Professional Documents
Culture Documents
Lecture 9.1-Gene Expression Protein Synthesis
Uploaded by
Mariah Jane Taladua0 ratings0% found this document useful (0 votes)
15 views55 pagesCopyright
© © All Rights Reserved
Available Formats
PDF, TXT or read online from Scribd
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
© All Rights Reserved
Available Formats
Download as PDF, TXT or read online from Scribd
0 ratings0% found this document useful (0 votes)
15 views55 pagesLecture 9.1-Gene Expression Protein Synthesis
Uploaded by
Mariah Jane TaladuaCopyright:
© All Rights Reserved
Available Formats
Download as PDF, TXT or read online from Scribd
You are on page 1of 55
26
General, Organic, and
Biochemistry, 8e
Bettelheim, Brown,
© 2006 Thomson Learning, Inc.
Campbell, and Farrell
All rights reserved
26-1
26 Chapter 26
Gene Expression and
Protein Synthesis
© 2006 Thomson Learning, Inc.
All rights reserved
26-2
26 Introduction
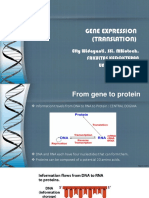
• The central dogma of molecular biology
• Information contained in DNA molecules is expressed
in the structure of proteins.
• Gene expression is the turning on or activation of a
gene.
© 2006 Thomson Learning, Inc.
All rights reserved
26-3
26 Transcription
• Transcription: the process by which information
encoded in a DNA molecule is copied into an
mRNA molecule.
• Transcription starts when the DNA double helix begins
to unwind near the gene to be transcribed.
• Only one strand of the DNA is transcribed.
• Ribonucleotides assemble along the unwound DNA
strand in a complementary sequence.
• Enzymes called polymerases (poly) catalyze
transcription: poly I for rRNA formation, poly II for
mRNA formation, and poly III for tRNA formation.
© 2006 Thomson Learning, Inc.
All rights reserved
26-4
26 Transcription
Transcription of Gene
© 2006 Thomson Learning, Inc.
All rights reserved
26-5
26 Transcription
• A eukaryotic gene has two parts:
• A structural gene that is transcribed into RNA; the
structural gene is made of exons and introns.
• A regulatory gene that controls transcription; the
regulatory gene is not transcribed but has control
elements, one of which is the promoter.
• A promoter is unique to each gene.
• There is always a sequence of bases on the DNA strand
called an initiation signal.
• Promoters also contain consensus sequences, such as
the TATA box, in which the two nucleotides T and A are
repeated many times.
© 2006 Thomson Learning, Inc.
All rights reserved
26-6
26 Transcription
• A TATA box lies approximately 25 base pairs
upstream (Figure 25.2).
• All three RNA polymerases interact with their
promoter regions via transcription factors that
are binding proteins.
• After initiation, RNA polymerase zips up the
complementary bases in a process called
elongation.
• Elongation is in the 5’ -> 3’ direction.
• At the end of each gene is a termination
sequence.
© 2006 Thomson Learning, Inc.
All rights reserved
26-7
26 Transcription
• Poly II has two different forms:
• At its C-terminal domain, it has Ser and Thr
repeats that can be phosphorylated.
• When poly II starts initiation, it is
unphosphorylated.
• Upon phosphorylation, it catalyzes elongation.
• After termination of the transcription, it is
dephosphorylated by a phosphatase.
• In this manner, poly II is constantly recycled
between its initiation and elongation roles.
© 2006 Thomson Learning, Inc.
All rights reserved
26-8
26 Transcription
• The RNA products of transcription are not
necessarily functional RNAs.
• They are made functional by post-transcription
modification.
• Transcribed mRNA is capped at both ends.
• The 5’ end acquires a methylated guanine.
• The 3’ end acquires a polyA tail that may contain from
100 to 200 adenine residues.
• Once the two ends are capped, the introns are spliced
out.
• tRNA is similarly trimmed, capped, and methylated.
• Functional rRNA also undergoes post-transcription
methylation.
© 2006 Thomson Learning, Inc.
All rights reserved
26-9
26 RNA in Translation
• mRNA, rRNA, and tRNA all participate in translation.
• Protein synthesis takes place on ribosomes.
• A ribosome dissociates into larger and a smaller body.
• In higher organisms, the larger body is called a 60S
ribosome; the smaller body is called a 40S ribosome.
• The 5’ end of the mature mRNA is bonded to the 40S
ribosome and this unit then joined to the 60S ribosome.
• Together the 40S and 60S ribosomes form a unit on
which mRNA is stretched out.
• Triplets of bases on mRNA are called codons.
• The 20 amino acids are then brought to the mRNA-
ribosome complex, each amino acid by its own
particular
© 2006 Thomson Learning, Inc. tRNA.
All rights reserved
26-10
26 Key Players
• mRNA carries the
information from a
gene in DNA.
• Ribosomes, made of
rRNA, consist of
subunits and carry
out an enzyme-like
role.
• tRNA carries specific
amino acids to the
ribosome.
© 2006 Thomson Learning, Inc.
All rights reserved
26-11
26 RNA in Translation
RNA Translation
© 2006 Thomson Learning, Inc.
All rights reserved
26-12
26 tRNA
• Each tRNA is specific for only one amino acid.
• Each cell carries at least 20 specific enzymes, each
specific for one amino acid.
• Each enzyme recognizes only one tRNA.
• The enzyme bonds the activated amino acid to the 3’
terminal -OH group of the appropriate tRNA by an ester
bond.
• At the opposite end of the tRNA molecule is a codon
recognition site.
• The codon recognition site is a sequence of three
bases called an anticodon.
• This triplet of bases aligns itself in a complementary
fashion to the codon triplet on mRNA.
© 2006 Thomson Learning, Inc.
All rights reserved
26-13
26 tRNA
• Figure 26.4 The three-dimensional structure of a
tRNA.
© 2006 Thomson Learning, Inc.
All rights reserved
26-14
26 The Genetic Code
• Assignments of triplets is based on several types
of experiments.
• One of these used synthetic mRNA.
• If mRNA is polyU, polyPhe is formed; the triplet UUU,
therefore, must code for Phe.
• If mRNA is poly ---ACACAC---, poly(Thr-His) is formed;
ACA must code for Thr, and CAC for His.
• By 1967, the genetic code was broken.
© 2006 Thomson Learning, Inc.
All rights reserved
26-15
26 The genetic code matches each RNA codon with its amino acid or function.
© 2006 Thomson Learning, Inc.
All rights reserved
26-16
26 Features of the Code
• All 64 codons have been assigned.
• 61 code for amino acids.
• 3 (UAA, UAG, and UGA) serve as termination signals.
• AUG also serves as an initiation signal.
• Only Trp and Met have one codon each.
• More than one triplet can code for the same amino
acid; Leu, Ser, and Arg, for example, are each coded for
by six triplets.
• The third base is irrelevant for Leu, Val, Ser, Pro, Thr,
Ala, Gly, and Arg.
© 2006 Thomson Learning, Inc.
All rights reserved
26-17
26 Features of the Code
• For the 15 amino acids coded for by 2, 3, or 4 triplets, it
is only the third letter of the codon that varies. Gly, for
example, is coded for by GGA, GGG, GGC, and GGU.
• The code is almost universal: it the same in viruses,
prokaryotes, and eukaryotes; the only exceptions are
some codons in mitochondria.
© 2006 Thomson Learning, Inc.
All rights reserved
26-18
26 Translation
• Translation: the process whereby a base
sequence of mRNA is used to create a protein.
• There are four major stages in protein synthesis:
• Activation
• Initiation
• Elongation
• Termination
© 2006 Thomson Learning, Inc.
All rights reserved
26-19
26
© 2006 Thomson Learning, Inc.
All rights reserved
26-20
26 Transcription to
Translation
© 2006 Thomson Learning, Inc.
All rights reserved
26-21
26 Protein Synthesis
• Molecular components of reactions at four stages
of protein synthesis:
© 2006 Thomson Learning, Inc.
All rights reserved
26-22
26 Amino Acid Activation
• Requires
• amino acids
• tRNAs
• aminoacyl-
tRNA
synthetases
• ATP, Mg2+
• Formation on
an aminoacyl-
tRNA
© 2006 Thomson Learning, Inc.
All rights reserved
26-23
26 Amino Acid Activation
• The activated amino acid is bound to its own
particular tRNA by an ester bond between the
carboxyl group of the amino acid and the 3’-OH of
the tRNA.
© 2006 Thomson Learning, Inc.
All rights reserved
26-24
26 Amino Acid Activation
• This two-stage reaction allows selectivity at two
levels
• the amino acid: The aminoacyl-AMP remains bound to
the enzyme and binding of the correct amino acid is
verified by an editing site on the tRNA synthetase.
• tRNA: There are specific binding sites on tRNAs that
are recognized by aminoacyl-tRNA synthetases.
© 2006 Thomson Learning, Inc.
All rights reserved
26-25
26 Chain Initiation
• Initiation begins with
a tRNA bearing
methionine (met)
attaching to one of
the ribosomal units.
The codon for
methionine is a
universal “start”
codon for “reading”
the mRNA strand.
© 2006 Thomson Learning, Inc.
All rights reserved
26-26
26
• The ribosomal unit
binds to mRNA
where the code for
met is located
(AUG). The
anticodon (UAC) of
the tRNA matches
the “start” codon
on mRNA (AUG).
© 2006 Thomson Learning, Inc.
All rights reserved
26-27
26 Elongation
• The larger ribosomal
subunit now binds to
the smaller unit,
forming a ribosomal
complex. The tRNA
binds to the first
active site on the
ribosome.
Translation may now
begin.
© 2006 Thomson Learning, Inc.
All rights reserved
26-28
26 Elongation
• The second codon
in mRNA (GUU)
matches the
anticodon of a tRNA
carrying the amino
acid valine (CAA).
The second tRNA
binds to the second
active site on the
large subunit.
© 2006 Thomson Learning, Inc.
All rights reserved
26-29
26
• A catalytic site on
the larger subunit
binds the two
amino acids
together using
dehydration
synthesis, forming
a peptide bond
between them.
© 2006 Thomson Learning, Inc.
All rights reserved
26-30
26
• The first tRNA now
detaches and goes
of to find another
met in the
cytoplasm. The
mRNA chain shifts
over one codon,
placing the second
codon (CAU) over
the second active
site.
© 2006 Thomson Learning, Inc.
All rights reserved
26-31
26
• A tRNA with an
anticodon (GUA)
matching the
exposed codon
(CAU) moves onto
the ribosome. This
tRNA carries
histidine (his).
© 2006 Thomson Learning, Inc.
All rights reserved
26-32
26
• A new peptide bond
forms between val
and his on the
catalytic site. The
tRNA that carried val
will detach and find
another val in the
cytoplasm. The mRNA
strand will then shift
over one more codon.
© 2006 Thomson Learning, Inc.
All rights reserved
26-33
26 Peptide Bond Formation
• Figure 26.8 Peptide bond formation in protein
synthesis.
© 2006 Thomson Learning, Inc.
All rights reserved
26-34
26 Termination
• The process continues
until the ribosome finds a
“stop” codon. The
subunits detach from one
another, the mRNA is
released, and the
polypeptide chain moves
down the ER for further
processing. The initial
met is removed and the
chain is folded into its
final shape.
© 2006 Thomson Learning, Inc.
All rights reserved
26-35
26• Chain termination requires:
• Termination codons (UAA, UAG, or UGA) of mRNA.
• Releasing factors that cleave the polypeptide chain
from the last tRNA and release the tRNA from the
ribosome.
© 2006 Thomson Learning, Inc.
All rights reserved
26-36
26
© 2006 Thomson Learning, Inc.
All rights reserved
26-37
26 Gene Regulation
• Gene regulation: the various methods used by
organisms to control which genes will be
expressed and when.
• Some regulations operate at the transcriptional level
(DNA -> RNA)
• Others operate at the translational level (mRNA ->
protein).
© 2006 Thomson Learning, Inc.
All rights reserved
26-38
26 Transcriptional Level
• In eukaryotes, transcription is regulated by three
elements: promoters, enhancers, and response
elements.
• Promoters
• located adjacent to the transcription site.
• are defined by an initiator and conserved sequences
such as TATA or GC boxes.
• Different transcription factors bind to different modules
of the promoter.
• Transcription factors allow the rate of synthesis of
mRNA (and from there the target protein) to vary by a
factor of up to a million.
© 2006 Thomson Learning, Inc.
All rights reserved
26-39
26 Promoters
• Transcription factors find their targeted sites by
twisting their protein chains so that a certain amino
acid sequence is present at the surface.
• One such conformational twist is provided by metal-
binding fingers (next screen).
• Two other prominent transcription factor
conformations are the helix-turn-helix and the leucine
zipper.
• Transcription factors also possess repressors, which
reduce the rate of transcription.
© 2006 Thomson Learning, Inc.
All rights reserved
26-40
26 Promoters
• A schematic of a metal-binding finger.
© 2006 Thomson Learning, Inc.
All rights reserved
26-41
26 Enhancers
• Enhancers speed up transcription:
• They may be several thousand nucleotides away from
the transcription site; a loop in DNA brings the
enhancer to the initiation site.
© 2006 Thomson Learning, Inc.
All rights reserved
26-42
26 Response Elements
• Response elements are activated by their
transcription factors in response to an outside
stimulus.
• The stimulus may be heat shock, heavy metal toxicity,
or a hormonal signal.
• The response element of steroids is in front of and
about 250 base pairs upstream from the starting point
of transcription.
© 2006 Thomson Learning, Inc.
All rights reserved
26-43
26 Translational Level
• During translation, there are a number of
mechanisms that ensure quality control.
• AARS control
• Each amino acid must bond to the proper tRNA.
• Enzymes called aminoacyl-tRNA synthase (AARS)
catalyze this bonding.
• Each AARS recognizes its tRNA by specific nucleotide
sequences.
• Further, the active site of each AARS has two sieving
sites.
© 2006 Thomson Learning, Inc.
All rights reserved
26-44
26 Translational Level
• Termination control
• The stop codons must be recognized by release
factors.
• The release factor combines with GTP and binds to the
ribosomal A site when that site is occupied by the
termination codon.
• Post-translational control
• In most proteins, the Met at the N-terminal end is
removed by Met-aminopeptidase.
• Certain proteins called chaperones help newly
synthesized proteins to fold properly.
• If rescue by chaperones fails, proteasomes may
© 2006 Thomson degrade
All rights reserved
Learning, Inc. the misfolded protein.
26-45
26 Mutations and Mutagens
• Mutation: a heritable change in the base
sequence of DNA.
• It is estimated that, on average, there is one copying
error for every 1010 bases.
• Mutations can occur during replication.
• Base errors can also occur during transcription in
protein synthesis (a nonheritable error).
• Consider the mRNA codons for Val, which are CAT,
CAC, CAG, and CAA.
• If the original codon is CAT, it may be transcribed onto
mRNA as GUC which codes for Val.
• Other errors in replication may lead to a change in
© 2006 Thomson protein
All rights reserved
Learning, Inc. structure and be very harmful.
26-46
26 Mutations and Mutagens
• Mutagen: a chemical that causes a base change
in DNA.
• Many changes in base sequence caused by
radiation and mutagens do not become mutations
because cells have repair mechanisms called
nucleotide excision repair (NER).
• NER can prevent mutations by cutting out damaged
areas and resynthesizing them.
• Not all mutations are harmful.
• Certain ones may be beneficial because they enhance
the survival rate of the species.
© 2006 Thomson Learning, Inc.
All rights reserved
26-47
26 Recombinant DNA
• Recombinant DNA: DNA from two sources that
have been combined into one molecule.
• One example of the technique begins with
plasmids found in the cells of Escherichia coli.
• plasmid: a small, circular, double-stranded DNA
molecule of bacterial origin.
• A class of enzymes called restriction endonucleases
cleave DNA at specific locations.
• One, for example, may be specific for cleavage of the
bond between A-G in the sequence -CTTAAAG-.
© 2006 Thomson Learning, Inc.
All rights reserved
26-48
26 Recombinant DNA
• In this example “B ” stands for bacterial gene, and “H
for human gene.
• The DNA is now double-stranded with two “sticky
ends”, each with free bases that can pair with a
complementary section of DNA.
• Next, we cut a human gene with the same restriction
endonuclease; for example, the gene for human
insulin.
© 2006 Thomson Learning, Inc.
All rights reserved
26-49
26 Recombinant DNA
• The human gene is now spliced into the plasmid by the
enzyme DNA ligase.
• Splicing takes place at both ends of the human gene
and the plasmid is once again circular.
• The modified plasmid is then put back into the bacterial
cell where it replicates naturally every time the cell
divides.
• These cells now manufacture the human protein, in our
example human insulin, by transcription and
translation.
© 2006 Thomson Learning, Inc.
All rights reserved
26-50
26 Recombinant DNA
• Figure 26.16 The recombinant
DNA technique.
© 2006 Thomson Learning, Inc.
All rights reserved
26-51
26 Cloning DNA
• Figure 26.17 The
cloning of human
DNA fragments
with a viral vector.
© 2006 Thomson Learning, Inc.
All rights reserved
26-52
26 Gene Therapy
• Figure 26.18 Gene therapy
via retroviruses.
© 2006 Thomson Learning, Inc.
All rights reserved
26-53
26
© 2006 Thomson Learning, Inc.
All rights reserved
26-54
26 Protein Synthesis
Gene Expression and
Protein Synthesis
End
Chapter 26
© 2006 Thomson Learning, Inc.
All rights reserved
26-55
You might also like
- Gene ExpressionDocument72 pagesGene ExpressionMika ForwardNo ratings yet
- Translation: Translation: Assembly of Polypeptides On A RibosomeDocument36 pagesTranslation: Translation: Assembly of Polypeptides On A RibosomemahtabNo ratings yet
- Chapter 32 Lippincott BiochemistryDocument35 pagesChapter 32 Lippincott BiochemistryMeysam SajjadiNo ratings yet
- Protein Synthesis: Dr. Gertude Kiwanuka Biochemistry Dept MustDocument41 pagesProtein Synthesis: Dr. Gertude Kiwanuka Biochemistry Dept MustRonnieNo ratings yet
- Protein SynthesisDocument54 pagesProtein SynthesisGemmie Jn PierreNo ratings yet
- TranslationDocument37 pagesTranslationsanofazal786No ratings yet
- CHapter 14Document10 pagesCHapter 14alaskamofoNo ratings yet
- Translation in Prokaryotes and Genetic CodeDocument35 pagesTranslation in Prokaryotes and Genetic Codevanita garg100% (1)
- Lecture 2Document33 pagesLecture 2SUNDAS FATIMANo ratings yet
- Unit 2: The Molecular and Biochemical Basis of An OrganismDocument13 pagesUnit 2: The Molecular and Biochemical Basis of An OrganismMeghaNo ratings yet
- DNA Replication, Protein SynthesisDocument20 pagesDNA Replication, Protein SynthesisGarnetNo ratings yet
- L14 BioDocument7 pagesL14 BioAhmed MohamedNo ratings yet
- Lecture # 20 The Genetic CodeDocument41 pagesLecture # 20 The Genetic CodethomasNo ratings yet
- Biochemistry - Biosynthesis of Protein & Protein ChemistryDocument113 pagesBiochemistry - Biosynthesis of Protein & Protein ChemistryAdella AnggrainiNo ratings yet
- Transciption and TranslationDocument31 pagesTransciption and TranslationDuduetsang MosalakataneNo ratings yet
- 3.10 Protein-SynthesisDocument20 pages3.10 Protein-SynthesisPratika MNo ratings yet
- Central-Dogma 2Document22 pagesCentral-Dogma 2Milena De CresentNo ratings yet
- Unit 6. Proteins SynthesisDocument57 pagesUnit 6. Proteins Synthesismunyanezaolivier422No ratings yet
- Lecture 9 Translational, Protein Synthesis and Post Translational Modifications MD 2 2023 by DR Mohamed AbdelbakyDocument56 pagesLecture 9 Translational, Protein Synthesis and Post Translational Modifications MD 2 2023 by DR Mohamed Abdelbakysoushinelall2007No ratings yet
- Dogma Central Biology Pertemuan 5Document11 pagesDogma Central Biology Pertemuan 5Rizky BluesNo ratings yet
- Central DogmaDocument50 pagesCentral DogmaIvilyn ManguladNo ratings yet
- TranslationDocument17 pagesTranslationVishalNo ratings yet
- Protein Synthesis: DECEMBER 13, 2010 Cape Biology Unit 1 Mrs. HaughtonDocument45 pagesProtein Synthesis: DECEMBER 13, 2010 Cape Biology Unit 1 Mrs. HaughtonAshley MorganNo ratings yet
- Translation Initiation: April 2017 BIO 3421Document109 pagesTranslation Initiation: April 2017 BIO 3421Lucy ZuluNo ratings yet
- Foundations in Microbiology: TalaroDocument68 pagesFoundations in Microbiology: Talaromertx013No ratings yet
- Translation (Protein Synthesis)Document55 pagesTranslation (Protein Synthesis)seada JemalNo ratings yet
- Transcription and TranslationDocument30 pagesTranscription and TranslationNgô Minh KhuêNo ratings yet
- BiochemDocument66 pagesBiochemReendelle JamiatreNo ratings yet
- Genetic CodeDocument15 pagesGenetic Codekratikaagrawal50No ratings yet
- Chap Viii - Gene Expression and Protein SynthesisDocument63 pagesChap Viii - Gene Expression and Protein SynthesisNgaya AnthonyNo ratings yet
- 6.2 - TranslationDocument21 pages6.2 - TranslationAna JuatasNo ratings yet
- Genetic CodeDocument17 pagesGenetic Codegopikupuchitty50% (2)
- Presentation ON TranslationDocument21 pagesPresentation ON TranslationSuraiya Siraj NehaNo ratings yet
- RNA and Protein SynthesisDocument33 pagesRNA and Protein SynthesisRaphael Ongodia100% (2)
- Lecture 5 Diagnostic GenomicsDocument33 pagesLecture 5 Diagnostic Genomicssales zfNo ratings yet
- Central Dogma: Faculty of Medicine, Sriwijaya UniversityDocument51 pagesCentral Dogma: Faculty of Medicine, Sriwijaya UniversityIlsya PertiwiNo ratings yet
- Protein Synthesis NotesDocument17 pagesProtein Synthesis Notesapi-655732711100% (1)
- Chap12 Lecture 2019-SDocument61 pagesChap12 Lecture 2019-SPaulNo ratings yet
- Protein Synthesis: - The Formation of New Proteins Using The Code Carried On DNADocument20 pagesProtein Synthesis: - The Formation of New Proteins Using The Code Carried On DNABien Emilio B NavarroNo ratings yet
- November 26 Cell and Molecular Biology NOVEMBER 26 2021Document21 pagesNovember 26 Cell and Molecular Biology NOVEMBER 26 2021shelahomero3No ratings yet
- Protein SynthesisDocument60 pagesProtein Synthesislucky mbaselaNo ratings yet
- RNA and Protein Synthesis Notes - HONORS-Website PDFDocument30 pagesRNA and Protein Synthesis Notes - HONORS-Website PDFSphumelele AndiswaNo ratings yet
- Microbial GeneticsDocument72 pagesMicrobial GeneticsKhánh Trà QuốcNo ratings yet
- 6 Lecture-TranslationDocument25 pages6 Lecture-Translationmoazzanrana678No ratings yet
- 24 LN Protein Synthesis CNRAADocument48 pages24 LN Protein Synthesis CNRAADakshitha DharmakeerthiNo ratings yet
- Translation 160204065032Document79 pagesTranslation 160204065032Sai SundeepNo ratings yet
- 106 Ceng w10Document62 pages106 Ceng w10mertNo ratings yet
- 11b. Protein Synthesis - 2Document24 pages11b. Protein Synthesis - 2Arisha razaNo ratings yet
- IB Diploma Biology: Essential Idea: Transcription and Translation Allow The GeneticDocument37 pagesIB Diploma Biology: Essential Idea: Transcription and Translation Allow The GeneticClàudia VicenteNo ratings yet
- Meridian Protein SynthesisDocument7 pagesMeridian Protein SynthesisjosewaltNo ratings yet
- Protein Metabolism: Presented By: Danica Alyssa Cruz, RMTDocument39 pagesProtein Metabolism: Presented By: Danica Alyssa Cruz, RMTDanica Alyssa CruzNo ratings yet
- Sintesis DNADocument32 pagesSintesis DNAZhafran DarwisNo ratings yet
- 1.3.3. Ekspresi GenDocument28 pages1.3.3. Ekspresi GenAinuzzahraNo ratings yet
- Translation of mRNA (Protein Synthesis)Document80 pagesTranslation of mRNA (Protein Synthesis)Cok AnggaNo ratings yet
- Rna Structure, Transcription, Translation, Rna Synthesis, Processing, and ModificationDocument18 pagesRna Structure, Transcription, Translation, Rna Synthesis, Processing, and ModificationfidhavfathimaNo ratings yet
- Unit 11 PDFDocument15 pagesUnit 11 PDFcarlette11No ratings yet
- TranslasiDocument73 pagesTranslasitamara hannestoNo ratings yet
- Dna, Rna, Transcription, Translation & Protein Synthesis PDFDocument14 pagesDna, Rna, Transcription, Translation & Protein Synthesis PDFSaeed Khan100% (1)
- Topic 12 - Protein SynthesisDocument45 pagesTopic 12 - Protein SynthesisKaye LapizNo ratings yet
- Preclinical Biochemistry and Medical Genetics Review 2023: For USMLE Step 1 and COMLEX-USA Level 1From EverandPreclinical Biochemistry and Medical Genetics Review 2023: For USMLE Step 1 and COMLEX-USA Level 1No ratings yet
- Chapter 20Document5 pagesChapter 20Mariah Jane TaladuaNo ratings yet
- Rectal and Vaginal Suppository InsertionDocument2 pagesRectal and Vaginal Suppository InsertionMariah Jane TaladuaNo ratings yet
- Chapter 37Document10 pagesChapter 37Mariah Jane TaladuaNo ratings yet
- Chapter 36Document7 pagesChapter 36Mariah Jane TaladuaNo ratings yet
- Topic Outline: Nasal Cannula TentDocument2 pagesTopic Outline: Nasal Cannula TentMariah Jane TaladuaNo ratings yet
- CatheterizationDocument3 pagesCatheterizationMariah Jane TaladuaNo ratings yet
- Imci: Assess and Classify Fever: MeaslesDocument4 pagesImci: Assess and Classify Fever: MeaslesMariah Jane TaladuaNo ratings yet
- General, Organic, and Biochemistry, 8e: Bettelheim, Brown Campbell, and FarrellDocument43 pagesGeneral, Organic, and Biochemistry, 8e: Bettelheim, Brown Campbell, and FarrellMariah Jane TaladuaNo ratings yet
- Topic Outline: Week 2 / January 24, 2022Document2 pagesTopic Outline: Week 2 / January 24, 2022Mariah Jane TaladuaNo ratings yet
- Integrated Management of Childhood IllnessDocument2 pagesIntegrated Management of Childhood IllnessMariah Jane TaladuaNo ratings yet
- Family Nursing Care Plan: 2 Major Types of Nursing AssessmentDocument3 pagesFamily Nursing Care Plan: 2 Major Types of Nursing AssessmentMariah Jane TaladuaNo ratings yet
- Overview of Public Health NursingDocument6 pagesOverview of Public Health NursingMariah Jane TaladuaNo ratings yet
- Factors Affecting HealthDocument4 pagesFactors Affecting HealthMariah Jane TaladuaNo ratings yet
- Chapter 32 NotesDocument4 pagesChapter 32 NotesMariah Jane TaladuaNo ratings yet
- Preparation For ChildbirthDocument3 pagesPreparation For ChildbirthMariah Jane Taladua100% (1)
- Chapter 30 NotesDocument4 pagesChapter 30 NotesMariah Jane TaladuaNo ratings yet
- Community AssessmentDocument3 pagesCommunity AssessmentMariah Jane TaladuaNo ratings yet
- Chapter 31: Adrenergic Antagonists: Drug ListDocument7 pagesChapter 31: Adrenergic Antagonists: Drug ListMariah Jane TaladuaNo ratings yet
- Protein Synthesis WorksheetDocument2 pagesProtein Synthesis WorksheetJuan Esteban Escandon AgudeloNo ratings yet
- Molecular Basis of Inheritance: Free Sample ChapterDocument33 pagesMolecular Basis of Inheritance: Free Sample ChapterJiya ShahNo ratings yet
- IIsc Biological SciencesDocument17 pagesIIsc Biological SciencesdhurvasNo ratings yet
- Translation NotesDocument4 pagesTranslation NotesRishikesh BhintadeNo ratings yet
- Genome, Transcriptome and Proteome PDFDocument17 pagesGenome, Transcriptome and Proteome PDFAdn CodeNo ratings yet
- Genetic Polymorphisms Dan Its Role On Clinical ApplicationDocument104 pagesGenetic Polymorphisms Dan Its Role On Clinical ApplicationyuslipahNo ratings yet
- Physiological Functions of Heat Shock ProteinsDocument10 pagesPhysiological Functions of Heat Shock ProteinsDerfel CardanNo ratings yet
- Ribosome: Our Updated Will Become Effective On May 25, 2012Document15 pagesRibosome: Our Updated Will Become Effective On May 25, 2012Ashok Kumar0% (1)
- Sem.1 Revision HLDocument88 pagesSem.1 Revision HLRawanMazen SharifNo ratings yet
- Exercise 5 Molecular Basis of HeredityDocument6 pagesExercise 5 Molecular Basis of HeredityHiroshi MatsushimaNo ratings yet
- DLL 10.jazDocument19 pagesDLL 10.jazkaycin DuzonNo ratings yet
- Rediscovering Biology TextbookDocument203 pagesRediscovering Biology TextbookJuiced-IN itNo ratings yet
- Fun With DNA Worksheet IDocument6 pagesFun With DNA Worksheet IHafsa KhanNo ratings yet
- BIO 216-Molecular Biology-Sec1-Muhammad Tariq-Shaper Mirza-Syed Shahzad Ul HussanDocument3 pagesBIO 216-Molecular Biology-Sec1-Muhammad Tariq-Shaper Mirza-Syed Shahzad Ul HussanAnonymous sF8ZuiGNo ratings yet
- SBG150127 GenEZ HandbookDocument18 pagesSBG150127 GenEZ HandbookRNo ratings yet
- Transcription: From DNA To RNADocument74 pagesTranscription: From DNA To RNAmd habibur rahmanNo ratings yet
- Dplex ManualDocument44 pagesDplex ManualTrang DoNo ratings yet
- Performance Task #1 - Aceron, Barrina, VelasDocument6 pagesPerformance Task #1 - Aceron, Barrina, VelasCalil AceronNo ratings yet
- TranscriptionDocument30 pagesTranscriptionAizelle TarataraNo ratings yet
- Screenshot 2565-12-27 at 17.12.37Document61 pagesScreenshot 2565-12-27 at 17.12.37Kittipat TeeraphatpornchaiNo ratings yet
- The Cell Biology of Rabies Virus: Using Stealth To Reach The BrainDocument11 pagesThe Cell Biology of Rabies Virus: Using Stealth To Reach The BrainFlavius AnghelNo ratings yet
- Mod 9 CC of Proteins Part 1Document9 pagesMod 9 CC of Proteins Part 1Benson PaglinawanNo ratings yet
- DNA Structure and Gene FunctionDocument12 pagesDNA Structure and Gene FunctionJiro SarmientoNo ratings yet
- MRCP Notes 2006Document117 pagesMRCP Notes 2006Walaa Ismail HidirbiNo ratings yet
- Bio FinalDocument8 pagesBio FinalChris SmithNo ratings yet
- Protein Synthesis Extra CreditDocument2 pagesProtein Synthesis Extra Creditcmillica1176No ratings yet
- MYCO VIRO 1-MergedDocument29 pagesMYCO VIRO 1-MergedGennelyn Ross Delos ReyesNo ratings yet
- Biological MoleculesDocument98 pagesBiological MoleculesSuyashNo ratings yet
- Grade 10 Ac Vity Sheet On Protein Synthesis Ac Vity 7: Decoding DNA Segment Objec VeDocument5 pagesGrade 10 Ac Vity Sheet On Protein Synthesis Ac Vity 7: Decoding DNA Segment Objec VeChristian Jay GuillermoNo ratings yet
- Goodsell-2009-Biochemistry and Molecular Biology Education PDFDocument8 pagesGoodsell-2009-Biochemistry and Molecular Biology Education PDFJeremy HutchinsonNo ratings yet