Professional Documents
Culture Documents
Pnas 20161dfhd9944si
Uploaded by
PaolaMorochoOriginal Description:
Original Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
Pnas 20161dfhd9944si
Uploaded by
PaolaMorochoCopyright:
Available Formats
Supporting Information
Edge et al. 10.1073/pnas.1619944114
Fig. S1. Allelic imputation accuracies for 431 non-CODIS tetranucleotide STR loci. The plot considers the partition of the data represented in Fig. 1. Beagle
imputation accuracy is obtained by imputing the STR genotype assigned the highest imputation probability by Beagle. Null imputation accuracy is obtained by
imputing the same STR genotype for all people, irrespective of nearby SNP genotypes. Markers are sorted from left to right by null accuracy. Across all loci, the
mean null accuracy is 0.497, and the mean Beagle accuracy is 0.624. Note that ref. 11 compared 432 rather than 431 non-CODIS tetranucleotides with the CODIS
loci; we omitted TPO-D2S, an alias for the CODIS locus TPOX.
Edge et al. www.pnas.org/cgi/content/short/1619944114 1 of 9
Fig. S2. The median proportion of test-set CODIS and SNP records matched correctly as a function of the sizes of the training and test sets. We divided the
data into training and test sets in 1,000 ways, examining training sets of sizes 436, 545, 654, and 763—representing 50, 62.5, 75, and 87.5% of the data. For
each training-set size, we used test-set sizes that were multiples of 109 (1/8 of 872), so that the sum of training-set and test-set sizes did not exceed 872. For
each of 10 possible schemes for the proportions representing the training and test sets, we considered 100 random divisions of the data, using the same
100 partitions in all analyses for a given scheme. (A) One-to-one matching. (B) One-to-many matching selecting the STR profile that best matches a query SNP
profile. (C) One-to-many matching selecting the SNP profile that best matches a query STR profile. (D) Needle-in-haystack matching. In D, the vertical axis has
the same scale as in the other panels.
Edge et al. www.pnas.org/cgi/content/short/1619944114 2 of 9
0
0
A 1 B 1 C 1 D 1
1/3
1/3
1/3
1/3
Inc
Inc
Inc
Inc
ct
ct
ct
ct
2/3 2/3 2/3 2/3
rre
rre
rre
rre
or r
or r
or r
or r
Co
Co
Co
Co
ec
ec
ec
ec
t
t
2/3
2/3
2/3
2/3
1/3 1/3 1/3 1/3
1
0 0 0 0
2/3
1/3
2/3
1/3
2/3
1/3
2/3
1/3
0
0
1
1
Unassigned Unassigned Unassigned Unassigned
0
0
E 1 F 1 G 1
1/3
1/3
1/3
Inc
Inc
Inc
ct
ct
ct
2/3 2/3 2/3
rre
rre
rre
orr
orr
orr
Co
Co
Co
ec
ec
ec
t
t
2/3
2/3
2/3
1/3 1/3 1/3
1
1
0 0 0
2/3
1/3
2/3
1/3
2/3
1/3
0
0
1
1
Unassigned Unassigned Unassigned
0
H 1 I 1
1/3
1/3
Inc
Inc
ct
ct
2/3 2/3
rre
rre
orr
orr
Highest accuracy
Co
Co
ec
ec
Median accuracy
t
t
2/3
2/3
1/3 1/3 Lowest accuracy
1
0 0
2/3
1/3
2/3
1/3
0
0
1
Unassigned Unassigned
0
J 1
1/3
Inc
ct
2/3
rre
orr
Co
ec
t
2/3
1/3
1
0
2/3
1/3
0
1
Unassigned
Fig. S3. Proportions of the sample unassigned, correctly assigned, and incorrectly assigned as a function of the match-score threshold under one-to-one
matching using the Hungarian method. Each panel considers different proportions (training, test) of the total data (n = 872) allocated into training and test
sets, with 100 allocations according to those proportions. (A) 1/2, 1/8. (B) 1/2, 1/4. (C ) 1/2, 3/8. (D) 1/2, 1/2. (E ) 5/8, 1/8. (F ) 5/8, 1/4. (G) 5/8, 3/8. (H) 3/4, 1/8.
(I) 3/4, 1/4. (J) 7/8, 1/8. The figure design follows Fig. 3.
Edge et al. www.pnas.org/cgi/content/short/1619944114 3 of 9
Fig. S4. The median value of the mean allelic imputation accuracy across 13 CODIS markers as a function of the size of the training set. Beagle and null
imputation accuracies follow Fig. 1. The median is taken across 100 partitions into training and test sets. Imputation accuracies are plotted for all 10 schemes
for the sizes of training and test sets; multiple test-set sizes produce similar values at a fixed training-set size, and they are represented by overlapping plotted
points. The lines connect the median values for the test-set sizes at given training-set sizes.
Edge et al. www.pnas.org/cgi/content/short/1619944114 4 of 9
0
0
A 1 B 1 C 1 D 1
1/3
1/3
1/3
1/3
Inc
Inc
Inc
Inc
ct
ct
ct
ct
2/3 2/3 2/3 2/3
rre
rre
rre
rre
or r
or r
or r
or r
Co
Co
Co
Co
ec
ec
ec
ec
t
t
2/3
2/3
2/3
2/3
1/3 1/3 1/3 1/3
1
0 0 0 0
2/3
1/3
2/3
1/3
2/3
1/3
2/3
1/3
0
0
1
1
Unassigned Unassigned Unassigned Unassigned
0
0
E 1 F 1 G 1
1/3
1/3
1/3
Inc
Inc
Inc
ct
ct
ct
2/3 2/3 2/3
rre
rre
rre
orr
orr
orr
Co
Co
Co
ec
ec
ec
t
t
2/3
2/3
2/3
1/3 1/3 1/3
1
1
0 0 0
2/3
1/3
2/3
1/3
2/3
1/3
0
0
1
1
Unassigned Unassigned Unassigned
0
H 1 I 1
1/3
1/3
Inc
Inc
ct
ct
2/3 2/3
rre
rre
orr
orr
Highest accuracy
Co
Co
ec
ec
Median accuracy
t
t
2/3
2/3
1/3 1/3 Lowest accuracy
1
0 0
2/3
1/3
2/3
1/3
0
0
1
Unassigned Unassigned
0
J 1
1/3
Inc
ct
2/3
rre
orr
Co
ec
t
2/3
1/3
1
0
2/3
1/3
0
1
Unassigned
Fig. S5. Proportions of the sample unassigned, correctly assigned, and incorrectly assigned as a function of the match-score threshold under one-to-many
matching that attempts to find the CODIS profile that matches a query SNP profile. Each panel considers different proportions (training, test) of the total data
(n = 872) allocated into training and test sets, with 100 allocations according to those proportions. (A) 1/2, 1/8. (B) 1/2, 1/4. (C) 1/2, 3/8. (D) 1/2, 1/2. (E) 5/8, 1/8.
(F) 5/8, 1/4. (G) 5/8, 3/8. (H) 3/4, 1/8. (I) 3/4, 1/4. (J) 7/8, 1/8. The figure design follows Fig. 3.
Edge et al. www.pnas.org/cgi/content/short/1619944114 5 of 9
0
0
A 1 B 1 C 1 D 1
1/3
1/3
1/3
1/3
Inc
Inc
Inc
Inc
ct
ct
ct
ct
2/3 2/3 2/3 2/3
rre
rre
rre
rre
or r
or r
or r
or r
Co
Co
Co
Co
ec
ec
ec
ec
t
t
2/3
2/3
2/3
2/3
1/3 1/3 1/3 1/3
1
0 0 0 0
2/3
1/3
2/3
1/3
2/3
1/3
2/3
1/3
0
0
1
1
Unassigned Unassigned Unassigned Unassigned
0
0
E 1 F 1 G 1
1/3
1/3
1/3
Inc
Inc
Inc
ct
ct
ct
2/3 2/3 2/3
rre
rre
rre
orr
orr
orr
Co
Co
Co
ec
ec
ec
t
t
2/3
2/3
2/3
1/3 1/3 1/3
1
1
0 0 0
2/3
1/3
2/3
1/3
2/3
1/3
0
0
1
1
Unassigned Unassigned Unassigned
0
H 1 I 1
1/3
1/3
Inc
Inc
ct
ct
2/3 2/3
rre
rre
orr
orr
Highest accuracy
Co
Co
ec
ec
Median accuracy
t
t
2/3
2/3
1/3 1/3 Lowest accuracy
1
0 0
2/3
1/3
2/3
1/3
0
0
1
Unassigned Unassigned
0
J 1
1/3
Inc
ct
2/3
rre
orr
Co
ec
t
2/3
1/3
1
0
2/3
1/3
0
1
Unassigned
Fig. S6. Proportions of the sample unassigned, correctly assigned, and incorrectly assigned as a function of the match-score threshold under one-to-many
matching that attempts to find the SNP profile that matches a query CODIS profile. Each panel considers different proportions (training, test) of the total data
(n = 872) allocated into training and test sets, with 100 allocations according to those proportions. (A) 1/2, 1/8. (B) 1/2, 1/4. (C) 1/2, 3/8. (D) 1/2, 1/2. (E) 5/8, 1/8.
(F) 5/8, 1/4. (G) 5/8, 3/8. (H) 3/4, 1/8. (I) 3/4, 1/4. (J) 7/8, 1/8. The figure design follows Fig. 3.
Edge et al. www.pnas.org/cgi/content/short/1619944114 6 of 9
0
0
A 1 B 1 C 1 D 1
1/3
1/3
1/3
1/3
Inc
Inc
Inc
Inc
ct
ct
ct
ct
2/3 2/3 2/3 2/3
rre
rre
rre
rre
or r
or r
or r
or r
Co
Co
Co
Co
ec
ec
ec
ec
t
t
2/3
2/3
2/3
2/3
1/3 1/3 1/3 1/3
1
0 0 0 0
2/3
1/3
2/3
1/3
2/3
1/3
2/3
1/3
0
0
1
1
Unassigned Unassigned Unassigned Unassigned
0
0
E 1 F 1 G 1
1/3
1/3
1/3
Inc
Inc
Inc
ct
ct
ct
2/3 2/3 2/3
rre
rre
rre
orr
orr
orr
Co
Co
Co
ec
ec
ec
t
t
2/3
2/3
2/3
1/3 1/3 1/3
1
1
0 0 0
2/3
1/3
2/3
1/3
2/3
1/3
0
0
1
1
Unassigned Unassigned Unassigned
0
H 1 I 1
1/3
1/3
Inc
Inc
ct
ct
2/3 2/3
rre
rre
orr
orr
Highest accuracy
Co
Co
ec
ec
Median accuracy
t
t
2/3
2/3
1/3 1/3 Lowest accuracy
1
0 0
2/3
1/3
2/3
1/3
0
0
1
Unassigned Unassigned
0
J 1
1/3
Inc
ct
2/3
rre
orr
Co
ec
t
2/3
1/3
1
0
2/3
1/3
0
1
Unassigned
Fig. S7. Proportions of the sample unassigned, correctly assigned, and incorrectly assigned as a function of the match-score threshold under needle-in-
haystack matching. Each panel considers different proportions (training, test) of the total data (n = 872) allocated into training and test sets, with 100 allo-
cations according to those proportions. (A) 1/2, 1/8. (B) 1/2, 1/4. (C) 1/2, 3/8. (D) 1/2, 1/2. (E) 5/8, 1/8. (F) 5/8, 1/4. (G) 5/8, 3/8. (H) 3/4, 1/8. (I) 3/4, 1/4. (J) 7/8, 1/8. The
figure design follows Fig. 3.
Edge et al. www.pnas.org/cgi/content/short/1619944114 7 of 9
Table S1. Sample sizes by population
Continental region and population Sample size
Sub-Saharan Africa 76
Bantu (Kenya) 11
Bantu (southern Africa) 7
Biaka Pygmy 8
Mandenka 19
Mbuti Pygmy 11
Yoruba 20
Europe 151
Adygei 17
Basque 23
French 24
Italian 12
Orcadian 15
Russian 25
Sardinian 28
Tuscan 7
Middle East 155
Bedouin 45
Druze 41
Mozabite 23
Palestinian 46
Central/South Asia 198
Balochi 24
Brahui 25
Burusho 25
Hazara 22
Kalash 23
Makrani 25
Pathan 21
Sindhi 23
Uygur 10
East Asia 227
Cambodian 10
Dai 10
Daur 9
Han 34
Han (North China) 10
Hezhen 8
Japanese 28
Lahu 8
Miao 10
Mongola 10
Naxi 7
Oroqen 9
She 10
Tu 10
Tujia 10
Xibo 9
Yakut 25
Yi 10
Oceania 26
Melanesian 9
Papuan 17
America 39
Colombian 6
Karitiana 5
Maya 14
Pima 13
Surui 1
Genotypes on 660,918 SNPs typed previously in 1,043 individuals from the
Human Genome Diversity Panel (10) were submitted to quality control pro-
cedures described in ref. 39. In particular, we excluded 409 SNPs with >10%
missing data among 1,043 individuals, 67 monomorphic SNPs, 696 SNPs with
fewer than five alleles present in at least 1 of 52 worldwide populations, and
641 autosomal SNPs with departures from Hardy–Weinberg equilibrium.
After removing these 1,813 SNPs, 659,105 SNPs remained for analysis,
642,563 of which were autosomal. We excluded relatives from the dataset
of 1,043 on which SNP quality control was conducted, leaving 938 unrelated
individuals (10), among whom 872 had STR data available.
Edge et al. www.pnas.org/cgi/content/short/1619944114 8 of 9
Table S2. Allelic imputation accuracies and expected
heterozygosities for 13 CODIS loci
Beagle imputation Null imputation Expected
Locus accuracy accuracy heterozygosity
D18S51 0.326 0.294 0.877
FGA 0.411 0.342 0.869
D21S11 0.509 0.381 0.850
D8S1179 0.589 0.397 0.828
TH01 0.826 0.411 0.794
VWA 0.537 0.417 0.810
D13S317 0.606 0.450 0.807
D16S539 0.610 0.461 0.787
D7S820 0.631 0.475 0.792
D5S818 0.601 0.498 0.772
D3S1358 0.592 0.523 0.742
CSF1PO 0.603 0.541 0.725
TPOX 0.849 0.585 0.691
Mean 0.591 0.444 0.796
The Beagle and null accuracies are taken from Fig. 1. Expected heterozy-
gosities are taken from figure 1A of ref. 11. Less heterozygous loci tend to
produce higher accuracies (for Beagle accuracies, Pearson r = −0.746, t =
−3.71, and p = 0.0034; for null accuracies, r = −0.973, t = −13.96, and p =
2.42 × 10−8).
Table S3. Mean match scores for matching and nonmatching pairs of individuals subdivided by geographic region
Source of STR profile
Source of nonmatching SNP profile Sample size Africa Europe Middle East Central/South Asia East Asia Oceania America
America 15 −17.14 −19.45 −18.55 −18.22 −18.42 −13.19 −10.21
Oceania 3 −14.03 −20.83 −18.72 −18.01 −18.63 −15.83 −15.06
East Asia 59 −15.26 −18.73 −17.50 −17.92 −15.86 −16.32 −16.26
Central/South Asia 56 −14.73 −17.53 −17.04 −17.36 −18.45 −16.21 −16.78
Middle East 41 −15.49 −18.56 −18.33 −19.17 −20.67 −17.86 −19.22
Europe 30 −14.64 −15.62 −15.55 −16.14 −18.60 −14.29 −15.93
Africa 14 −19.11 −27.23 −26.04 −27.45 −28.47 −24.39 −27.00
Mean across matching profiles 0.24 8.40 8.90 9.02 9.25 1.95 13.52
Mean across matching profiles minus 19.35 24.01 27.23 26.38 25.11 17.77 23.73
mean across nonmatching profile pairs
from the same region
Numbers are all calculated based on the values plotted in the match-score matrix in Fig. 2A. Each mean is a mean of all matching or nonmatching pairs of
matrix entries from a specific pair of geographic regions. Note that Oceania has a small sample size in these computations.
Edge et al. www.pnas.org/cgi/content/short/1619944114 9 of 9
You might also like
- Laporan FSE May 2022Document14 pagesLaporan FSE May 2022Andre PrimaNo ratings yet
- SCE Matrix InstructionsDocument4 pagesSCE Matrix InstructionsBrian FinnertyNo ratings yet
- PapersWithImage FinalExamHistoryDocument6 pagesPapersWithImage FinalExamHistoryJames Brian Garcia GarayNo ratings yet
- Ariel Feedback ReportDocument19 pagesAriel Feedback ReportFenias BoaneNo ratings yet
- What Is Life? A Guide To Biology: Second EditionDocument62 pagesWhat Is Life? A Guide To Biology: Second Editionanother dbaNo ratings yet
- Achievement Chart TemplateDocument4 pagesAchievement Chart TemplateedmontiaNo ratings yet
- National Core Arts Standards LKDocument4 pagesNational Core Arts Standards LKMary MaxwellNo ratings yet
- UntitledDocument16 pagesUntitledvishnu jNo ratings yet
- Preanalytical Quality Improvement - Dr. Yenny Surjawan, SP - PK, PH.DDocument55 pagesPreanalytical Quality Improvement - Dr. Yenny Surjawan, SP - PK, PH.Dgonteng sadyogaNo ratings yet
- Thai 11203514 AsmDocument22 pagesThai 11203514 AsmThái TranNo ratings yet
- Program Bus PitDocument1 pageProgram Bus PitTudor AlinNo ratings yet
- Acordes e InversõesDocument2 pagesAcordes e Inversõesbrunoluis1No ratings yet
- 3 BenzDocument1 page3 BenzRay LatinoNo ratings yet
- Hypnobirthing Study AustraliaDocument7 pagesHypnobirthing Study AustraliaRia Ulfha IndrianyNo ratings yet
- 070 EN Leak Detection11Document36 pages070 EN Leak Detection11jack macNo ratings yet
- SOIC-Top 10 Mid-Caps in IndiaDocument52 pagesSOIC-Top 10 Mid-Caps in IndiaPrateekNo ratings yet
- Case Study EdFinancial Services Achieves NIST Compliance For Service AccountsDocument2 pagesCase Study EdFinancial Services Achieves NIST Compliance For Service AccountsRadu CostinNo ratings yet
- Crawford County Drug Overdose Fatality Review's 2021 ReportDocument13 pagesCrawford County Drug Overdose Fatality Review's 2021 ReportGere Goble0% (1)
- EVR General Product CatalogueDocument16 pagesEVR General Product CatalogueAnthonyNo ratings yet
- National Standards For Music Education LKDocument1 pageNational Standards For Music Education LKMary Maxwell100% (1)
- The Problem and A Review of Related LiteratureDocument36 pagesThe Problem and A Review of Related Literaturechariot1729No ratings yet
- Amorn Ngammongkolrat - Food Industry in Thailand1Document22 pagesAmorn Ngammongkolrat - Food Industry in Thailand1Girish S ReddyNo ratings yet
- Zhou Bicycle Company Cal State La CompressDocument11 pagesZhou Bicycle Company Cal State La CompressjohanhadiwNo ratings yet
- Alvin Answer Sheet02Document16 pagesAlvin Answer Sheet02Dileep Kumar MotukuriNo ratings yet
- Week 11 Data AnalyticsDocument36 pagesWeek 11 Data AnalyticsJoya Labao Macario-BalquinNo ratings yet
- Customer-Facing Project Portfolio DashboardDocument11 pagesCustomer-Facing Project Portfolio DashboardAwais ZafarNo ratings yet
- 03 AlvinthePM - Mock Exam01 Answer SheetDocument18 pages03 AlvinthePM - Mock Exam01 Answer SheetDileep Kumar MotukuriNo ratings yet
- Ar Ar: Er-Softw Er-SoftwDocument1 pageAr Ar: Er-Softw Er-SoftwKitsune AovNo ratings yet
- Valve ListDocument1 pageValve ListRains MpcNo ratings yet
- Dashboard Project Management 1Document19 pagesDashboard Project Management 1Cris demaalaNo ratings yet
- UntitledDocument31 pagesUntitledDiego QuinteroNo ratings yet
- Pre and Post Test Forms and Answer KeyDocument6 pagesPre and Post Test Forms and Answer KeyLaura ElenaNo ratings yet
- Frenchik VIN vPICDocument14 pagesFrenchik VIN vPICBharghava Sasanka DesuNo ratings yet
- Tabel Elemen MesinDocument144 pagesTabel Elemen MesinRio Pradipta100% (3)
- R6 QR'FTR: DR+ I & (T (Itr"fr FTRDocument3 pagesR6 QR'FTR: DR+ I & (T (Itr"fr FTRtapuNo ratings yet
- Akashdeep Singh Stock Analysis AssignmentDocument7 pagesAkashdeep Singh Stock Analysis Assignmentapi-644716103No ratings yet
- Motor Starting MethodsDocument1 pageMotor Starting MethodsAhmed Abd El WahabNo ratings yet
- Nemesis Instructiones (ES)Document24 pagesNemesis Instructiones (ES)Rebeca CH DobantonNo ratings yet
- Surface Anatomy: Clinical Correlations: - Gray's Pp. 200-208Document14 pagesSurface Anatomy: Clinical Correlations: - Gray's Pp. 200-208speedy.catNo ratings yet
- Route 292 6 22 03 MAPDocument1 pageRoute 292 6 22 03 MAPJason BentleyNo ratings yet
- CGAXIS 28 Trees IIIDocument31 pagesCGAXIS 28 Trees IIIAKASH SINGHNo ratings yet
- DM 011 S 2023 ONLINE ORIENTATION ON THE CONDUCT OF NAT FOR GR 12 SY 2022 2023 01029Document4 pagesDM 011 S 2023 ONLINE ORIENTATION ON THE CONDUCT OF NAT FOR GR 12 SY 2022 2023 01029James CorpuzNo ratings yet
- Combined Science: Paper 3Document16 pagesCombined Science: Paper 3Vinieysha LoganathanNo ratings yet
- Drug Presentation Evaluation FormDocument1 pageDrug Presentation Evaluation FormHemantNo ratings yet
- GOW Debate Scoring SheetDocument3 pagesGOW Debate Scoring SheetBishal ThakurNo ratings yet
- Online Travel Agency (OTA) Market Share in Europe 2017 - StatistaDocument3 pagesOnline Travel Agency (OTA) Market Share in Europe 2017 - StatistaPermaculture Farm FamilyNo ratings yet
- Methods of Calculating Total Organic Carbon PDFDocument7 pagesMethods of Calculating Total Organic Carbon PDFJanaMaulanaSupriatnaNo ratings yet
- Mohamed AliDocument3 pagesMohamed AliMohamed AliNo ratings yet
- Study Stone Durability 2019Document4 pagesStudy Stone Durability 2019mz zhenNo ratings yet
- Schoolinterim 2 ReportDocument1 pageSchoolinterim 2 Reportapi-354407024No ratings yet
- Co Un T) : Shree Amar Kalyan Secondary School, Maijogmai 1 Nayabazar, IlamDocument4 pagesCo Un T) : Shree Amar Kalyan Secondary School, Maijogmai 1 Nayabazar, IlamDB BhandariNo ratings yet
- Concept in Engineering DesignDocument1 pageConcept in Engineering DesignAdheshNo ratings yet
- Modelo de RevestimentosDocument1 pageModelo de RevestimentosMarcos RodriguesNo ratings yet
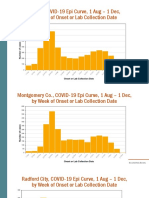
- NRHD, COVID-19 Epi Curve, 1 Aug - 1 Dec, by Week of Onset or Lab Collection DateDocument6 pagesNRHD, COVID-19 Epi Curve, 1 Aug - 1 Dec, by Week of Onset or Lab Collection DatePat ThomasNo ratings yet
- RGD Ebara Direct FiredDocument12 pagesRGD Ebara Direct FiredMuhammed HusseinNo ratings yet
- It's So Easy Going Green: An Interactive, Scientific Look at Protecting Our EnvironmentFrom EverandIt's So Easy Going Green: An Interactive, Scientific Look at Protecting Our EnvironmentNo ratings yet
- 2005 - Recommendations For DX and TTMT of Acute PorphyriasDocument13 pages2005 - Recommendations For DX and TTMT of Acute PorphyriasDavidArangoNo ratings yet
- PGLO GFP Purification Electrophoresis and ChromatographyDocument37 pagesPGLO GFP Purification Electrophoresis and ChromatographyImaniah Bazlina WardaniNo ratings yet
- Conrph 2022Document56 pagesConrph 2022Ming GamonganNo ratings yet
- Ucu Curriculum Planning Premed All Courses FellowDocument3 pagesUcu Curriculum Planning Premed All Courses FellowPaula Medio TorrubianoNo ratings yet
- Class X Science Practice Test Ak 2022-23Document15 pagesClass X Science Practice Test Ak 2022-23Tanish MehtaNo ratings yet
- 6m PracExmA.B 1 Unit 7 EcologyDocument19 pages6m PracExmA.B 1 Unit 7 Ecologywalid ben aliNo ratings yet
- MCQ Chapter 1 Page 2Document27 pagesMCQ Chapter 1 Page 2Leen Alodat100% (1)
- Plastida Mitokondria 11 PDFDocument29 pagesPlastida Mitokondria 11 PDFelsaNo ratings yet
- Scientists That Contributed in The Field of EmbryologyDocument3 pagesScientists That Contributed in The Field of EmbryologyJhestine AbayanNo ratings yet
- Apocalypse Humans ProjectDocument18 pagesApocalypse Humans ProjectaubweeNo ratings yet
- Exit Exam G10.Document6 pagesExit Exam G10.John Keneth Baysac CadeliniaNo ratings yet
- Enzymes:: "Helper" Protein MoleculesDocument27 pagesEnzymes:: "Helper" Protein MoleculesLyan Joy PalmesNo ratings yet
- Producing Malonate in Saccharomyces Cerevisiae Via The Alanine Pathwaysystems Microbiology and BiomanufacturingDocument11 pagesProducing Malonate in Saccharomyces Cerevisiae Via The Alanine Pathwaysystems Microbiology and BiomanufacturingWendy SarmientoNo ratings yet
- BIOLOGY 10 REVIEWER - 3rd QuarterDocument10 pagesBIOLOGY 10 REVIEWER - 3rd QuarterGeromme TudNo ratings yet
- Haematology Dissertation TopicsDocument6 pagesHaematology Dissertation TopicsCollegePapersWritingServiceCanada100% (1)
- Full Download Genetic Analysis An Integrated Approach 3rd Edition Sanders Test BankDocument15 pagesFull Download Genetic Analysis An Integrated Approach 3rd Edition Sanders Test Bankdopemorpheanwlzyv100% (42)
- WS4 EvolutionDocument2 pagesWS4 Evolutiondelossantosprincejoshua3No ratings yet
- Stephen Bailey - Academic Writing - A Handbook For International Students-Routledge (2004) - 34-36Document3 pagesStephen Bailey - Academic Writing - A Handbook For International Students-Routledge (2004) - 34-36selma gpdNo ratings yet
- Extra Nuclear InheritanceDocument22 pagesExtra Nuclear InheritanceCeya JoseNo ratings yet
- NNPC Aptitude Test Past Questions and Answers PDFDocument105 pagesNNPC Aptitude Test Past Questions and Answers PDFADEWOLE EMANUELNo ratings yet
- Polymorphism in CnidariaDocument7 pagesPolymorphism in Cnidariabhavak bajajNo ratings yet
- 11.5 Speciation Through IsolationDocument3 pages11.5 Speciation Through IsolationOmar AlwaerNo ratings yet
- 2024-3.0 Hour Review Test-4 (Gen - 1) - Paper - 1Document14 pages2024-3.0 Hour Review Test-4 (Gen - 1) - Paper - 1RishuNo ratings yet
- New - Micro-2023-Part-1Document15 pagesNew - Micro-2023-Part-1Trisha RomeoNo ratings yet
- Bacterial Cytology and Physiology - BacteriologyDocument6 pagesBacterial Cytology and Physiology - BacteriologyMary Joy HersaliaNo ratings yet
- Secondary MetabolitesDocument4 pagesSecondary MetabolitesEliud NgangaNo ratings yet
- 978 1 62703 038 0 PDFDocument546 pages978 1 62703 038 0 PDFNelson Hernan Parada RoaNo ratings yet
- Jurnal SepsisDocument13 pagesJurnal SepsisDwx PrasetyaNo ratings yet
- Biology B - Paper 1 (9BI0 - 01)Document40 pagesBiology B - Paper 1 (9BI0 - 01)Priya Kumar100% (1)
- Psych 1106-01 Assignment 1 - Anthonisha Martin and Renee Gauthier 1Document7 pagesPsych 1106-01 Assignment 1 - Anthonisha Martin and Renee Gauthier 1api-535493143No ratings yet