with 0.5 mL 1X PBS buffer to release bacteria cells from soil particles, resistance levels
influence plant growth, human health, and agricultural practices and the bacteria was incubated at room temperature for 48 hours on an agar plate.
which affect our food source.1 Antibiotic resistance genes not
only affect our food source, but also influence evolutionary
paths of the microbiome as well as other interacting organisms
in the ecosystem. Since there is sufficient research conducted on
antibiotic resistance in microbiomes, our study aims to analyze
the antibiotic resistance specifically in human-maintained
microbiomes in dry soil and compare our findings to the
antibiotic resistance in non-human maintained microbiomes in Analyzing Morphological Traits Analyzing Molecular Traits
dry soil. Human maintenance introduces factors such as Measuring Morphological Diversity (Figure 2) Analyzing Nitrogen-Fixing Genes (Figure 4)
fertilizer, pesticides, and industrial outputs that can potentially • Colonies were distinguished by pigmentation, • Separate PCRs were conducted for amplifying
lead to an increased presence of antibiotic resistance genes whole colony shape, margin shape, and surface nifH genes, amoA genes, and 16S rDNA.
through selection and gene transfer.1 The dryness of the soil is appearance. • A secondary PCR was run on each product for
also important to take into account because previous studies Calculating Bacterial Abundance (Figure 3) further amplification, which were run on gel
have found that soil moisture was significantly correlated (p • A transparency grid with 163 squares was laid electrophoresis.
<0.05) with tetracycline and ciprofloxacin resistance on top of each plate to estimate the percent Sequencing16S rDNA for Molecular Analysis
levels.2 Thus, our research question for this project is, do dry abundance of each colony type. • The 16S rDNA products were sent to the Conclusions and Future Directions
soil microbiome communities in human-maintained Measuring Antibiotic Resistance (Figure 4) Georgia Genomics and Bioinformatics Core
environments have higher or lower levels of antibiotic • Zone of inhibition was measured using ImageJ facility for sequencing. Conclusions
resistance than those in non-human maintained environments? after two antibiotic disks were placed into agar • The sequences were analyzed using NCBI’s
The findings from this study provides us with critical plates each with one colony type and incubated BLAST and the Ribosomal Database Project •Average zone of inhibition (ZOI) of colonies cultured from
information about how to maintain soil microbiomes in at room temperature for 24-48 hours. (RDP). dry, non-human maintained soil was approximately 0.15 cm
common dry environments, such as residential and industrial larger than that of dry, human maintained soil.
areas, that will influence our agriculture and health. •All but one human maintained sample had a higher ZOI than
Objectives & Hypothesis the smallest ZOI of the non-human maintained. The highest
ZOI in our dataset also came from human maintained soil. This
Objectives: indicates certain non-human maintained bacteria may have
• To compare the effect of human maintenance on soil had more resistant antibiotic genes.
microbiome bacterial diversity and abundance, antibiotic •The distribution of colony abundance was lower in the dry,
resistance, nitrogen fixation, and 16S rDNA sequences. human-maintained soil than in the dry, non-human maintained
• To understand the impact of human maintenance on soil Results soil, suggesting that certain colonies in human-maintained
microbiome antibiotic resistance. environments may dominate among other bacterial colonies
Hypothesis: We hypothesize that human-maintained 1) Dry, human maintained had less distinct colonies and higher colony abundance present.
environments will have higher soil microbiome antibiotic •Our data is in accordance with a study investigating the
resistance than non-human maintained environments. Multiple impact of manure-borne microorganisms on antibiotic
studies have shown that human use and application of manure resistance; manure-borne microorganisms were found to have
significantly increased antibiotic resistance gene presence in dry contributed largely to the elevation of antibiotic resistance
soil microbiomes. 3,4 Studies have also revealed genes due to both the addition of “manure-borne antibiotic
that surrounding human activities such as industry, resistant bacteria (ARB) in soil and potential horizontal gene
manufacturing, and utility use in urban areas increase antibiotic transfer via mobile genetic elements from manure-borne ARB
resistance presence. 5 to indigenous soil microorganisms". 3
•The lack of distribution of colony types in non-human
References maintained soil may be attributed to pesticide contamination,
1. Martinez, J. L. “Environmental Pollution by Antibiotics and or other substances entering the soil as a result of human
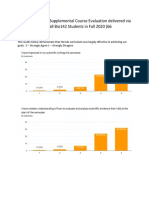
by Antibiotic Resistance Determinants.” Environmental Pollution, 2009, Figure 6. Percent colony abundance for all bacterial colonies identified in dry, Human maintained sample soil maintenance. Studies have found that bacterial communities in
vol. 157(11), pp. 2893-2902. Doi: 10.1016/j.envpol.2009.05.051 microbiome sites (A) and dry, non-human maintained sample soil microbiome sites (B).
2. Negreanu, Y., Pasternak, Z., Jurkevitch, E., & Cytryn, E. (2012).
pesticide contaminated soil experience a reduction of
Impact of Treated Wastewater Irrigation on Antibiotic Resistance in 2) Dry, human maintained had more colonies 3) Sequence identification and abundance of microbial diversity, while enriching only
Agricultural Soils. Environmental Science & Technology, 46(9), 4800- with nitrogen fixation gene presence morphological characteristics of specific communities. Meanwhile, bacterial communities in
4808. doi:10.1021/es204665b bacterial colonies non-pesticide contaminated soil were found to have abundance
3. Chen, Q. L., X. L. An, H. Li, Y. G. Zhu, J. Q. Su, and L. Cui. “Do among a larger number of phyla. 6
Manure-Borne or Indigenous Soil Microorganisms Influence the Spread
of Antibiotic Resistance Genes in Manured Soil?” Soil Biology & Future Directions
Biochemistry, 2017, vol. 114, pp. 229-237.
4. Pu, Q., L. X. Zhao, Y. T. Li, and J. Q. Su. “Manure Fertilization • Comparing the antibiotic resistance of a specific bacterial
Increase Antibiotic Resistance in Soils from Typical Greenhouse colony in non-human versus human maintained soil (bacterial
Vegetable Production Bases, China.” Journal of Hazardous Materials, species is a controlled variable)
2020, vol. 391.
• Comparing antibiotic resistance of colonies from similar
5. Yan, Z. Z., Q. L. Chen, Y. J. Zhang, J. Z. He, and H. W. Hu.
“Antibiotic Resistance in Urban Green Spaces Mirrors the Pattern of environments but different maintenance (nonhuman versus
Industrial Distribution.” Environmental International, 2019, vol. 132. human maintained) — type of area is a controlled variable
Doi: 10.1016/j.envint.2019.105106. • Further examine trends in percent abundance/distribution of
6. Regar, R.K., Gaur, V.K., et al. Science of The Total Environment, colonies cultured from human maintained versus non-human
2019; 1(681): p. 413-423. maintained.
You might also like
- Stau Dinger 2011Document9 pagesStau Dinger 2011inforNo ratings yet
- HHS Public Access: Interactions Between The Microbiota and The Immune SystemDocument16 pagesHHS Public Access: Interactions Between The Microbiota and The Immune Systemsandra pabonNo ratings yet
- Characterization of Soil Microorganism From Humus and Indigenous Microorganism AmendmentsDocument8 pagesCharacterization of Soil Microorganism From Humus and Indigenous Microorganism AmendmentsrahmanNo ratings yet
- Sociomicrobiology Biofilms and Quorum SensingDocument42 pagesSociomicrobiology Biofilms and Quorum Sensingtummalapalli venkateswara raoNo ratings yet
- Whipps Et Al, 2008Document12 pagesWhipps Et Al, 2008joyeeta8No ratings yet
- Amon Et Sanderson 2017 - What Is The MicrobiomeDocument5 pagesAmon Et Sanderson 2017 - What Is The MicrobiomeLaura DecockNo ratings yet
- 2008 Article 4 PDFDocument6 pages2008 Article 4 PDFnilaNo ratings yet
- STSrevDocument5 pagesSTSrevAnn Margarette De LeonNo ratings yet
- Perspectives and Vision For Strain Selection in BioaugmentationDocument4 pagesPerspectives and Vision For Strain Selection in BioaugmentationAndrew SingerNo ratings yet
- Biodiv 3 Human2020Document44 pagesBiodiv 3 Human2020Amina ArnautNo ratings yet
- Review: Genomic Approaches To Studying The Human MicrobiotaDocument7 pagesReview: Genomic Approaches To Studying The Human MicrobiotaJing XueNo ratings yet
- Clin Bacte Lec - Host Microorganism InteractionsDocument6 pagesClin Bacte Lec - Host Microorganism InteractionsnikkiNo ratings yet
- 1 Metagenomics Principles and Applications PPintoDocument44 pages1 Metagenomics Principles and Applications PPintograhamhollandpattersonNo ratings yet
- Sts FinalsDocument4 pagesSts Finals7xnc4st2g8No ratings yet
- MicroPara ReviewerDocument22 pagesMicroPara ReviewerAina HaravataNo ratings yet
- Rhizomicrobiome Engineering Through Indigenous Biofertilizers Toward Climate Resilient Sustainable Agriculture IndonesiaDocument46 pagesRhizomicrobiome Engineering Through Indigenous Biofertilizers Toward Climate Resilient Sustainable Agriculture IndonesiaDea NirmalaNo ratings yet
- HHS Public Access: Structure and Function of The Human Skin MicrobiomeDocument20 pagesHHS Public Access: Structure and Function of The Human Skin MicrobiomeEugeniaNo ratings yet
- Prelim MicrobioDocument8 pagesPrelim MicrobioAmalyn OmarNo ratings yet
- Lesson 1Document2 pagesLesson 1Ordeniza Llanz MherNo ratings yet
- DermatosisDocument8 pagesDermatosisDⒶrk OtⒶkuNo ratings yet
- Developments in Microbiology: Learning ObjectivesDocument289 pagesDevelopments in Microbiology: Learning Objectivesலலிதா மீனாட்சிசுந்தரம்No ratings yet
- Disease Suppression by Mycorrhizal FungiDocument68 pagesDisease Suppression by Mycorrhizal FungiMuhammad MushtaqNo ratings yet
- Microbiome JournalDocument14 pagesMicrobiome JournalTiara Rachmaputeri AriantoNo ratings yet
- Micro LecDocument26 pagesMicro Lecangelshizu0No ratings yet
- A Screening Tool For Quality of Mycorrhizal Bio FertilizerMycorrhiza Inoculum PotentialDocument4 pagesA Screening Tool For Quality of Mycorrhizal Bio FertilizerMycorrhiza Inoculum PotentialarunknbioNo ratings yet
- 1 s2.0 S2452231718300058 MainDocument10 pages1 s2.0 S2452231718300058 MainNahida Chowdhury ChainaNo ratings yet
- 1 Human Microbiome (A)Document11 pages1 Human Microbiome (A)Lenard MerlinNo ratings yet
- Minireview: Sami Aito Eishi Keda Iroshi Zura Iwamu InamisawaDocument13 pagesMinireview: Sami Aito Eishi Keda Iroshi Zura Iwamu InamisawaBekele OljiraNo ratings yet
- Microbes in Human WelfareDocument58 pagesMicrobes in Human WelfareAYUSHI SINGHNo ratings yet
- A Preliminary Evaluation of The Potential of Beauveria Bassiana For Bed Bug ControlDocument4 pagesA Preliminary Evaluation of The Potential of Beauveria Bassiana For Bed Bug ControlxidegaNo ratings yet
- Science PaperDocument2 pagesScience PaperMicael PortoNo ratings yet
- Biodiversity and Functional Genomics in The Human Microbiome PDFDocument8 pagesBiodiversity and Functional Genomics in The Human Microbiome PDFabcder1234No ratings yet
- Hunter Cevera1998Document8 pagesHunter Cevera1998Redd ZhuangNo ratings yet
- Antagonistic Activity Against Plant Pathogenic Fungus by Various Indigenous Microorganisms From Different Cropping Systems in Soc Trang Province, VietnamDocument8 pagesAntagonistic Activity Against Plant Pathogenic Fungus by Various Indigenous Microorganisms From Different Cropping Systems in Soc Trang Province, VietnamrahmanNo ratings yet
- He Et Al 2019Document26 pagesHe Et Al 2019rini susilowatiNo ratings yet
- Chapter 1Document10 pagesChapter 1EDLE FAITH ANDREA CATABIANNo ratings yet
- 3 - Biotek Tanah Plant Growth-Promoting MicroorganismDocument15 pages3 - Biotek Tanah Plant Growth-Promoting MicroorganismErwin irawanNo ratings yet
- 3 Cutaneous MicrobiomeDocument10 pages3 Cutaneous MicrobiomeAna Li SingNo ratings yet
- Jurnal Internasional The Effect of ManureDocument11 pagesJurnal Internasional The Effect of ManureIkram AffandiNo ratings yet
- Gut MicrobiomeDocument23 pagesGut Microbiomejyotisingh7No ratings yet
- Maintenance of Soil Functioning Following Erosion of Microbial DiversityDocument8 pagesMaintenance of Soil Functioning Following Erosion of Microbial DiversityAde Rizky FajrullohNo ratings yet
- Jimb0514 PDFDocument9 pagesJimb0514 PDFTitik SuratiNo ratings yet
- Pnas 1220608110Document6 pagesPnas 1220608110Glorifya JulitaNo ratings yet
- Microbial Physiology Notes 1Document9 pagesMicrobial Physiology Notes 1Genalin M. Escobia-Bagas100% (1)
- 2 Introduction To MicrobiologyDocument40 pages2 Introduction To MicrobiologyViah Angela CaasiNo ratings yet
- Do Not Copy Penalties Apply: The Role of Cutaneous Microbiota Harmony in Maintaining A Functional Skin BarrierDocument7 pagesDo Not Copy Penalties Apply: The Role of Cutaneous Microbiota Harmony in Maintaining A Functional Skin BarriereaudreyliaNo ratings yet
- Department of Soil Science and Agricultural ChemistryDocument38 pagesDepartment of Soil Science and Agricultural ChemistryGUMMALLA ANIL KUMARNo ratings yet
- Trivedi Effect - Antimicrobial Susceptibility of Proteus Mirabilis: Impact of Biofield Energy TreatmentDocument5 pagesTrivedi Effect - Antimicrobial Susceptibility of Proteus Mirabilis: Impact of Biofield Energy TreatmentTrivedi EffectNo ratings yet
- Microbiome AnalysisDocument13 pagesMicrobiome AnalysisJulius Memeg PanayoNo ratings yet
- Normal Flora, Bacteria, and DiseaseDocument8 pagesNormal Flora, Bacteria, and DiseaseDeepbluexNo ratings yet
- Sun 2009Document6 pagesSun 2009Django BoyeeNo ratings yet
- Jurnal 5Document7 pagesJurnal 5kalstarNo ratings yet
- 1-Introduction To MicroorganismsDocument23 pages1-Introduction To Microorganismsguestiam29No ratings yet
- Immune Homeostasis, Dysbiosis and Therapeutic Modulation of The Gut MicrobiotaDocument15 pagesImmune Homeostasis, Dysbiosis and Therapeutic Modulation of The Gut MicrobiotaAjit Kumar BaidyaNo ratings yet
- Evolution of Antibiotic Resistance at Non-Lethal Drug ConcentrationsDocument11 pagesEvolution of Antibiotic Resistance at Non-Lethal Drug ConcentrationsT. Jonathan SianturiNo ratings yet
- Introduction To Environmental MicrobiologyDocument44 pagesIntroduction To Environmental MicrobiologyGermaineNo ratings yet
- Cross BiomeDocument6 pagesCross BiomeALEXANDRA ANTONIETA URRUTIA ZEGARRANo ratings yet
- Linking Species Richness, Biodiversity and Ecosystem Function in Soil SystemsDocument19 pagesLinking Species Richness, Biodiversity and Ecosystem Function in Soil SystemsroseNo ratings yet
- Biodiversity MicrobialDocument21 pagesBiodiversity MicrobialShuddhodana MukthaNo ratings yet
- Reflection On Revisions Assignment SheetDocument2 pagesReflection On Revisions Assignment Sheetapi-331864368No ratings yet
- Annotated-Bio142 20preposal 20reflectionDocument2 pagesAnnotated-Bio142 20preposal 20reflectionapi-331864368No ratings yet
- 5 - Intro Bingo ActivityDocument1 page5 - Intro Bingo Activityapi-331864368No ratings yet
- Emails Lab Activity and Lesson PlanDocument11 pagesEmails Lab Activity and Lesson Planapi-331864368No ratings yet
- Prepare For Linkage - Review From 141Document2 pagesPrepare For Linkage - Review From 141api-331864368No ratings yet
- Student Feedback That Influenced My Teaching PhilosophyDocument1 pageStudent Feedback That Influenced My Teaching Philosophyapi-331864368No ratings yet
- Example Online DiscussionDocument3 pagesExample Online Discussionapi-331864368No ratings yet
- 5 - Information From Supplemental Course Evaluation Delivered Via Google Forms For All Bio142 Students in Fall 2020Document2 pages5 - Information From Supplemental Course Evaluation Delivered Via Google Forms For All Bio142 Students in Fall 2020api-331864368No ratings yet
- Ors Poster Spring 2021 To ShareDocument1 pageOrs Poster Spring 2021 To Shareapi-331864368No ratings yet
- 1 - Exp ApplicationDocument5 pages1 - Exp Applicationapi-331864368No ratings yet
- 6 - Example Student Final PaperDocument22 pages6 - Example Student Final Paperapi-331864368No ratings yet
- 3 - Two Examples of Active Learning in The Bio142 Online ClassroomDocument2 pages3 - Two Examples of Active Learning in The Bio142 Online Classroomapi-331864368No ratings yet
- 7 - Example Asynchronous Lesson PlanDocument3 pages7 - Example Asynchronous Lesson Planapi-331864368No ratings yet
- Ors Poster Summer 2021Document1 pageOrs Poster Summer 2021api-331864368No ratings yet
- Emily Mclean Curriculum VitaeDocument5 pagesEmily Mclean Curriculum Vitaeapi-331864368No ratings yet
- Catalog en Eurocoustic 0Document132 pagesCatalog en Eurocoustic 0Vikash KumarNo ratings yet
- Diana ACCOMPLISHMENTDocument8 pagesDiana ACCOMPLISHMENTMarycon MaapoyNo ratings yet
- VEIKK A15PRO Instruction Manual 0714Document20 pagesVEIKK A15PRO Instruction Manual 0714Corny777 UwUNo ratings yet
- BIS Ventilation Brochure enDocument16 pagesBIS Ventilation Brochure enBruno SantosNo ratings yet
- 200mm 250mm 300 MM: Project Name Location Bill No. - 00 (Rev. 0) Consultant Contractor Buildiong Name DATEDocument2 pages200mm 250mm 300 MM: Project Name Location Bill No. - 00 (Rev. 0) Consultant Contractor Buildiong Name DATEganesh gundNo ratings yet
- SurveyDocument1 pageSurveyJainne Ann BetchaidaNo ratings yet
- Manual Servicio Soxte 2050Document46 pagesManual Servicio Soxte 2050Quimica JordanlabNo ratings yet
- Vocabulary June v22Document2 pagesVocabulary June v22Wiston TonwisNo ratings yet
- 432.01 Managing HSE in A Geophysical Nov 2017Document138 pages432.01 Managing HSE in A Geophysical Nov 2017Andrei Savu100% (1)
- Fulltext PDFDocument55 pagesFulltext PDFManikandan VpNo ratings yet
- ND 0108 SpongeDiver SymiDocument2 pagesND 0108 SpongeDiver SymiPoki MokiNo ratings yet
- Argumentative EssayDocument5 pagesArgumentative EssayJoshua MontoyaNo ratings yet
- Cell Division-Mitosis Notes: 2 New CellsDocument21 pagesCell Division-Mitosis Notes: 2 New CellsCristina MariaNo ratings yet
- Lead and Manage People TTLMDocument41 pagesLead and Manage People TTLMHenok Mehari100% (1)
- Water Distiller - ManualDocument2 pagesWater Distiller - ManualSanjeevi JagadishNo ratings yet
- UNIT-5 International Dimensions To Industrial Relations: ObjectivesDocument27 pagesUNIT-5 International Dimensions To Industrial Relations: ObjectivesManish DwivediNo ratings yet
- 06 - Flexible Operation of Thermal Power Plants - OEM Perspective and Experiences PDFDocument22 pages06 - Flexible Operation of Thermal Power Plants - OEM Perspective and Experiences PDFRavishankarNo ratings yet
- Modelling The Effects of Condensate Banking On High CGR ReservoirsDocument11 pagesModelling The Effects of Condensate Banking On High CGR ReservoirslikpataNo ratings yet
- Summary of Germ LayersDocument2 pagesSummary of Germ Layersaichiii.bearNo ratings yet
- Health Insurance BookDocument3 pagesHealth Insurance BookHarish SihareNo ratings yet
- Midas Tutorial Fea 7Document3 pagesMidas Tutorial Fea 7sasiNo ratings yet
- Ducted Exhaust Ventilation Fans: Low Noise, High Performance Air and Moisture ExtractionDocument4 pagesDucted Exhaust Ventilation Fans: Low Noise, High Performance Air and Moisture ExtractionNicolas BaquedanoNo ratings yet
- 5 Natural Sore Throat RemediesDocument5 pages5 Natural Sore Throat Remedieslisa smithis100% (1)
- 6 Human Diseases That Cause by VirusesDocument7 pages6 Human Diseases That Cause by VirusesJefry JapNo ratings yet
- 1101259L 580.752830 Pressure Washer ManualDocument64 pages1101259L 580.752830 Pressure Washer Manualgork1roguesNo ratings yet
- AE Board Review Part3 - MSWord2003 Long Paper 2011Document12 pagesAE Board Review Part3 - MSWord2003 Long Paper 2011Jan14No ratings yet
- Book Summary - I Hear You Summary Michael Sorensen - Read in 7 MinutesDocument10 pagesBook Summary - I Hear You Summary Michael Sorensen - Read in 7 MinutesNogüey JoseNo ratings yet
- Surveys For Maint'Ce ClassDocument7 pagesSurveys For Maint'Ce ClassSuhe EndraNo ratings yet
- Jokes and Their Relation To The Unconscious: Laurence HenkelmanDocument3 pagesJokes and Their Relation To The Unconscious: Laurence HenkelmanMilos VisnjicNo ratings yet
- Final Workshop Report On Value Addition and AgroprocessingDocument31 pagesFinal Workshop Report On Value Addition and AgroprocessingBett K. BernardNo ratings yet